
| Version | Summary | Created by | Modification | Content Size | Created at | Operation |
|---|---|---|---|---|---|---|
| 1 | Waseem Asghar | + 3788 word(s) | 3788 | 2021-07-02 08:06:23 | | | |
| 2 | Amina Yu | Meta information modification | 3788 | 2021-07-05 03:25:59 | | |
Video Upload Options
The dengue virus (DENV) is a vector-borne flavivirus that infects around 390 million individuals each year with 2.5 billion being in danger. Having access to testing is paramount in preventing future infections and receiving adequate treatment. Currently, there are numerous conventional methods for DENV testing, such as NS1 based antigen testing, IgM/IgG antibody testing, and Polymerase Chain Reaction (PCR).
1. Introduction
The DENV is a flavivirus that is primarily spread by the femaleAedes aegyptusmosquito, which lives mostly in an urban environment [1]. The first virologic proved case of dengue in the USA was in Philadelphia in 1780 [2]. However, the first dengue like symptoms in the Americas were reported in 1635 [2]. Once a non-infected mosquito bites an infected human, the incubation period lasts for 4–10 days inside mosquito [3].
The DENV from 1960 to 2010 has seen a 30-fold upsurge due to an increase in global population, global warming, ineffective mosquito control, and inadequate medical facilities [4]. Every year, around 100 million people become sick, requiring medical attention and around 22,000 people die due to the dengue virus infection globally [5]. However, currently, 70% of the endangered countries are Asian, and a large number of infections are still being recorded in countries like Bangladesh, Malaysia, Vietnam, and the Philippines [3]. Besides, 2,163,354 dengue cases of infections has been reported in Americas in 2020 with 872 deaths [6].
Some typical infection signs are a high fever running around 104°F, aches behind the eyes, severe headache, muscle/joint pain, vomiting, enflamed glands, and rash [7]. However, severe dengue fever (DF) can be potentially life-threatening in part due to plasma leaking, fluid accumulation, ascites, pleural effusions, severe bleeding, low platelets, and/or organ impairment [7]. Nevertheless, patients infected with DENV-2 & DENV-4 shows acute illness due to dengue hemorrhagic fever (DHF), but infection due to DENV-1 & DENV-3 is mild, sometimes inapparent [8]. DHF can be staged in four grades according to the guidelines of World Health Organization (WHO):
The viral load measurement by detecting genomic nucleic acids in infected patients is considered the gold standard for diagnosing dengue infection at the preliminary stage of infection [9]. As a result, IgM and IgG response from the immune system is also considered a possible medium of dengue virus diagnostic [9]. Besides, the DENV infection cocirculates with other flaviviruses, especially with Zika virus; thus, specific detection of DENV plays a vital role in DENV outbreak management [10]. Most of the developed tests are not suitable for resource limited settings although DENV is distributed mostly in countries with limited resources.
In this review, we discuss the current conventional technologies utilized for testing DENV at resource available settings. Moreover, we also discuss the currently developed state of the art biosensing technologies, evaluated their performance and outline strategies to address challenges posed by disease and provide future guidelines for the improved usage of diagnostics during recurrence or future outbreaks of DENV. We focus mainly on the most recent advancements in POC based biosensing assays that were developed since 2015. We also highlight recent advancements in dengue sensing, including CRISPR-based methods, which were not covered previously.
2. Dengue Structure
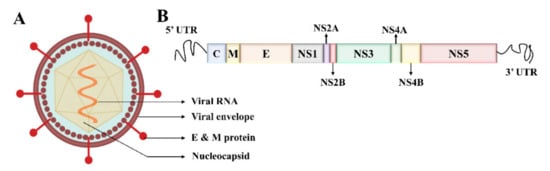
Electron microscopy imaging demonstrates that DENV has a relatively smooth surface, 40–60 nm size, and contains 25–30 nm nucleocapsid protein covered with a lipid bilayer (Figure 1A) [11][12]. The DENV is a positive-sense ssRNA virus that can be translated directly to proteins [1]. The complete genome is ~11 kb-long, encoding for three structural and seven nonstructural proteins, and the subtypes of DENV share about 65% of the complete amino acid sequences (Figure 1B) [1][13]. The virus is structured with Envelope/E protein with 495 amino acids and consists of three domains that mainly interact with the host cell for invasion [14].
