Cutaneous Melanoma is a form of skin cancer characterized by an elevated mutational load that favors high spread to distant organs and resistance to therapies. The identification of biomarkers, able to dissect normal and pathogenic biological processes and response to therapeutic intervention, is necessary to describe melanoma as accurately as possible, having a positive impact on early diagnosis, in turn selecting the best therapeutic option. Recently, a great number of new biomarkers were evaluated, in order to identify those patients who may have clinical benefit from a therapeutic choice, particularly for immunotherapy. At present, these new biomarkers wait to be validated before clinical use. Hence, the requirement to look at and periodically update the advances in this field.
1. Introduction
In the last decade, research in melanoma treatment saw two historical moments, defined in 2011 by the FDA approval of ipilimumab and vemurafenib for BRAF proto-oncogene (BRAF) mutated metastatic melanoma treatment, and in 2014 by the authorization of pembrolizumab and nivolumab for unresectable and metastatic melanoma. These two different approaches, targeted therapy and immunotherapy, are able to attack the advanced stages of melanoma that, until those dates, accounted for the majority of skin cancer related death, although representing less than 5% of all cutaneous malignancies
[1].
This new picture in the control of advanced melanoma, even if representing a milestone in cancer treatment, still evidences some dark zones, since patients suffer from drug resistance or are non-responders, and progression free or overall survival are still low
[2]. Certainly, we need to increase knowledge of the molecular pathways responsible for melanoma pathogenesis and progression, but also understand why effectiveness of therapies is so different among patients. Just after a full comprehension of the underlying specificities, we will be able to select the more promising therapeutic approach recognizing the risk of disease progression or the actual chance of response.
At diagnosis, 85% of patients have circumscribed disease, 15% show regional lymph nodal involvement, about 2–5% present distant metastases
[3]. Determination of melanoma staging results from a complex histopathological examination of lesions that also considers the absence of specific and sensitive biomarkers with exclusive features and that is capable of obtaining univocal impact for prognosis assessment and treatment planning.
The American Joint Committee on Cancer (AJCC) has recently revised melanoma classification based on Breslow thickness. In this implementation, patients can be stratified in risk category by the TNM (tumor, lymph node, metastasis) system, individuating four stages, stage I–IV, comprising different localizations of the disease with different impact on the overall survive and profound effects on prognosis. Stage I and II identify localized disease, while stage III and IV diagnose disease progression with nodal or distant metastases respectively. In AJCC classification, primary tumors are also divided in four classes (T1–T4) based of tumor thickness, again subdivided in a or b for absence or presence of ulcerations. Nodal category indicates whether melanoma has moved from the primary site to the vicinal lymph nodes and is individuated by different subgroups, designated by 0 to 3, indicating the absence (N0) or presence of lymph nodal metastasis. With M are classified only melanoma that have developed distant metastases and represent the most advanced staging of disease
[4]. In this context, the accuracy and improvement of melanoma diagnosis, staging and risk assessment are essential for adequate prognostication, eventually allowing for the best therapeutic choice. Classical diagnostic and prognostic methods have evidenced accuracy failure in patient stratification. However, if we look at the last 20 years, the evolution of melanoma biomarkers discovery went from around 900 biomarker publications in 1999 to 51,000 in 2019
[5]. Indeed, the recent combination with new omics technologies from tumor or sera of patients, have produced an impressive increment of new possible biomarkers. However, all these potential biomarkers need to be investigated in large-scale studies for validation and standardization.
2. Prognostic and Predictive Biomarkers
To complete the presentation of biomarkers for melanoma, categorization is essential to define a group of them capable to dissect the impact on tumor biology and disease course (prognostic biomarkers) and a second group defining tumor response to treatment (predictive biomarkers) as well as the improvement in overall survival (OS), disease free survival (DFS) and progression free survival (PFS). The last aspects are essential to define those patients that can have clinical benefit from therapeutic choice, particularly for immunotherapy that, if it has revolutionized the melanoma cure, still presents many questions to be addressed related to the limited efficacy and high toxicity
[6]. To date, many possible biomarkers have been evaluated essentially in retrospective studies. Therefore, in the AJCC staging system, intrinsic limitations in the number of the categorized biomarkers were observed. Thus, many different parameters are normally utilized to obtain more accurate disease prognosis and prediction, although their implementation in larger data set is necessary to support a validation for routine procedures in clinical practice.
