Changes in structural and functional neuroplasticity have been implicated in various neurological disorders. Sterol regulatory element-binding protein (SREBP)-1c is a critical regulatory molecule of lipid homeostasis in the brain. Recently, our findings have shown the potential involvement of SREBP-1c deficiency in the alteration of novel modulatory molecules in the hippocampus and occurrence of schizophrenia-like behaviors in mice. However, the possible underlying mechanisms, related to neuronal plasticity in the hippocampus, are yet to be elucidated. In this study, we investigated the hippocampus-dependent memory function and neuronal architecture of hippocampal neurons in SREBP-1c knockout (KO) mice. During the passive avoidance test, SREBP-1c KO mice showed memory impairment. Based on Golgi staining, the dendritic complexity, length, and branch points were significantly decreased in the apical cornu ammonis (CA) 1, CA3, and dentate gyrus (DG) subregions of the hippocampi of SREBP-1c KO mice, compared with those of wild-type (WT) mice. Additionally, significant decreases in the dendritic diameters were detected in the CA3 and DG subregions, and spine density was also significantly decreased in the apical CA3 subregion of the hippocampi of KO mice, compared with that of WT mice. Alterations in the proportions of stubby and thin-shaped dendritic spines were observed in the apical subcompartments of CA1 and CA3 in the hippocampi of KO mice. Furthermore, the corresponding differential decreases in the levels of SREBP-1 expression in the hippocampal subregions (particularly, a significant decrease in the level in the CA3) were detected by immunofluorescence.
Alterations in neuronal structure and function have been implicated in the pathophysiology of various neurological disorders
[1]. In the brain, the hippocampus is one of the few remaining regions capable of undergoing neuronal plasticity even during adulthood, rendering it especially susceptible to structural and functional alterations
[2][3]. These hippocampal alterations further translate to abnormalities of emotions, learning, and memory
[4]. These hippocampal abnormalities in volume
[5][6], synaptic circuitry
[7], and cytoarchitecture
[8] have been reported in patients with neurological diseases, including schizophrenia. Rodent models of hippocampal lesions also demonstrate congruent pharmacological, anatomical, and behavioral phenotypes of schizophrenia
[9].
Lipids are vital for brain function as they constitute the bulk of the brain and they function in cellular signaling, myelin generation, and neuroplasticity
[10]. The brain is largely dependent on de novo lipid synthesis as the blood–brain barrier limits the exogenous lipid transport into the brain
[11][12]. Sterol regulatory element-binding proteins (SREBPs) are considered the master regulators of lipid synthesis
[13]. SREBPs are transcription factors, controlling both cholestero- and lipogenesis, via the activation of enzymatic cascades required for the synthesis of endogenous cholesterol, fatty acids, and triglycerides
[14]. In mammals, three SREBP isoforms have been described, of which SREBP-1a and SREBP-1c are both produced from different transcription sites (exon 1a and exon 1c) of the same
Srebf-1 gene
[15], while SREBP-2 originates from the
Srebf-2 gene
[16]. SREBP-1c is the predominant isoform expressed in most tissues, including the white adipose tissue, liver, and brain. SREBP-1a is predominantly expressed in cellular proliferative tissues, such as the spleen and intestine
[17], while SREBP-2 is ubiquitously expressed throughout the body at amounts approximately equal to the total of the SREBP-1a and 1c expression
[18]. Although there is a commonality in their activities, SREBP-1c favors the control of fatty acid and triglyceride synthesis, while SREBP-2 mainly regulates cholesterol synthesis and SREBP-1a does not display any particular preference
[13].