3. Conventional Methods of Laboratory Diagnostics
Serological tests are most widely used to detect dengue infection due to low cost and operative simplicity compared to molecular or culture-based testing. However, plasma and whole blood samples are used occasionally as testing specimens. IgG level persists in the human body after primary infection. It can exhibit inconclusive results for secondary infection and false positive in those with infection/immunization against other flaviviruses (West Nile virus (WNV), yellow fever virus (YFV), or Zika virus (ZIKV)).
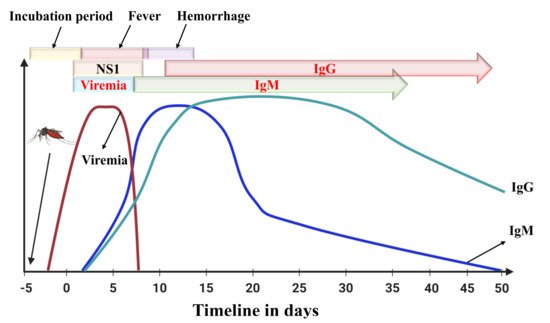
Antibodies are proteins generated by the immune system to fight against the attack of an antigen [15]. Immunoglobulin proteins, for example Ig(G, M, A, E, and D) are conventional antibodies along with B-cells and T-cells are produced in response to antigens. For devices that detect Dengue antibodies, the major problem is that samples are collected before the five-day negative window period (Figure 2). An enzyme-linked immunosorbent assay capturing the IgM antibody (MAC-ELISA) test, first developed by the Armed Forces Research Institute of Medical Sciences, can be utilized to sense IgM antibodies in dengue [16].
There are numerous commercially available MAC-ELISA testing kits that are developed and marketed for testing dengue infection globally. To perform the test, usually, the dengue IgM antibody is captured from the patient’s serum specimen by anti-human IgM antibodies on a solid phase. It provides qualitative detection by capturing the human IgM antibody to DENV recombinant antigens on a 96 well plate [17]. The first clinical study showed that 2–3 days of onset fever and 4–5 days of onset fever showed only a 28.7% (2–3 days) and 40.3% (4–5) chance of a positive result.
There are several inherent drawbacks to these IgM rapid tests and IgM-based ELISA tests. The low sensitivity of the POC tests can be a considerable drawback in clinics. In the United States, only one such POC test is approved, and before six days, the test is unreliable to a certain extent. Due to some cross-reactivity, a PRNT based test is needed to confirm the results in some cases.
The incubation period for dengue fever is usually five days, and the acute illness can be observed from 0–7 days of disease infection, and sometime after that period, some patients experience hemorrhagic shock. During the early stage of infection, viral load measurement can be done using RT-PCR or any other molecular detection method. Simultaneously, NS1 can also be diagnosed up to 10 days of infection by ELISA method. IgM or IgG based testing is useful to diagnose dengue infection after a few days of infection.
IgG-based tests can be used for determining past or present infections. The IgG antibody is produced after IgM, and IgG antibodies can persist for a more extended period (Figure 2), even lifelong [18]. A four-fold spike in IgG antibodies is often attributed to a recent infection [19]. Standard Diagnostics (Seoul, South Korea) and Panbio Inc (Brisbane, Australia now Alere Inc., Waltham, WA, USA) have commercialized IgG-based ELISA tests [20].
Panbio, on the other hand, had a sensitivity of 63.5 and specificity of 95.3% [20]. A study was done comparing these tests with their NS1 and IgM/IgG antibodies and observed that the IgG tests had a high rate of false positives and high cross-reactivity [21]. IgG tests can be useful in certain instances, but IgM and NS1 ELISA are more useful for acute infections. Due to cross-reactivity with other flavivirus IgG antibodies, it is challenging to determine primary infection just by testing IgG-based assays [22].
An IgM/IgG ratio test is usually differentiating test between first time and later infections [23]. A ratio greater than 1.32 is considered as primary infection, while a ratio below 1.32 is considered secondary [24]. The use of both IgM and IgG can be useful to determine the type of infection present. Depending on the IgM and IgG test performed, the cutoff ratio will vary.
The hemagglutination Inhibition (HI) test is a standard test for distinctive primary and later DENV infections, but it cannot detect early infection [25][26]. Response to a primary infection usually has low levels of antibodies, while secondary infections have high antibody titers that usually exceed 1:1280. Values below 1:1280 are usually interpreted as a primary infection [27]. Any value below 1 was characterized as a primary infection, and any amount over 1 was a secondary infection [28].
