Your browser does not fully support modern features. Please upgrade for a smoother experience.

Submitted Successfully!
Thank you for your contribution! You can also upload a video entry or images related to this topic.
For video creation, please contact our Academic Video Service.
| Version | Summary | Created by | Modification | Content Size | Created at | Operation |
|---|---|---|---|---|---|---|
| 1 | Mei-Ru Chen | + 2005 word(s) | 2005 | 2021-05-18 07:51:48 | | | |
| 2 | Peter Tang | Meta information modification | 2005 | 2021-07-08 11:31:14 | | |
Video Upload Options
We provide professional Academic Video Service to translate complex research into visually appealing presentations. Would you like to try it?
Cite
If you have any further questions, please contact Encyclopedia Editorial Office.
Chen, M. Nuclear Import of EBV Proteins. Encyclopedia. Available online: https://encyclopedia.pub/entry/11777 (accessed on 07 February 2026).
Chen M. Nuclear Import of EBV Proteins. Encyclopedia. Available at: https://encyclopedia.pub/entry/11777. Accessed February 07, 2026.
Chen, Mei-Ru. "Nuclear Import of EBV Proteins" Encyclopedia, https://encyclopedia.pub/entry/11777 (accessed February 07, 2026).
Chen, M. (2021, July 07). Nuclear Import of EBV Proteins. In Encyclopedia. https://encyclopedia.pub/entry/11777
Chen, Mei-Ru. "Nuclear Import of EBV Proteins." Encyclopedia. Web. 07 July, 2021.
Copy Citation
Epstein–Barr virus (EBV) is a human pathogen that infects most of the world’s population and is associated with several well-known malignancies. Within the nucleus, the replicated viral DNA is packaged into capsids, which subsequently egress from the nucleus into the cytoplasm for tegumentation and final envelopment. There is increasing evidence that viral lytic gene expression or replication contributes to the pathogenesis of EBV. Various EBV lytic proteins regulate and modulate the nuclear envelope structure in different ways, especially the viral BGLF4 kinase and the nuclear egress complex BFRF1/BFRF2.
Epstein–Barr virus
BGLF4 kinase
nuclear egress
BFRF1
nuclear envelope modulation
1. The Structure and Function of the Nuclear Envelope
In higher eukaryotic cells, the nucleus is surrounded by an envelope composed of the inner and outer nuclear membranes, the perinuclear space between the membranes, and the underlying nuclear lamina network (Figure 1A). These three-layer structures are held in place by the nuclear pore complex (NPC), which functions as a transport gatekeeper for macromolecules trafficking in or out of the nucleus. The outer nuclear membrane is continuous with the rough endoplasmic reticulum. The nuclear lamina is a thin, electron-dense meshwork lining the nucleoplasmic face of the inner nuclear membrane (INM) [1][2] and provides firm structural support for the major components of the nuclear envelope (NE) [3][4]. The integrity of the envelope provides the structural support for nuclear morphology and functions as a platform for coordinating several cellular processes, including DNA repair, cell signaling, migration, and gene expression. The lamina is connected to the inner nuclear membrane through INM proteins (emerin and lamin B receptor), the chromatin proteins (histone H2A/H2B), and also the cytoskeleton-interacting LINC (linker of nucleoskeleton and cytoskeleton) complexes (reviewed in [5]). The physical linkages of LINC are instrumental in the plasticity of cellular organization and are required for processes such as nuclear migration and anchorage, meiotic chromosome movements, centrosome positioning, and the global organization of the cytoskeleton.