Previously, we found that mice lacking SREBP-1c demonstrate schizophrenia-like behavioral symptoms
[19]. Furthermore, we found that novel genes (e.g., glucagon-like peptide 2 receptor (
Glp2r), necdin (
Ndn), and Erb-B2 receptor tyrosine kinase 4) were significantly altered in the hippocampus of SREBP-1c knockout (KO) mice, compared with those in wild-type (WT) mice
[20]. However, the mechanisms possibly underlying these abnormal behaviors and altered molecular expressions need further examination. Thus, we sought to investigate the potential changes in functional and structural neuroplasticity of the hippocampus of SREBP-1c KO mice, which may provide new insights into the role of SREBP-1c in the hippocampus and its potential relevance to the development of animal experimental systems for neurological disorders such as schizophrenia.
1. SREBP-1c KO Mice Demonstrate Significantly Decreased Hippocampus-Dependent Memory Retention
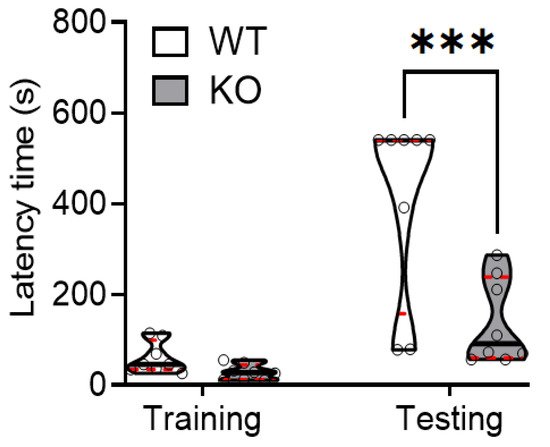
Previously, we observed that SREBP-1c KO mice had altered behaviors suggestive of the positive and negative symptoms of schizophrenia, but not its cognitive symptoms. To further assess cognitive symptoms, we used the passive avoidance task as a hippocampus-dependent paradigm. During the training period, both WT and KO mice had short cross-over latencies before entering the dark compartment where they received a foot shock. Interestingly, during the testing period, memory retention was significantly reduced in KO mice, compared with WT mice. This was evidenced by the significantly shorter increase in the cross-over latency during the testing period (Figure 1).
Figure 1. SREBP-1c deficiency significantly impairs learning and memory function in mice. Using the passive avoidance test, WT and SREBP-1c KO mice were trained using mild electrical foot shocks. Both groups were tested 24 h after training. Cross-over latency (s) was assessed to evaluate memory retention. SREBP-1c KO mice showed a significantly lesser increase in the cross-over latency before entering the dark room during testing than WT mice (WT: 406 ± 208.77 s, KO: 138.3 ± 94.40 s, n = 8 per group; Finteraction (1,14) = 10.17, p = 0.0002). The upper and lower dashed red lines signify the upper and lower quartiles, respectively, and the median is represented by a solid black line within a violin plot. *** p < 0.001 vs. WT. WT, wild-type littermate; KO, SREBP-1c KO group.
2. SREBP-1c KO Mice Display Altered Dendritic Morphology in the Hippocampus
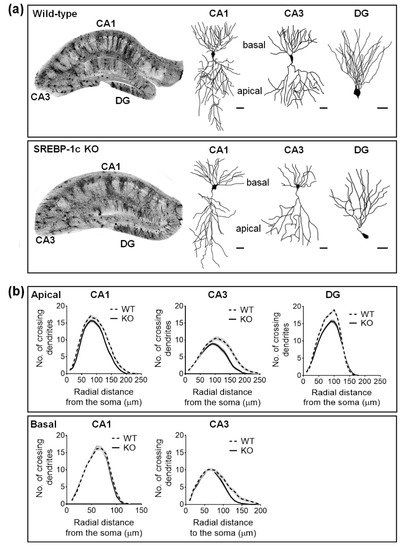
Sholl analysis was used to quantify the dendritic complexity of the Golgi-stained neurons. The total number of dendritic intersections was recorded per 10 µm incremental radial distance from the soma. Overall, SREBP-1c KO mice, compared with WT mice, had decreased dendritic complexity (Figure 2a). The complexity was significantly reduced in apical CA1, apical and basal CA3, and DG, but not in the basal subcompartment of CA1 (Figure 2b). Apical CA1 dendrites in SREBP-1c KO mice demonstrated significantly fewer intersections than WT mice at the Sholl radii of 130–150 μm from the