A PRNT allows an individual to determine exactly the source of infection in IgM positive individuals by detecting precise neutralizing antibodies. When the virus is neutralized, plaques cannot form, so in theory, if the antibody inactivates the virus, then plaques do not form [29]. To start the test, cells are coated with semi-solid media on a test tube or plate that does not allow the spread of the progeny virus. Then the plaques are compared to determine the amount decrease entire virus infectivity to the concentration of the virus [30].
The PRNT can be done at the Centers for Disease Control and Prevention (CDC, Atlanta, GA, USA) or a lab nominated by CDC. However, worldwide, there is no set standard for PRNT assay [30]. Without a set standard, comparing the results with PRNT can be cumbersome and can alter the interpretations of results [31]. PRNT is also used to effectively determine asymptomatic DENV infections, although it showed cross-reactivity with other DENV serotypes [32].
The antigen is produced and discharged into the bloodstream of the infected patients; as a result, it is considered as an essential biomarker for detecting flavivirus infection at an earlier stage [33][34][35]. According to CDC, the NS1 based tests show similar outcomes as molecular tests in the first week of infection [31]. There is currently one FDA approved NS1 test in the USA, the DENV The seven tests are Dengue NS1 Ag STRIP (BioRad, Marnes-la-Coquette, France), Platelia Dengue NS1 Ag ELISA (BioRad), Dengue NS1 Detect Rapid Test (InBios International), DENV Detect NS1 ELISA
However, the Detect NS1 ELISA (InBios International), which is the only FDA approved product in the market, has a sensitivity of 95.9%, which makes it the superior product among its peers (89.4% by BioRad and 85.6% by Panbio) [36]. The sensitivity varied by DENV stereotype with DNEV-1 reporting the highest sensitivity. However, different studies show different specificities and sensitivities, more or less, closer to the studies mentioned above [37][21][38][39][40][41][42][43].
Rapid tests can be done in a resource-limited settings where access to adequate health care facilities might not be available. However, all the rapid tests showed a statistically significant decrease in sensitivity compared to the FDA-approved Detect NS1 ELISA by InBios International. The highest sensitivity for the rapid tests was for NS1 Ag STRIP by BioRad, which showed a sensitivity of 81.0 to 84.8 percent for all serotypes and a specificity of 100% [44]. The InBios and SD Bioline Dengue NS1 Ag rapid test presented 76.5% and 72.4% sensitivity, respectively [38].
A positive NS1 confirms a DENV infection, while a negative result does not eliminate infection possibility. If a negative test result occurs, then an IgM based test should be performed [31]. A notable downside of NS1-based tests is that they are not recommended after seven days [45].
Nucleic acid tests (NATs) are molecular tests done in a centralized lab location and require expensive equipment and skilled personnel. NATs diagnose and quantify viral RNA/DNA with higher accuracy and sensitivity, allowing detection in the acute phase. The viral genome can be detected within five days of symptom onset. Several assays have already been developed that may exhibit quantitative detection or provide semi-quantitative or qualitative detection.
In RT-PCR, the viral RNA is initially extracted from different samples, including plasma, blood, urine, and serum. Recently, a multiplex RT-PCR assay was developed to determine serotypes from the blood sample, and it can be used during blood transfusion [46]. To automatize the sample extraction to amplification, an insulated isothermal PCR assay has been developed to detect DENV [47]. They developed POCKIT combo central system which accommodates a cartridge where serum samples are loaded and the cartridge contains all the extraction, amplification reagent and the POCKIT system automatically provides the qualitative result with a lowest detection points of 1 and 10 PFU/mL for DENV-1,3 and DENV-2,4 respectively.
The observed LOD in serum and plasma was reported as 1 × 103PFU/ In the retrospective study, the FDA indicated that it could detect RNA in 98.04% of the samples and 98.5 in negative percent agreement [48]. Furthermore, a study compared the CDC assay and a laboratory-built assay [49]. This study showed that the lab assay had a higher sensitivity of 97.4% compared to 87.1% of the CDC assay.
The Trioplex is only usable for symptomatic patients, not for blood donors. The test can detect Dengue in serum, cerebrospinal fluid, and blood but not in urine. The test showed no cross-reactivity with other flaviviruses [50]. In comparison, ZIKV differed slightly for both positive and negative percent agreement showing 95% and 99.1 % agreement, respectively [51].