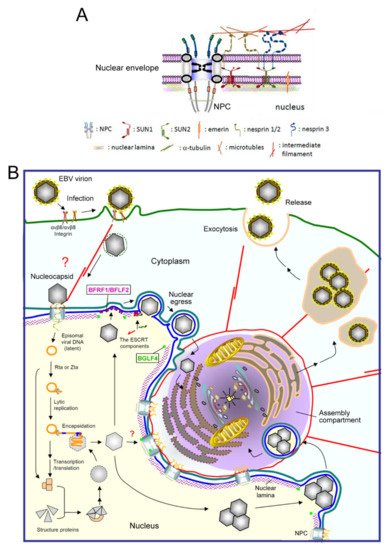
Figure 1. A diagram of EBV virion replication, production, and maturation. (A) An overview of the nuclear envelope (NE) structure with selected NE-associated proteins and nuclear pore complexes. The NE is composed of two lipid bilayers, inner and outer nuclear membranes (INM and ONM), and the nuclear lamina formed by lamin filaments. The nuclear pore complexes (NPCs) not only serve as gates for material transport, but also structurally hold the membranes and lamina together. The ONM, which is continuous with the ER, is characterized by cytoskeleton-associated nesprin proteins tethered by SUN1 and SUN2 in the INM. Emerin is an INM integrated protein that anchors on the nuclear lamina. (B) After EBV infects cells, the viral genome is injected into the nucleus through nuclear pore complexes. The linear genome is circularized into the episomal form through the terminal repeats and maintained as an episomal form in the latently infected cells. With chemical stimulations or reactivation signals, the viral transactivator Zta or Rta activates replication-related viral proteins and lytic viral DNA replication. Along with the production and assembly of viral capsid proteins, the viral genome is encapsidated into the preformed procapsid to form the mature nucleocapsid. Simultaneously, the viral BGLF4 protein kinase mediates the phosphorylation and partial disassembly of nuclear membrane-underlying nuclear lamina, allowing the access of nucleocapsids to the nuclear membrane. In our observations, the nucleocapsids then traverse through the nuclear envelope with the coordination of the viral nuclear egress complex BFRF1/BFLF2, cellular ESCRT machinery, and Nedd4-like ubiquitin ligases. The cytoplasm-distributed nucleocapsids may subsequently transport into the juxtanuclear “viral assembly compartment”. This specialized compartment, containing highly reorganized membrane structures and organelles, may provide a site for efficient tegumentation and secondary envelopment of the nucleocapsids. Finally, the mature virion is transported close to the cell margins and released from the infected cells by exocytosis. NPC, nuclear pore complex.
NE integrity is essential for cell homeostasis. Alteration of NE structure or components may impact the behavior and the phenotypes of cells in many ways. For decades, deleterious morphological changes of the NE in tumor cells have been recognized as an important parameter for the malignant criteria of tumors [5][6][7]. Many NE-associated diseases, collectively designated nuclear envelopathies or laminopathies, have been identified. Most of these disorders come from mutations in INM proteins or their interacting partners, diseases including muscular dystrophy, lipodystrophy, neuropathy, and progeria (premature aging) syndromes [8][9]. Diseases may be due to mutations or altered expression of alleles, the mechanical stability of the nucleus, or cell cycle regulation. However, the molecular mechanisms that give rise to these diseases and whether pathogen infection may modulate the nuclear structure and contribute to the pathogenesis remain poorly understood [10][11].
2. Nuclear Pore Complexes (NPC) Function as Gates and Guardians for Nuclear–Cytoplasmic Transport
The gatekeeper function of the NPC is important for preventing the abnormal distribution of cytoplasmic components or genome-toxic insults to the cellular chromatin. The NPC has a molecular mass of ~125 megaDaltons (MDa) in vertebrates and is composed of multiple copies of a set of 30 diverse proteins, termed nucleoporins (NUPs) [12]. In the middle of the nuclear pore, FG-NUPs, containing Phe-Gly (FG) repeats, play the most relevant role in controlling the nuclear transport receptors (NTR) that shuttle through the NPC. FG-NUPs and NTRs interact through hydrophobic contacts for the translocation step of nuclear transport through the NPC. The canonical transport of most macromolecules (>40 kDa) into the nucleus is controlled by nuclear localization signals (NLS) that are recognized by specific members of the importin receptor family. Proteins containing a bipartite motif or a stretch of basic amino acids bind to an importin alpha and beta heterodimeric receptor complex [13]. During transport, importin α binds to the NLS-bearing protein, and importin β mediates the binding of the transport complex to the FG-NUPs, which subsequently slide through the NPC [14][15]. The small GTPase Ran regulates the association and dissociation of the importin-cargo complex between GTP- and GDP-bound forms [16][17]. For the export of nuclear protein, XPO1 (Exportin-1/Chromosome Region Maintenance 1/CRM1) is the most known mediator. Nuclear export signals (NESs) are short leucine-rich sequences that can be found in many shuttling proteins. Cargo proteins containing NES are recognized by XPO1, which interacts with nucleoporins NUP214 and NUP88 in NPC to transport out of the nucleus in a RanGAP-dependent manner [18][19].