In primary melanoma, the prognosis is strongly dependent on tumor thickness, ulceration, mitotic rate, vasculature development, presence of tumor-infiltrating lymphocytes (TILs) and melanoma subtype. Other characteristics are intrinsic to the patients, such as age, sex and anatomical site of the lesion. In general, young and females are associated with a more favorable prognosis.
2.1. Genetic Prognostic Factors
The genetic background of a tumor, beyond addressing patients to specific treatment, has a clear prognostic value. BRAF mutation is the most common mutation of CM, generally affecting young people and characterized by decrease of CD8
+ number and increased release of immunosuppressive cytokines. BRAF mutation displays unfavorable prognostic value compared to wild type melanomas, although may benefit of approved combination therapy with BRAF and MEK inhibitors (dabrafenib plus trametinib or vemurafenib plus cobimetinib). Unfortunately, these patients are often subjected to acquired resistance
[7][8][9]. NRAS mutations identify a more aggressive disease and these mutations are predictor of poorer outcomes in view of the lack of any novel specific therapy, being progress limited to MEK inhibitors with modest clinical benefit in PFS and substantially inefficacious to increase the OS
[10].
Other molecular markers with important prognostic and predictive value are guanine nucleotide-binding protein alpha subunits G(q) /11 (GNAQ/11) and TP53 mutations. GNAQ/11 are involved in signaling via G-protein-coupled receptors. GNAQ/11 mutations, found in uveal melanoma and in little percentage of the so-called triple wild type subtype of CM, induce overexpression of RAS Guanyl releasing protein 3 (RasGRP3) with consequent constitutive activation of RAS, event associated with poor OS. Currently, therapies for treatment of this mutation are essentially inadequate
[11]. Associated with poor OS and PFS, the TP53 mutations are present in 20% of CM and characterized by senescence evasion and autophagy activation. TP53 mutations characterize older group of patients, mainly presenting head and neck melanomas
[12]. Familial melanoma susceptibility is a sporadic event that occur in 2% of all melanomas and the incidence strongly increases with increased number of familiar cases. The screening among the familial population of the CDKN2A mutation is necessary and essential
[13].
2.2. Prognostic Factors in Lymphangiogenesis
Melanoma cells secrete, among others, some vascular endothelial growth factors (VEGF), namely VEGF-C and -D that, together with their receptor VEGFR3, have been proposed as lymphangiogenetic markers, albeit with contradictory results
[14]. Furthermore, lymphatic invasion is characterized by cancer cells ability to adhere to endothelial cells and to migrate along the lymphatic vasculature. In this regard, protein phosphatase 2 regulatory subunit A (PPP2R1A), responsible of active interaction between melanoma and lymphatic endothelial cells, has been proposed as a new biomarker in melanoma metastatization
[15]. Overall, the immunohistochemical approach, although very expensive and time-consuming, provides a representative picture of the lymphohematological status of the primary tumor, yielding information for diagnosis and prognosis. In recent years, however, researchers have begun to look at lymph as a greater reservoir of cancer biomarkers compared to plasma liquid biopsy. In this respect, the use of postoperative lymphatic exudate from metastatic melanoma patients is becoming a powerful non-invasive clinical practice for the identification of tumor-derived factors, including extracellular vesicles charged of proteins and miRNAs reminiscent of metastatic progression
[16][17].
2.3. Lymph Node Prognostic Role
Sentinel lymph node status was indicated as the major prognostic factor for disease free survival and choice of therapy in adjuvant setting
[18]. Lymph nodal involvement correlated with tumor thickness and their biopsy was justified for thick melanomas (>4 mm), in presence of ulceration, tumor lymphocyte infiltration and mitotic rate >1 mm
2. In the recent past, the possibility that patients with lymph nodal metastatic disease could benefit from complete surgical eradication of lymph nodal station/basin has generated a deep scientific debate that produced a number of studies
[19]. Among these studies, two important international multicenter trials led to similar conclusions. In these trials, there appears to be no survival benefit associated with complete lymph node dissection, but for a major regional node control of the disease
[20][21]. Consequently, surgical lymph node dissection for patients with sentinel lymph node-positive melanoma is no longer routinely recommended, mainly when melanoma had spread to distant sites (stage III and IV), also considering the major efficacy of targeted and immune strategies introduced in the adjuvant setting
[22].