Another form of molecular diagnostics is isothermal amplification of genomic DENV RNA, where the amplification process requires only one temperature. In rural areas, for example, access to lab-based PCR tests might not be possible. Unfortunately, to date, there is no CDC approved isothermal product for the detection of DENV. Although, several platforms are already being developed that utilized isothermal amplification method, including reverse transcriptase loop-mediated isothermal application (RT-LAMP), Reverse Transcription Recombinase Polymerase Amplification (RT-RPA) and nucleic acid sequence-based amplification (NASBA).
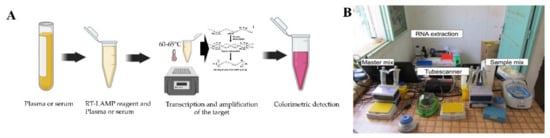
This method has been developed as a promising pathogen detection technique that uses only water bath or a simple heating block to amplify the viral target at a constant temperature, usually in the range of 60–65 °C [52]. The RT-LAMP assay they designed showed a 100% and 98.9% success rate for diagnosing clinical strains of DENV and infected patients, respectively. In their study, the RT-PCR had a sensitivity of 93% for clinical strains and 84.2% for infected patients. However, in the case of multiplexing and viral quantification applications, PCR is superior to LAMP-based assays [53].
Another commonly used isothermal amplification method is NASBA. Silica is used to extract the RNA from the clinical serum or plasma samples, which is then amplified without a thermocycler at 41°C [54]. DENV-1 target RNA is amplified by NASBA amplification followed by generation of sandwich complex by AuNP probe and DENV-1 capture probe. mL. Although NASBA is simple, cost-effective, and it increases the detection sensitivity of the biosensor used but it requires sample pre-processing and cautious handling of the viral targets [55].
A point of need based mobile RPA unit (Figure 3B) has been developed and deployed in Thailand and Senegal to ensure sensitive detection of DENV rapidly [56]. The mobile unit facilitates necessary equipment and reagents for RNA extraction to fluorescent detection. Moreover, another RT-RPA based assay was developed where the detection was done by lateral flow dipsticks with a detection capability of 1 to 106copies Considering all the abovementioned RPA based assays it is clear that this platform requires minimally expensive equipment and can be employed in resource constraint settings for high and accurate detection of DENV.
4. Recently Emerged State-of-the-Art Sensing Technologies for POC DENV Detection
Currently, there is a need for POC technologies that can be cheaper and reliable in various health care settings. This emerging market has many technologies that have the potential to be used in these settings. Technologies such as electrochemical impedance spectroscopy (EIS) based sensing, surface plasmon resonance (SPR), microarray-based sensing, paper-based lateral flow assays, and lab-on-chip (LOC) are currently being researched to become POC technologies so they can be clinically used in these settings. Nanoparticles are often combined with these technologies to enhance performance.
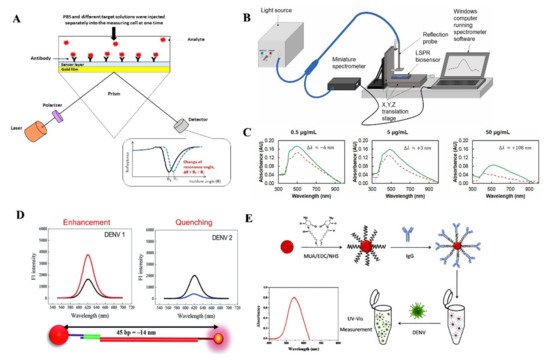
SPR detects changes in the RI when a biomolecular interaction between antigen and antibody occurs [58]. One recently developed SPR based sensing device can detect NS1 antigen of DENV in human plasma samples within 30 min [59]. A silver nanostructure was created using thermal annealing method, which enabled the blood plasma separation using a polyethersulfone membrane in a microfluidic device. The presence of IgM antibodies in the serum was correlated with an increase in resonance angle [60].