Nuclear transport is also an important step that controls specific signaling pathways. For example, activators of transcription, including IRF3 and NF-κB, are regularly expressed and sequestered in the cytoplasm. They are translocated into the nucleus through phosphorylation-mediated regulation and activate specific innate immune response genes upon stimulation [20][21]. Such regulation could occur in as little as 30 min to activate gene transcription in response to challenge by various pathogens.
3. Overview of EBV Lytic Replication
Epstein–Barr virus (EBV) is a human gamma-herpesvirus that infects most of the world’s population. It has a 170–175 kb linear double-stranded DNA genome which encodes approximately 90 open reading frames. The genome is packaged within an icosahedral capsid, approximately 100–120 nm in diameter, surrounded by a proteinaceous tegument and a lipid bilayer envelope. EBV generally establishes two phases in its life cycle, latency and lytic replication [22]. EBV latent proteins help B cell proliferation, including LMP1 (latent membrane protein 1), which can mimic CD40 signaling to activate NF-κB signaling, and EBNA2 (EBV nuclear antigen 2), which can turn on Myc expression. EBV can transform primary B cells into lymphoblastoid cell lines in vitro and may cause post-transplantation lymphoproliferative disease (PTLD). Increasing evidence indicates that lytic phase gene expression also contributes to EBV oncogenesis [23].
EBV mainly infects naïve B lymphocytes and epithelial cells through different receptors, including CR2, HLA class II, and integrins [24]. B cell entry may occur via membrane fusion or endocytosis followed by fusion of the viral membrane with the membrane of the endocytic vesicle [25]. Ephrin receptor A2 was recently identified as the EBV receptor on epithelial cells [24][26][27]. The various steps of EBV replication are illustrated in Figure 1B. After the initial infection, mediated by sequential interactions of viral glycoproteins with host surface receptors, the nucleocapsid is internalized through endocytosis or membrane fusion [28][29][30]. It has been postulated that the nucleocapsid might undergo some structural changes and slide along the microtubule or actin cytoskeleton to the NPC boundary [31]. The linear viral genome is then injected into the nucleus [32]. However, how the EBV genome gets through the NPC and whether viral factors modify the NPC structure to facilitate transport of the viral genome remain unclear. The genome is then circularized and maintained by the EBV encoded nuclear antigen EBNA1 as an episomal form in the nucleus, remaining latent in the host [24].
Upon chemical or stress stimulation, or Ig cross-linking on the surface of B cells, EBV is reactivated and genome replication ensues. Viral lytic genes are expressed in a temporal and sequential order that is divided into three stages, immediate-early (IE), early (E), and late (L). Two immediate-early transactivators, Zta and Rta, are transcribed before viral protein synthesis and can turn on complete viral lytic replication [22]. Most of the early genes are required for viral DNA replication, such as the polymerase processivity factor BMRF1 and CDK1 (cyclin-dependent kinase 1)-like protein kinase BGLF4. The late viral products are mainly structural proteins that are expressed after viral DNA synthesis, such as viral capsid protein VCA and glycoprotein gp350/220 [33]. After DNA replication, the viral genome is packaged into the preformed capsid in the nucleus. With the coordination of cellular machinery and the viral egress complex BFRF1 and BFLF2, the intranuclear nucleocapsids subsequently move towards the inner leaflet of the nuclear membrane and bud from the NE [34][35]. In our unpublished observations, the transported nucleocapsids may obtain their tegument and secondary envelope at the “cytoplasmic assembly compartment”, a specialized cell compartment containing highly reorganized membrane apparatus, cell organelles, and cytoskeletons. Finally, the mature virions containing the cis-Golgi derived membrane are transported and released from the cell through exocytosis [36] (Figure 1B).