soma (Figure 2b, upper left line graphs). The number of intersections of the CA3 dendrites in SREBP-1c KO mice, compared with those in WT mice, were significantly reduced in the apical subcompartment at the Sholl radii of 110–180 μm from the soma (Figure 2b, upper middle line graphs) and in the basal subcompartment at the Sholl radii of 110–140 μm from the soma (Figure 2b, lower right line graphs). The DG granule cell dendrites of KO mice also had significantly less intersections than those of WT mice at Sholl radii of 70–110 μm from the soma (Figure 2b, upper right line graphs). However, no significant differences between SREBP-1c KO and WT mice in the basal compartment of the CA1 subregion were observed (Figure 2b, lower left line graphs). Table 1 shows the results of the two-way ANOVA tests for the effects of the genotype and distance on dendritic arborization (mean no. of crossing dendrites) in each hippocampal subregion of WT and KO mice measured by Sholl analysis (n = 40 neurons/group).
Figure 2. SREBP-1c KO mice display impaired dendritic complexity in the CA1, CA3, and DG subregions of the hippocampus. The figures show the representative images of the Golgi-stained hippocampus and subregional neurons in WT and SREBP-1c KO mice (a). The line graphs show the mean number of intersections per 10 µm radial unit distance from the soma (0) for apical ((b); upper) and basal ((b); lower) dendrites. Data are expressed as means ± standard errors (SEs) of 10 neurons of each subregion in each mouse (n = 40 dendrites/group). The scale bars in (a) represent 25 µm. CA, cornu ammonis; DG, dentate gyrus; WT, wild-type littermate; KO, SREBP-1c KO group.
Table 1. Results of two-way ANOVA tests for the effects of the genotype and distance on dendritic arborization (mean no. of crossing dendrites) in each hippocampal subregion of WT and KO mice measured by Sholl analysis (n = 40 neurons/group).
| |
Sholl Radii Distance (μm) 1 |
Šidăk’s Post Hoc Test per Sholl Radii Distance |
Two-Way ANOVA |
| CA1 apical |
130, 140, 150 |
p = 0.0001, p = 0.0007, p = 0.0121 |
Finteraction (23, 1794) = 1.741, p = 0.0161 |
| basal |
n.s. |
n.s. |
Finteraction (12, 936) = 0.8516, p = 0.5969 |
| CA3 apical |
110, 120, 130, 140, 150, 160, 170, 180 |
p = 0.0371, p = 0.0046, p = 0.0004, p < 0.0001, p < 0.0001, p < 0.0001, p = 0.0004, p = 0.0257 |
Finteraction (24,1872) = 3.985, p < 0.0001 |
| basal |
110, 120, 130, 140 |
p = 0.0243, p = 0.0023, p = 0.0002, p = 0.0490 |
Finteraction (19,1482) = 2.098, p = 0.0037 |
| DG |
70, 80, 90, 100, 110 |
p = 0.0014, p = 0.0149, p = 0.0189, p < 0.0001, p = 0.0025 |
Finteraction (17, 1326) = 4.031, p < 0.0001 |
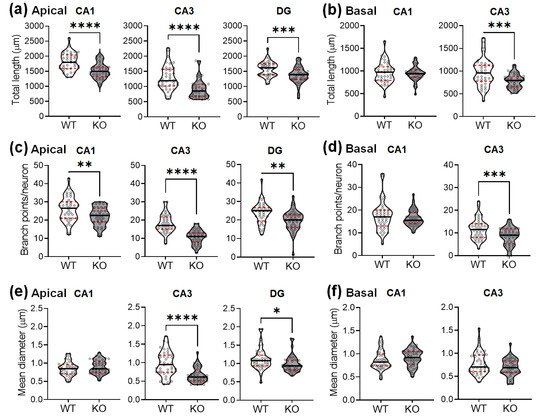
The total dendritic length was also significantly reduced in the hippocampi of SREBP-1c KO mice, compared with those of WT mice (Figure 3a,b). Deficiencies were observed in the apical CA1 (Figure 3a, left violin plots), apical CA3 (Figure 3a, middle violin plots), DG subregions (Figure 3a, right violin plots), and basal CA3 (Figure 3b, right violin plots), but not in the basal subcompartment of CA1 (Figure 3b, left violin plots).