Another group of researchers developed a LSPR-based sensing mechanism using serotype-specific nanoprobe bound to CdSeTeS quantum dots and Au nanoparticles to detect different serotypes of DENV [61]. Another easier to operate approach for detecting DENV serotypes using surface modified AuNPs based LSPR sensor was reported recently (Figure 4E) [62]. SPR possesses features to be used in POC settings due to its high sensitivity, real time response and low sample consumption. However, the lower sensitivity of SPR based assays compared to some IgM tests could pose some issues for health care providers.
e.g., impedance, voltage, and current) from the biomolecular interactions on the surface of different biosensors, which has been used for detecting various viral targets [64][65][66][67][68][69]. Till now, different electrochemical based biosensor development approaches have been reported, including carbon nanotube, screen printing, gold nanoparticles, and peptides [70][71][72][73]. This can lead to higher sensitivity, low detection limits, and long-term stabilities for biosensors [74]. Their device has a sensitivity of 45.7 ± 1.7 Ω/order of the magnitude of antibody concentration change in PBS.
Another commonly used analytical sensing technique is voltammetric sensing. On the other hand, DPV measures the before and after application of linear increment of amplitude potential pulses to analyze the redox process [75]. Recently, various voltammetric sensing assays have been reported for detecting DENV; for example, Palomar et al. developed an electrochemical sensor with Au and carbon nanotube based composites [76]. A composite of Au and nitrogen, sulfur co-doped graphene quantum dots has been used to detect DENV DNA using four dye-combined probes with a LOD of 9.4 femto molar [77].
The development of electrochemical sensing based techniques has become a powerful tool for DENV detection in POC application for their diverse and multitude advantages. Besides, the detection sensitivity can be improved by using different types of nanostructures. However, some chemical sensor development requires use of clean room facilities which increases the cost of the sensor. Further, significant sample pre-processing is required before electrochemical methods can be used.
SERS is a method that enhances Raman spectroscopy by improving the electron cloud around metallic nanostructures, which is done through either electromagnetic or chemical enhancement [78]. It was recently showed that SERS active enhancement factors could achieve 106 to 108 M. It provides a chemical fingerprint on an object to ensure higher specificity, requiring a less complex sample preparation mechanism, free from interference of signals capability, high reproductivity and potential to be implemented in POC settings. SERS can be used for various areas such as various viral pathogen diagnostics, including HIV, Influenza virus, Respiratory viruses, SARS-CoV-2 and flaviviruses [79][80][81][82][83][84].
Recently SERS has been adapted to detect infectious diseases such as Zika and DENV [84]. This low LOD shows the promising use of SERS to detect infectious diseases like DENV. The testing for clinical samples using a SERS platform based on Ag nanorod array was carried out in a study [85]. The team of investigators was successfully able to differentiate infected and non-infected patients based on the recorded SERS spectra.
SERS currently holds promise as a POC in the future, but it still inherently lacks robustness and repeatability compared to the current gold standard assays for DENV. A considerable limitation of SERS is the high initial cost of portable Raman readers that might not be available in many rural and developing areas [86]. However, SERS shows promise in detecting DENV at early onset as it provides low LOD. SERS can also lower the cost of testing over the long term due to the fact that it does not require a washing step where reagents are used.
There are several LOC technologies that have been developed to detect various viral targets, as well as devices that incorporate micro/nanoparticles [87][88][89][90]. Often microfluidic-based sensing incorporates paper material and nanotechnology together to enhance detection. To ensure rapid detection, 4G2-coated beads were used and coupled with a fluorescent probe to ensure fluorescence detection. The developed chip is highly reusable and requires a very small sample volume, which reduced the cost of detection per assay.
On the other hand, LODc is another microfluidic device that utilizes centrifugal force to pump the reagents in different chambers inside a disc [91]. It also requires a minimal amount of reagent and is cost-effective due to non-photolithographic fabrication techniques. It also facilitates different types of detection capabilities [92]. Moreover, a multilayer PMMA based LODc has been developed where microspheres and microfluidic disc formed a hybrid that is used to detect DENV.
Zhang et al. developed a lateral flow based assay to detect DENV target antibody (IgM and IgA) from a large volume of saliva samples using a simple stacking flow based 2D platform [93]. This device accommodates the whole LAMP-based NAT process from the dried reagents by completing serum treatment, automated flow control, and finally, the RNA amplification. Moreover, micro size paper-based analytical devices commonly known as μPADs have been incorporated with smartphones to detect the DENV [89][94]. A microfluidic pump was developed to control the flow inside the chambers of the device.
There are some other microfluidic-based technologies that are being used for DENV detection with a potentiality to be used at POC. In this section, we discuss microarray and thread based assays for their application in DENV detection.