4. The Nuclear Import of EBV Proteins
To initiate viral DNA replication, Zta and Rta, coordinately bind to the origin of lytic replication (oriLyt) and recruit essential lytic replication proteins to form a core replication complex [37][38]. The viral DNA replication machinery includes the DNA polymerase (BALF5), single-stranded DNA (ssDNA) binding protein (BALF2), DNA polymerase processivity factor (BMRF1), helicase (BBLF4), primase (BSLF1), primase accessory proteins (BBLF2/BBLF3), and uracil DNA glycosylase (BKRF3). Viral DNA replicates in a manner very similar to the mammalian DNA replication system [37][38][39][40]. For oriLyt binding and transactivator function, Zta contains a classical NLS for nuclear targeting. Interestingly, several EBV genes involved in viral DNA replication lack the canonical NLS signals in their coding region. For example, viral DNA polymerase (BALF5) and uracil DNA glycosylase (BKRF3) are transported into the nucleus through interaction with the DNA polymerase processivity factor (BMRF1) [39][41]. Interestingly, the three EBV primase/helicase complex components do not contain canonical NLS but can be imported into the nucleus through interaction with the origin binding protein Zta [42], suggesting EBV evolved with novel pathways for nuclear targeting. In addition, we found the EBV protein kinase BGLF4 enters the nucleus through the direct interaction of its carboxyl-terminal alpha-helical region with the FG-NUP proteins NUP60 and NUP153, independent of importins [43]. BGLF4 also modulates the structure and transport preference of the NPC to facilitate the nuclear import of several EBV lytic proteins, including all three primase/helicase complex proteins and the viral capsid protein BcLF1 [44]. We postulated that the novel nuclear targeting mechanism and BGLF4-mediated NPC preference may interfere with the cellular antiviral response and ensure dominance of the virus for efficient replication.
5. The Nuclear Export (Egress) of EBV Nucleocapsids
After genome replication, EBV DNA is packaged into the preassembled procapsid, which primarily consists of the major capsid protein, VCA, and is held together by 320 triplexes formed by two minor capsid proteins, BDLF1 and BORF1, and a small capsid protein, BFRF3, on the hexameric capsomers [45]. The nucleocapsid size is about 100–130 nm, too large to go through the nucleopore. The NE structure is a significant hindrance for the transport of the nucleocapsid into the cytoplasm. Our and others’ studies demonstrated that at least three virus-encoded proteins, including BGLF4 protein kinase and nuclear egress complex BFRF1/BFLF2, modify the NE structure to facilitate the nucleocytoplasmic transport of nucleocapsids.
References
- Fawcett, D.W. On the occurrence of a fibrous lamina on the inner aspect of the nuclear envelope in certain cells of vertebrates. Am. J. Anat. 1966, 119, 129–145.
- Gruenbaum, Y.; Margalit, A.; Goldman, R.D.; Shumaker, D.K.; Wilson, K.L. The nuclear lamina comes of age. Nat. Rev. Mol. Cell Biol. 2005, 6, 21–31.
- McKeon, F.D.; Kirschner, M.W.; Caput, D. Homologies in both primary and secondary structure between nuclear envelope and intermediate filament proteins. Nature 1986, 319, 463–468.
- Moir, R.D.; Spann, T.P.; Goldman, R.D. The dynamic properties and possible functions of nuclear lamins. Int. Rev. Cytol. 1995, 162B, 141–182.
- Ungricht, R.; Kutay, U. Mechanisms and functions of nuclear envelope remodelling. Nat. Rev. Mol. Cell Biol. 2017, 18, 229–245.
- Chow, K.H.; Factor, R.E.; Ullman, K.S. The nuclear envelope environment and its cancer connections. Nat. Rev. Cancer 2012, 12, 196–209.
- de Las Heras, J.I.; Batrakou, D.G.; Schirmer, E.C. Cancer biology and the nuclear envelope: A convoluted relationship. Semin. Cancer Biol. 2013, 23, 125–137.
- Romero-Bueno, R.; de la Cruz Ruiz, P.; Artal-Sanz, M.; Askjaer, P.; Dobrzynska, A. Nuclear Organization in stress and aging. Cells 2019, 8, 664.
- Stiekema, M.; van Zandvoort, M.; Ramaekers, F.C.S.; Broers, J.L.V. Structural and mechanical aberrations of the nuclear lamina in disease. Cells 2020, 9, 1884.
- Worman, H.J.; Fong, L.G.; Muchir, A.; Young, S.G. Laminopathies and the long strange trip from basic cell biology to therapy. J. Clin. Investig. 2009, 119, 1825–1836.
- Liu, S.Y.; Ikegami, K. Nuclear lamin phosphorylation: An emerging role in gene regulation and pathogenesis of laminopathies. Nucleus 2020, 11, 299–314.
- Lin, D.H.; Hoelz, A. The structure of the nuclear pore complex (An update). Annu. Rev. Biochem. 2019, 88, 725–783.
- Robbins, J.; Dilworth, S.M.; Laskey, R.A.; Dingwall, C. Two interdependent basic domains in nucleoplasmin nuclear targeting sequence: Identification of a class of bipartite nuclear targeting sequence. Cell 1991, 64, 615–623.