Figure 3. SREBP-1c KO mice show decreases in the length, number of branch points, and diameter of the dendrites in the hippocampus. Violin plots show the total length (a,b), number of branch points (c,d), and mean diameter (e,f) of the dendrites in each subregion (CA1, CA3, and DG). The upper and lower dashed red lines signify the upper and lower quartiles, respectively, and the median is represented by a solid black line within the violin plots. * p < 0.05, ** p < 0.01, *** p < 0.001, and **** p < 0.0001 vs. WT (n = 40 dendrites/group). CA, cornu ammonis; DG, dentate gyrus; WT, wild-type littermate; KO, SREBP-1c KO group.
A corresponding decrease in the dendritic branching points was also observed in the hippocampi of KO mice compared with those of WT mice (Figure 3c,d). The branch points per neuron were reduced in the apical CA1 (Figure 3c, left violin plots), apical CA3 (Figure 3c, middle violin plots), DG subregions (Figure 3c, right violin plots), and basal CA3 (Figure 3d, right violin plots), but not in the basal subcompartment of CA1 (Figure 3d, left violin plots).
The Golgi-stained neurons were also examined for variations in the dendritic thickness in WT and KO mice (Figure 3e,f). The dendritic diameters were significantly reduced in the apical CA3 (Figure 3e, middle violin plots) and DG subregions (Figure 3e, right violin plots) of KO mice, but not in the other areas (CA1 apical: Figure 3e, left violin plots; CA1 basal: Figure 3f, left violin plots; CA3 basal: Figure 3f, right violin plots).
Table 2 shows the results of Student’s t-tests for total dendritic lengths, branch points, and thickness of the hippocampal neurons in each subregion between WT and KO mice (n = 40 neurons/group).
Table 2. Results of Student’s t-tests for total dendritic lengths, branch points, and thickness of the hippocampal neurons in each subregion between WT and KO mice (n = 40 neurons/group).
| |
WT |
KO |
Student’s t-Test |
| Total dendritic length (μm) |
| CA1 apical |
1811.3 ± 296.7 |
1516 ± 251.8 |
t(78) = 4.794, p < 0.0001 |
| basal |
964.3 ± 229.5 |
940.8 ± 160.4 |
t(78) = 0.531, p = 0.5971 |
| CA3 apical |
1281 ± 398 |
924 ± 308.2 |
t(78) = 4.482, p < 0.0001 |
| basal |
967.5 ± 301.2 |
783 ± 156.1 |
t(78) = 3.440, p = 0.0009 |
| DG |
1576 ± 245.1 |
1374 ± 278.5 |
t(78) = 3.448, p = 0.0009 |
| Dendritic branch points |
| CA1 apical |
25.9 ± 6.7 |
22.0 ± 5.0 |
t(78) = 2.956, p = 0.0041 |