Furthermore, the microarray had a better sensitivity with IgG DENV than with IgM, which can be attributed to sampling timing and prior infections. The researchers used the microarray to test DENV after 1 and 6 months of infection. Using their binomial model, they were able to have a 92% accuracy on primary past infections but only a 45% accuracy on past secondary infections [95]. Microarrays have many inherent benefits, such as high throughput and high sensitivity.
Threads can be an alternate mode of liquid transport using capillary wicking principle needing no microchannel fabrication and can be a cheaper alternative to paper-based devices [96]. Kosuke et al. has developed a microfluidic thread based analytical device which is hydrophobic, intertwisted and sewn inside two lamination films [97]. Bioluminescence resonance energy transfer-based technology with the integration of cellphone was utilized to detect multiple antibodies including anti-HIV-1, anti-hemagglutinin or anti-DENV-1 from the spike antibodies in whole blood. The lower limit of detection for anti-DENV-1 was 14.9 nM.
References
- Guzman, M.G.; Halstead, S.B.; Artsob, H.; Buchy, P.; Farrar, J.; Gubler, D.J.; Hunsperger, E.; Kroeger, A.; Margolis, H.S.; Martínez, E. Dengue: A continuing global threat. Nat. Rev. Microbiol. 2010, 8, S7–S16.
- Dick, O.B.; San Martín, J.L.; Montoya, R.H.; del Diego, J.; Zambrano, B.; Dayan, G.H. The history of dengue outbreaks in the Americas. Am. J. Trop. Med. Hyg. 2012, 87, 584–593.
- World Health Organization. Dengue and Severe Dengue; World Health Organization, Regional Office for the Eastern Mediterranean: Cairo, Egypt, 2014.
- Hasan, S.; Jamdar, S.F.; Alalowi, M.; Al Beaiji, S.M.A.A. Dengue virus: A global human threat: Review of literature. J. Int. Soc. Prev. Commun. Dent. 2016, 6, 1.
- CDC. About Dengue: What You Need to Know. 2020. Available online: (accessed on 20 May 2020).
- Pan American Health Organization. Epidemiological Update: Dengue in the context of COVID-19; PAHO/WHO: Washington, DC, USA, 3 December 2020.
- CDC. Symptoms and Treatment. 2020. Available online: (accessed on 27 April 2021).
- Halstead, S. Recent advances in understanding dengue. F1000Research 2019, 8, 1279.
- Simmons, C.P.; Farrar, J.J.; van Vinh Chau, N.; Wills, B. Dengue. N. Engl. J. Med. 2012, 366, 1423–1432.
- Herrada, C.A.; Kabir, M.; Altamirano, R.; Asghar, W. Advances in Diagnostic methods for Zika Virus infection. J. Med. Devices 2018, 12, 12.
- Dimmock, N.J.; Easton, A.J.; Leppard, K.N. Introduction to Modern Virology; John Wiley & Sons: Hoboken, NJ, USA, 2016.
- Paranjape, S.M.; Harris, E. Control of dengue virus translation and replication. Dengue Virus 2009, 338, 15–34.
- Huber, R.G.; Lim, X.N.; Ng, W.C.; Sim, A.Y.; Poh, H.X.; Shen, Y.; Lim, S.Y.; Sundstrom, K.B.; Sun, X.; Aw, J.G. Structure mapping of dengue and Zika viruses reveals functional long-range interactions. Nat. Commun. 2019, 10, 1–13.
- Kuhn, R.J.; Zhang, W.; Rossmann, M.G.; Pletnev, S.V.; Corver, J.; Lenches, E.; Jones, C.T.; Mukhopadhyay, S.; Chipman, P.R.; Strauss, E.G. Structure of dengue virus: Implications for flavivirus organization, maturation, and fusion. Cell 2002, 108, 717–725.
- Dwek, R. Antibodies and antigens: It’s all about the numbers game. Proc. Natl. Acad. Sci. USA 2009, 106, 2087–2088.
- Innis, B.L.; Nisalak, A.; Nimmannitya, S.; Kusalerdchariya, S.; Chongswasdi, V.; Suntayakorn, S.; Puttisri, P.; Hoke, C.H. An enzyme-linked immunosorbent assay to characterize dengue infections where dengue and Japanese encephalitis co-circulate. Am. J. Trop. Med. Hyg. 1989, 40, 418–427.