- Gorlich, D.; Henklein, P.; Laskey, R.A.; Hartmann, E. A 41 amino acid motif in importin-alpha confers binding to importin-beta and hence transit into the nucleus. EMBO J. 1996, 15, 1810–1817.
- Weis, K.; Ryder, U.; Lamond, A.I. The conserved amino-terminal domain of hSRP1 alpha is essential for nuclear protein import. EMBO J. 1996, 15, 1818–1825.
- Sorokin, A.V.; Kim, E.R.; Ovchinnikov, L.P. Nucleocytoplasmic transport of proteins. Biochemistry 2007, 72, 1439–1457.
- Wente, S.R.; Rout, M.P. The nuclear pore complex and nuclear transport. Cold Spring Harb. Perspect. Biol. 2010, 2, a000562.
- Fornerod, M.; Ohno, M.; Yoshida, M.; Mattaj, I.W. CRM1 is an export receptor for leucine-rich nuclear export signals. Cell 1997, 90, 1051–1060.
- Fung, H.Y.; Chook, Y.M. Atomic basis of CRM1-cargo recognition, release and inhibition. Semin. Cancer Biol. 2014, 27, 52–61.
- Fitzgerald, K.A.; McWhirter, S.M.; Faia, K.L.; Rowe, D.C.; Latz, E.; Golenbock, D.T.; Coyle, A.J.; Liao, S.M.; Maniatis, T. IKKepsilon and TBK1 are essential components of the IRF3 signaling pathway. Nat. Immunol. 2003, 4, 491–496.
- Hayden, M.S.; Ghosh, S. Signaling to NF-kappaB. Genes Dev. 2004, 18, 2195–2224.
- Longnecker, R.M.; Kieff, E.; Cohen, J.I. Epstein-Barr virus. In Fields Virology, 6th ed.; Lippincott Williams & Wilkins: Philadelphia, PA, USA, 2013; p. 1898.
- Rosemarie, Q.; Sugden, B. Epstein-Barr virus: How its lytic phase contributes to oncogenesis. Microorganisms 2020, 8, 1824.
- Chen, J.; Longnecker, R. Epithelial cell infection by Epstein-Barr virus. FEMS Microbiol. Rev. 2019, 43, 674–683.
- Miller, N.; Hutt-Fletcher, L.M. Epstein-Barr virus enters B cells and epithelial cells by different routes. J. Virol. 1992, 66, 3409–3414.
- Chen, J.; Sathiyamoorthy, K.; Zhang, X.; Schaller, S.; Perez White, B.E.; Jardetzky, T.S.; Longnecker, R. Ephrin receptor A2 is a functional entry receptor for Epstein-Barr virus. Nat. Microbiol. 2018, 3, 172–180.
- Zhang, H.; Li, Y.; Wang, H.B.; Zhang, A.; Chen, M.L.; Fang, Z.X.; Dong, X.D.; Li, S.B.; Du, Y.; Xiong, D.; et al. Ephrin receptor A2 is an epithelial cell receptor for Epstein-Barr virus entry. Nat. Microbiol. 2018, 3, 1–8.
- Backovic, M.; Jardetzky, T.S.; Longnecker, R. Hydrophobic residues that form putative fusion loops of Epstein-Barr virus glycoprotein B are critical for fusion activity. J. Virol. 2007, 81, 9596–9600.
- Backovic, M.; Longnecker, R.; Jardetzky, T.S. Structure of a trimeric variant of the Epstein-Barr virus glycoprotein B. Proc. Natl. Acad. Sci. USA 2009, 106, 2880–2885.
- Grove, J.; Marsh, M. The cell biology of receptor-mediated virus entry. J. Cell Biol. 2011, 195, 1071–1082.
- Valencia, S.M.; Hutt-Fletcher, L.M. Important but differential roles for actin in trafficking of Epstein-Barr virus in B cells and epithelial cells. J. Virol. 2012, 86, 2–10.
- Chesnokova, L.S.; Jiang, R.; Hutt-Fletcher, L.M. Viral entry. In Epstein Barr Virus Volume 2: One Herpes Virus: Many Diseases; Münz, C., Ed.; Springer: Cham, Switzerland, 2015; pp. 221–235.