| basal |
17.3 ± 6.3 |
16.4 ± 3.6 |
t(78) = 0.719, p = 0.4741 |
| CA3 apical |
18.1 ± 5.1 |
10.6 ± 3.9 |
t(78) = 7.201, p < 0.0001 |
| basal |
11.8 ± 4.5 |
8.5 ± 3.9 |
t(78) = 3.469, p = 0.0009 |
| DG |
28.3 ± 5.7 |
19.7 ± 5.5 |
t(78) = 3.282, p = 0.0015 |
| Dendritic thickness (μm) |
| CA1 apical |
0.845 ± 0.190 |
0.890 ± 0.185 |
t(78) = 1.031, p = 0.3056 |
| basal |
0.864 ± 0.206 |
0.916 ± 0.189 |
t(78) = 1.168, p = 0.2462 |
| CA3 apical |
0.976 ± 0.325 |
0.664 ± 0.202 |
t(78) = 5.162, p < 0.0001 |
| basal |
0.775 ± 0.244 |
0.689 ± 0.190 |
t(78) = 1.760, p = 0.0823 |
| DG |
1.117 ± 0.291 |
0.994 ± 0.228 |
t(78) = 2.013, p = 0.0387 |
3. SREBP-1c KO Mice Show Altered Dendritic Spine Density and Morphology in the Hippocampus
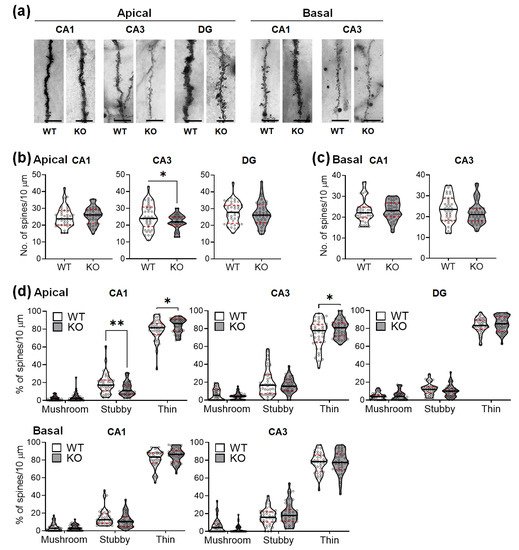
The Golgi-stained neurons were magnified and analyzed for possible changes in the dendritic spines. The mice lacking SREBP-1c had significant alterations in spine density and morphology only in the apical subcompartment of the CA1 and CA3 subregions (Figure 4). The spine densities (number of spines per 10 μm) were not altered in the apical CA1 (Figure 4b, left violin plots), DG (Figure 4b, right violin plots), basal CA1 (Figure 4c, left violin plots), and basal CA3 (Figure 4c, right violin plots), although they significantly decreased in the apical CA3 subregions (Figure 4b, middle violin plots). Table 3 shows the results of Student’s t-tests for the dendritic spine density (per dendritic segments of 10 μm) of the hippocampal neurons in each subregion between WT and KO mice (n = 40 dendritic segments/group).
Figure 4. SREBP-1c KO mice show altered dendritic spine density and morphological proportions in the hippocampus. Representative images from each cohort show dendritic segments and their spines along Golgi-impregnated neurons (a). Violin plots show the total number of spines in each subregion (CA1, CA3, and DG) (b,c). Violin plots show the proportions of the spines categorized by morphology in each subregion (CA1, CA3, and DG) (d). Upper and lower dashed red lines signify the upper and lower quartiles, respectively, and the median is represented by a solid black line within a violin plot. * p < 0.05, ** p < 0.01 vs. WT (n = 40 dendritic segment/group). The scale bars in the dendritic images represent 5 µm. CA, cornu ammonis; DG, dentate gyrus; WT, wild-type littermate; KO, SREBP-1c KO group.