- Tsai, J.-J.; Liu, C.-K.; Tsai, W.-Y.; Liu, L.-T.; Tyson, J.; Tsai, C.-Y.; Lin, P.-C.; Wang, W.-K. Seroprevalence of dengue virus in two districts of Kaohsiung City after the largest dengue outbreak in Taiwan since World War II. PLoS neglected tropical diseases 2018, 12, e0006879.
- Narayan, R.; Raja, S.; Kumar, S.; Sambasivam, M.; Jagadeesan, R.; Arunagiri, K.; Krishnasamy, K.; Palani, G. A novel indirect ELISA for diagnosis of dengue fever. Indian J. Med. Res. 2016, 144, 128–133.
- Andries, A.C.; Duong, V.; Ngan, C.; Ong, S.; Huy, R.; Sroin, K.K.; Te, V.; Y, B.; Try, P.L.; Buchy, P. Field evaluation and impact on clinical management of a rapid diagnostic kit that detects dengue NS1, IgM and IgG. PLoS Negl. Trop. Dis. 2012, 6, e1993.
- Blacksell, S.D.; Jarman, R.G.; Gibbons, R.V.; Tanganuchitcharnchai, A.; Mammen, M.P., Jr.; Nisalak, A.; Kalayanarooj, S.; Bailey, M.S.; Premaratna, R.; de Silva, H.J.; et al. Comparison of seven commercial antigen and antibody enzyme-linked immunosorbent assays for detection of acute dengue infection. Clin. Vaccine Immunol. 2012, 19, 804–810.
- Lee, H.; Ryu, J.H.; Park, H.S.; Park, K.H.; Bae, H.; Yun, S.; Choi, A.R.; Cho, S.Y.; Park, C.; Lee, D.G.; et al. Comparison of Six Commercial Diagnostic Tests for the Detection of Dengue Virus Non-Structural-1 Antigen and IgM/IgG Antibodies. Ann. Lab. Med. 2019, 39, 566–571.
- Calvert, A.E.; Boroughs, K.L.; Laven, J.; Stovall, J.L.; Luy, B.E.; Kosoy, O.I.; Huang, C.Y. Incorporation of IgG Depletion in a Neutralization Assay Facilitates Differential Diagnosis of Zika and Dengue in Secondary Flavivirus Infection Cases. J. Clin. Microbiol. 2018, 56, e00234-18.
- Prince, H.E.; Yeh, C.; Lapé-Nixon, M. Utility of IgM/IgG ratio and IgG avidity for distinguishing primary and secondary dengue virus infections using sera collected more than 30 days after disease onset. Clin. Vaccine Immunol. 2011, 18, 1951–1956.
- Kuno, G.; Gómez, I.; Gubler, D.J. An ELISA procedure for the diagnosis of dengue infections. J. Virol. Methods 1991, 33, 101–113.
- Nguyen, T.H.T.; Clapham, H.E.; Phung, K.L.; Nguyen, T.K.; Dlnh, T.T.; Nguyen, T.H.Q.; Tran, V.N.; Whitehead, S.; Simmons, C.; Wolbers, M.; et al. Methods to discriminate primary from secondary dengue during acute symptomatic infection. BMC Infect. Dis. 2018, 18, 375.
- Atchareeya, A.N.; Prakong, S.; Sa-Ngasang, A.; Chanama, S.; Sawanpanyalert, P.; Kurane, I.; Anantapreecha, S. Comparison between Haemagglutination Inhibition (HI) Test and IgM and IgG-capture ELISA in Determination of Primary and Secondary Dengue Virus Infections. Dengue Bull. 2006, 30, 141–145.
- Dengue, W. Guidelines for Diagnosis, Treatment. Prevention and Control; WHO: Geneva, Switzerland, 2009; Volume 1.
- Lukman, N.; Salim, G.; Kosasih, H.; Susanto, N.H.; Parwati, I.; Fitri, S.; Alisjahbana, B.; Widjaja, S.; Williams, M. Comparison of the Hemagglutination Inhibition Test and IgG ELISA in Categorizing Primary and Secondary Dengue Infections Based on the Plaque Reduction Neutralization Test. BioMed Res. Int. 2016, 2016, 5253842.
- Russell, P.K.; Nisalak, A.; Sukhavachana, P.; Vivona, S. A plaque reduction test for dengue virus neutralizing antibodies. J. Immunol. 1967, 99, 285–290.