- Lu, C.C.; Jeng, Y.Y.; Tsai, C.H.; Liu, M.Y.; Yeh, S.W.; Hsu, T.Y.; Chen, M.R. Genome-wide transcription program and expression of the Rta responsive gene of Epstein-Barr virus. Virology 2006, 345, 358–372.
- Lee, C.P.; Chen, M.R. Escape of herpesviruses from the nucleus. Rev. Med. Virol. 2010, 20, 214–230.
- Hellberg, T.; Passvogel, L.; Schulz, K.S.; Klupp, B.G.; Mettenleiter, T.C. Nuclear egress of herpesviruses: The prototypic vesicular nucleocytoplasmic transport. Adv. Virus Res. 2016, 94, 81–140.
- Nanbo, A. Epstein-Barr Virus Exploits the Secretory Pathway to Release Virions. Microorganisms 2020, 8, 729.
- Fixman, E.D.; Hayward, G.S.; Hayward, S.D. Replication of Epstein-Barr virus oriLyt: Lack of a dedicated virally encoded origin-binding protein and dependence on Zta in cotransfection assays. J. Virol. 1995, 69, 2998–3006.
- El-Guindy, A.; Ghiassi-Nejad, M.; Golden, S.; Delecluse, H.J.; Miller, G. Essential role of Rta in lytic DNA replication of Epstein-Barr virus. J. Virol. 2013, 87, 208–223.
- Su, M.T.; Liu, I.H.; Wu, C.W.; Chang, S.M.; Tsai, C.H.; Yang, P.W.; Chuang, Y.C.; Lee, C.P.; Chen, M.R. Uracil DNA glycosylase BKRF3 contributes to Epstein-Barr virus DNA replication through physical interactions with proteins in viral DNA replication complex. J. Virol. 2014, 88, 8883–8899.
- Fixman, E.D.; Hayward, G.S.; Hayward, S.D. trans-acting requirements for replication of Epstein-Barr virus ori-Lyt. J. Virol. 1992, 66, 5030–5039.
- Kawashima, D.; Kanda, T.; Murata, T.; Saito, S.; Sugimoto, A.; Narita, Y.; Tsurumi, T. Nuclear transport of Epstein-Barr virus DNA polymerase is dependent on the BMRF1 polymerase processivity factor and molecular chaperone Hsp90. J. Virol. 2013, 87, 6482–6491.
- Gao, Z.; Krithivas, A.; Finan, J.E.; Semmes, O.J.; Zhou, S.; Wang, Y.; Hayward, S.D. The Epstein-Barr virus lytic transactivator Zta interacts with the helicase-primase replication proteins. J. Virol. 1998, 72, 8559–8567.
- Chang, C.W.; Lee, C.P.; Huang, Y.H.; Yang, P.W.; Wang, J.T.; Chen, M.R. Epstein-Barr virus protein kinase BGLF4 targets the nucleus through interaction with nucleoporins. J. Virol. 2012, 86, 8072–8085.
- Chang, C.W.; Lee, C.P.; Su, M.T.; Tsai, C.H.; Chen, M.R. BGLF4 kinase modulates the structure and transport preference of the nuclear pore complex to facilitate nuclear import of Epstein-Barr virus lytic proteins. J. Virol. 2015, 89, 1703–1718.
- Wang, W.H.; Kuo, C.W.; Chang, L.K.; Hung, C.C.; Chang, T.H.; Liu, S.T. Assembly of Epstein-Barr virus capsid in promyelocytic leukemia nuclear bodies. J. Virol. 2015, 89, 8922–8931.
More
Information
Subjects:
Virology
Contributor
MDPI registered users' name will be linked to their SciProfiles pages. To register with us, please refer to https://encyclopedia.pub/register
:
View Times:
1.1K
Revisions:
2 times
(View History)
Update Date:
08 Jul 2021
Notice
You are not a member of the advisory board for this topic. If you want to update advisory board member profile, please contact office@encyclopedia.pub.
OK
Confirm
Only members of the Encyclopedia advisory board for this topic are allowed to note entries. Would you like to become an advisory board member of the Encyclopedia?
Yes
No
${ textCharacter }/${ maxCharacter }
Submit
Cancel
Back
Comments
${ item }
|
More
No more~
There is no comment~
${ textCharacter }/${ maxCharacter }
Submit
Cancel
${ selectedItem.replyTextCharacter }/${ selectedItem.replyMaxCharacter }
Submit
Cancel
Confirm
Are you sure to Delete?
Yes
No