Table 3. Results of Student’s t-tests for the dendritic spine density (per dendritic segments of 10 μm) of the hippocampal neurons in each subregion between WT and KO mice (n = 40 dendritic segments/group).
| |
WT |
KO |
Student’s t-Test |
| CA1 apical |
24.8 ± 6.0 |
25.7 ± 5.1 |
t(78) = 0.7302, p = 0.4675 |
| basal |
23.1 ± 5.7 |
23.2 ± 4.5 |
t(78) = 0.0146, p = 0.9884 |
| CA3 apical |
24.9 ± 7.5 |
22.0 ± 4.4 |
t(78) = 2.108, p = 0.0382 |
| basal |
23.6 ± 6.2 |
21.9 ± 5.5 |
t(78) = 1.362, p = 0.1770 |
| DG |
27.7 ± 6.5 |
27.1 ± 7.1 |
t(78) = 0.4203, p = 0.6754 |
Changes in the proportions of different dendritic spine morphologies were observed in the apical CA1 and apical CA3 subregions. Stubby spines were significantly reduced in the apical CA1 subregion (Figure 4d, upper left violin plots), but not in the other areas. In contrast, the thin spines were significantly increased in the apical CA1 (Figure 4d, upper left violin plots) and apical CA3 subregions (Figure 4d, upper middle violin plots), but not in the other subregions. No significant changes in the proportion of mushroom-shaped spines were observed. Table 4 shows the results of two-way ANOVA tests for the effects of the genotype and spine morphology on the proportion of dendritic spines in each hippocampal subregion of WT and KO mice (n = 40 dendritic segments (10 μm)/group).
Table 4. Results of two-way ANOVA tests for the effects of genotype and spine morphology on the proportion of dendritic spines in each hippocampal subregion of WT and KO mice (n = 40 dendritic segments (10 μm)/group).
| |
Mushroom Spines (%) |
Stubby Spines (%) |
Thin Spines (%) |
Two-Way ANOVA |
| CA1 apical |
| WT |
2.2 ± 2.6 |
18.7 ± 11.2 |
79.1 ± 11.8 |
Finteraction (2, 156) = 5.126, p = 0.0070 |
| KO |
3.4 ± 4.9 |
13.0 ± 7.8 |
83.6 ± 7.9 |
| |
p = 0.9026 1 |
p = 0.0082 1 |
p = 0.0487 1 |
|
| CA1 basal |
| WT |
4.0 ± 4.7 |
14.9 ± 9.3 |
81.0 ± 10 |
Finteraction (2, 156) = 3.003, p = 0.0525 |
| KO |
3.3 ± 3.2 |
11.7 ± 7.9 |
84.9 ± 8.2 |
| |
p = 0.9630 1 |
p = 0.1753 1 |
p = 0.0647 1 |
|
| CA3 apical |
| WT |
6.4 ± 6.3 |
19.7 ± 14.8 |
73.8 ± 15.8 |
Finteraction (2, 156) = 3.231, p = 0.0422 |
| KO |
3.9 ± 3.9 |
16.2 ± 8.4 |
79.9 ± 9.8 |
| |
p = 0.6465 1 |
p = 0.3666 1 |
p = 0.0357 1 |
|
| CA3 basal |
| WT |
6.3 ± 7.8 |
16.3 ± 9.4 |
77.4 ± 11.5 |
Finteraction (2, 156) = 1.599, p = 0.2054 |
| KO |
2.7 ± 4.3 |
19.7 ± 12 |
77.6 ± 12.8 |
| |
p = 0.3040 1 |
p = 0.3488 1 |
p = 0.9997 1 |
|
| DG |
| WT |
4.2 ± 3.9 |
12.3 ± 6.7 |
83.4 ± 8.3 |
Finteraction (2, 156) = 1.224, p = 0.2970 |
| KO |
5.1 ± 5.2 |
9.9 ± 6.8 |
84.9 ± 8.7 |
| |
p = 0.9285 1 |