2.4. Prognostic and Predictive Biomarkers and Immunotherapy
The recent use of ICBs as standard therapy for advanced melanomas evidenced efficacy only in a limited number of patients and adverse events, developed in some patients, were so severe to induce therapy blockage. Thus, it is now essential to characterize and validate standard biomarkers predicting response and toxicity before treatment, in order to achieve the best potential clinical benefit (, ). In melanoma, the high mutational burden correlates with more effective immunotherapy, considering that different somatic mutations can generate immunogenic tumor neoantigens able to sustain an immune response in ICB-responder melanoma patients
[23]. Riaz and coworkers demonstrated that during immunotherapy with Nivolumab, compared to baseline, the tumor mutation burden decreased in responder patients. Furthermore, in these patients, the loss of a number of certain neoantigens proportionally reflexed the clonal expansion of T cell populations
[24]. However, this association was able to predict improved survival but not treatment responses. In this respect, a transcriptomic study in biopsies of pretreated melanoma revealed that the mutational loads improved survival independently from the capacity to respond to immunotherapy, whereas responder patients were characterized by an enrichment for mutations in the DNA repair gene BRCA2. On the contrary, non-responder patients showed a transcriptional signature, referred as innate anti-programed cell death 1 (PD-1) resistance, with high expression of genes involved in regulation of epithelial mesenchymal transition, extracellular matrix remodeling, cell adhesion and angiogenesis ()
[23]. Several investigations found a direct correlation between gene expression associated to the interferon γ (IFNγ) pathway, T cell inflamed tumor microenvironment with abundant expression of chemokines and response to immune checkpoint blockers. Other studies indicated the deficiency of ICB clinical efficacy in melanomas characterized by PTEN mutations and functional deficiency of the IFNγ signaling triggered by loss-of-function mutations in Janus kinase JAK1/2 and β2 microglobulin ()
[25][26][27].
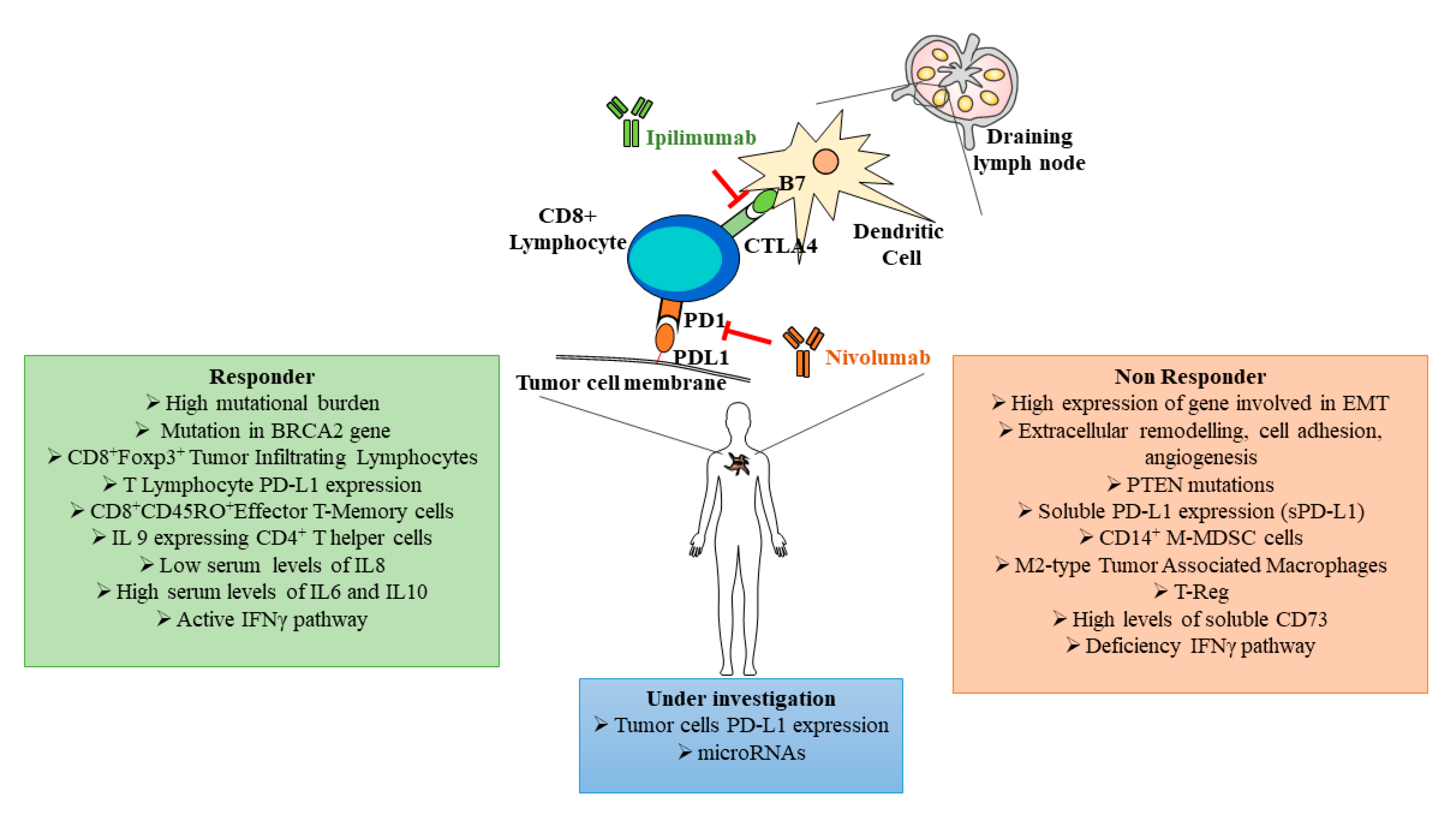 Figure 2.