p = 0.3283 1 |
p = 0.6861 1 |
|
4. The CA3 Subregion of SREBP-1c KO Mice Shows Significantly Greater Decreases in Dendritic and Spine Morphological Parameters Compared to Those of Other Subregions
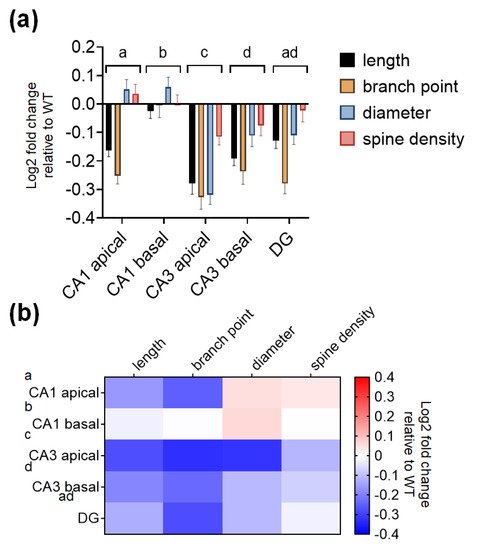
To determine if there are subregion-specific differences in the degree of morphological alterations observed, the log2 fold changes in WT mice were computed for each structural parameter. Of the different subregions, the CA3 subregion had the greatest decreases in dendritic length, number of branch points, diameter, and spines compared with the CA1 subregion and DG subregion (Figure 5a, bar graphs). Table 5 shows the results of two-way ANOVA tests for the effects of the hippocampal subregion and morphological parameters on log2 fold changes in KO mice relative to WT mice. Table 6 shows the summary of P-values from Šidăk’s post hoc analyses for the comparison of the different hippocampal subregions in Table 5. To better visualize the subregion-specific differences in the morphological changes, the same set of data is expressed as a heatmap in Figure 5b.
Figure 5. SREBP-1c KO mice show greater morphological alterations in the CA3 than in the other subregions of the hippocampus. Fold changes in dendritic and spine morphological characteristics in WT mice are represented as bar graphs (a) and heatmaps (b) to better visualize the degree of differences between the different hippocampal subregions (CA1, CA3, and DG). Data are expressed as means ± SEs, and different letters, “a,” “b,” “c,” and “d,” indicate statistically significant differences between the subregions at p < 0.05 (n = 40 neuron or dendrites/group). Bars with no common letters are significantly different (p < 0.05). CA, cornu ammonis; DG, dentate gyrus; WT, wild-type littermate group.
Table 5. Results of two-way ANOVA tests for the effects of the hippocampal subregion and morphological parameters on log2 fold changes in KO mice relative to WT mice.
| |
Dendritic Length |
Dendritic Branch Point |
Dendritic Diameter |
Spine Density |
Two-Way ANOVA |
| CA1 apical |
−0.16 ± 0.022 |
−0.25 ± 0.029 |
0.051 ± 0.035 |
0.037 ± 0.032 |
Finteraction (12, 780) = 3.718, p < 0.0001 |
| basal |
−0.02 ± 0.026 |
−0.004 ± 0.043 |
0.06 ± 0.035 |
0.0007 ± 0.031 |
| CA3 apical |
−0.28 ± 0.038 |
−0.33 ± 0.042 |
−0.32 ± 0.033 |
−0.12 ± 0.028 |
| basal |
−0.19 ± 0.026 |
−0.24 ± 0.045 |
−0.11 ± 0.039 |
−0.08 ± 0.037 |
| DG |
−0.13 ± 0.028 |
−0.28 ± 0.035 |
−0.11 ± 0.032 |
−0.02 ± 0.04 |
Table 6. Summary of the P-values from Šidăk’s post hoc analyses for the comparison of the different hippocampal subregions in Table 5.
| |
CA1 Apical |
CA1 Basal |
CA3 Apical |
CA3 Basal |
DG |
| CA1 apical |
|
0.0024 |
<0.0001 |
0.0330 |
0.2481 |
| CA1 basal |
|
|
<0.0001 |
<0.0001 |
<0.0001 |
| CA3 apical |
|
|
|
0.0001 |
<0.0001 |
| CA3 basal |
|
|
|
|
0.9978 |
| DG |
|
|
|
|
|
5. SREBP-1c KO Mice Show Significantly Reduced SREBP-1 Protein Expression in the CA3 Subregion of the Hippocampus
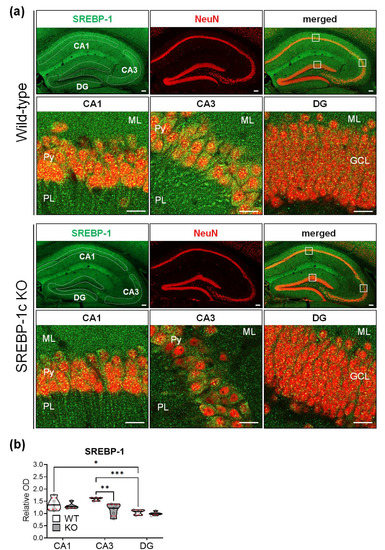
To evaluate the protein expression of SREBP-1 in the hippocampus, the sections were subjected to immunofluorescence staining, and the relative immunoreactive intensities were analyzed using ImageJ software (National Institutes of Health, Bethesda, MD, USA). The SREBP-1 protein was differentially expressed in the subregions of the hippocampi of WT and KO mice; the expressions were most intense in CA3, followed by CA1 and DG (Figure 6a,b). The relative intensities of SREBP-1 expression in the CA1 and DG subregions did not substantially differ in KO and WT mice. However, a significant decrease in SREBP-1 expression was detected in the CA3 subregion compared with that of WT mice of KO mice (Figure 6b, violin plots). Table 7 shows the results of two-way ANOVA tests for the effects of the genotype and hippocampal subregion on the relative immunoreactivity for SREBP-1 in WT and KO mice (n = 5 hippocampi/group).
Figure 6. SREBP-1c KO mice show different levels of SREBP-1 protein expressions in each subregion of the hippocampus. Representative immunofluorescence images from each group show expressions of SREBP-1 (green) and NeuN (red)—a neuron-specific marker—in the various hippocampal subregions (CA1, CA3, and DG) (a). KO mice showed relatively lesser levels of SREBP-1 expression in all subregions, especially in the CA3 subregion; SREBP-1 was mainly expressed in the pyramidal and granular cell layers of the CA and DG subregions, respectively. Violin plots show the relative intensities of SREBP-1 expression in the pyramidal cell layers (CA1 and CA3) and granular cell layer (DG) in each of the hippocampal subregions (CA1, CA3, and DG) of WT and KO mice (b). Upper and lower dashed red lines represent the upper and lower quartiles, respectively, and the median is represented by a solid black line within a violin plot. * p < 0.05, ** p < 0.01, *** p < 0.001 vs. WT (n = 5 animals/group). The scale bars represent 100 and 20 µm in the hippocampal and subregion images, respectively. CA, cornu ammonis; DG, dentate gyrus; GCL, granular cell layer; ML, molecular layer; PL, polymorphic cell layer; Py, pyramidal cell layer; SREBP, sterol regulatory element-binding protein; NeuN, neuronal nuclei; WT, wild-type littermate; KO, SREBP-1c KO group.
Table 7. Results of two-way ANOVA tests for the effects of the genotype and hippocampal subregion on the relative immunoreactivity for SREBP-1 in WT and KO mice (n = 5 hippocampi/group).
| |
WT |
KO |
Šidăk’s Post Hoc Test |
Two-Way ANOVA |
| CA1 |
1.40 ± 0.25 1 |
1.31 ± 0.13 1 |
p = 0.8182 |
Genotype: F(1, 8) = 6.392, p = 0.0354
Subregion: F(2, 16) = 18.86, p < 0.0001
Interaction: F(2, 16) = 5.725, p = 0.0133 |
| CA3 |
1.59 ± 0.07 1 |
1.15 ± 0.25 1 |
p = 0.001 |
| DG |
1.05 ± 0.10 1 |
1 ± 0.08 1 |
p = 0.9429 |