Figure 2. Schematic representation of some key immune checkpoint blocker functional links and predictive biomarkers associated to responder (green box) and non-responder (red box) patients. In the blue box are indicated melanoma biomarkers still under investigation. PD-L1: programed cell death ligand 1, PD-1: programed cell death 1, sPD-L1: soluble programed cell death ligand 1, CTLA4: cytotoxic T-lymphocyte associated protein 4, BRCA2: breast cancer type 2 susceptibility protein, Foxp3: forkhead box p3, IL: interleukin, IFN: interferon, EMT: epithelial–mesenchymal transition, PTEN: phosphatase and tensin homolog, MDSC: myeloid derived suppressor cells, M-MDSC: monocyte myeloid derived suppressor cells; T-Reg: regulatory T cells.

 Figure 2. Schematic representation of some key immune checkpoint blocker functional links and predictive biomarkers associated to responder (green box) and non-responder (red box) patients. In the blue box are indicated melanoma biomarkers still under investigation. PD-L1: programed cell death ligand 1, PD-1: programed cell death 1, sPD-L1: soluble programed cell death ligand 1, CTLA4: cytotoxic T-lymphocyte associated protein 4, BRCA2: breast cancer type 2 susceptibility protein, Foxp3: forkhead box p3, IL: interleukin, IFN: interferon, EMT: epithelial–mesenchymal transition, PTEN: phosphatase and tensin homolog, MDSC: myeloid derived suppressor cells, M-MDSC: monocyte myeloid derived suppressor cells; T-Reg: regulatory T cells.
Figure 2. Schematic representation of some key immune checkpoint blocker functional links and predictive biomarkers associated to responder (green box) and non-responder (red box) patients. In the blue box are indicated melanoma biomarkers still under investigation. PD-L1: programed cell death ligand 1, PD-1: programed cell death 1, sPD-L1: soluble programed cell death ligand 1, CTLA4: cytotoxic T-lymphocyte associated protein 4, BRCA2: breast cancer type 2 susceptibility protein, Foxp3: forkhead box p3, IL: interleukin, IFN: interferon, EMT: epithelial–mesenchymal transition, PTEN: phosphatase and tensin homolog, MDSC: myeloid derived suppressor cells, M-MDSC: monocyte myeloid derived suppressor cells; T-Reg: regulatory T cells.