- World Health Organization. Guidelines for Plaque Reduction Neutralization Testing of Human Antibodies to Dengue Viruses; World Health Organization: Geneva, Switzerland, 2007.
- Thomas, S.J.; Nisalak, A.; Anderson, K.B.; Libraty, D.H.; Kalayanarooj, S.; Vaughn, D.W.; Putnak, R.; Gibbons, R.V.; Jarman, R.; Endy, T.P. Dengue plaque reduction neutralization test (PRNT) in primary and secondary dengue virus infections: How alterations in assay conditions impact performance. Am. J. Trop. Med. Hyg. 2009, 81, 825–833.
- Chatchen, S.; Sabchareon, A.; Sirivichayakul, C. Serodiagnosis of asymptomatic dengue infection. Asian Pac. J. Trop. Med. 2017, 10, 11–14.
- Chan, H.B.Y.; How, C.H.; Ng, C.W.M. Definitive tests for dengue fever: When and which should I use? Singap. Med. J. 2017, 58, 632–635.
- Coleman, B.; Coarsey, C.; Kabir, M.A.; Asghar, W. Point-of-care colorimetric analysis through smartphone video. Sens. Actuators B Chem. 2019, 282, 225–231.
- Kabir, M.A.; Zilouchian, H.; Sher, M.; Asghar, W. Development of a flow-free automated colorimetric detection assay integrated with smartphone for Zika NS1. Diagnostics 2020, 10, 42.
- Anderson, N.W.; Jespersen, D.J.; Rollins, L.; Seaton, B.; Prince, H.E.; Theel, E.S. Detection of the dengue virus NS1 antigen using an enzyme immunoassay. Diagn. Microbiol. Infect. Dis. 2014, 79, 194–197.
- Raafat, N.; Blacksell, S.D.; Maude, R.J. A review of dengue diagnostics and implications for surveillance and control. Trans. R. Soc. Trop. Med. Hyg. 2019, 113, 653–660.
- Pal, S.; Dauner, A.L.; Mitra, I.; Forshey, B.M.; Garcia, P.; Morrison, A.C.; Halsey, E.S.; Kochel, T.J.; Wu, S.-J.L. Evaluation of dengue NS1 antigen rapid tests and ELISA kits using clinical samples. PLoS ONE 2014, 9, e113411.
- Blacksell, S.D. Commercial dengue rapid diagnostic tests for point-of-care application: Recent evaluations and future needs? J. Biomed. Biotechnol. 2012, 2012, 1–12.
- Vivek, R.; Ahamed, S.F.; Kotabagi, S.; Chandele, A.; Khanna, I.; Khanna, N.; Nayak, K.; Dias, M.; Kaja, M.K.; Shet, A. Evaluation of a pan-serotype point-of-care rapid diagnostic assay for accurate detection of acute dengue infection. Diagn. Microbiol. Infect. Dis. 2017, 87, 229–234.
- Kerkhof, K.; Falconi-Agapito, F.; Van Esbroeck, M.; Talledo, M.; Ariën, K.K. Reliable Serological Diagnostic Tests for Arboviruses: Feasible or Utopia? Trends Microbiol. 2020, 28, 276–292.
- Suputtamongkol, Y.; Avirutnan, P.; Wongsawat, E.; Kim, Y.-W. Performance of Two Commercial Dengue NS1 Rapid Tests for the Diagnosis of Adult Patients with Dengue Infection. Siriraj Medical Journal 2020, 72, 74–78.
- Parkash, O.; Shueb, R.H. Diagnosis of dengue infection using conventional and biosensor based techniques. Viruses 2015, 7, 5410–5427.
- Dussart, P.; Petit, L.; Labeau, B.; Bremand, L.; Leduc, A.; Moua, D.; Matheus, S.; Baril, L. Evaluation of two new commercial tests for the diagnosis of acute dengue virus infection using NS1 antigen detection in human serum. PLoS Negl. Trop. Dis. 2008, 2, e280.
- Martínez-Cuellar, C.; Lovera, D.; Galeano, F.; Gatti, L.; Arbo, A. Non-structural protein 1 (NS1) of dengue virus detection correlates with severity in primary but not in secondary dengue infection. J. Clin. Virol. 2020, 124, 104259.
- Tezuka, K.; Kuramitsu, M.; Okuma, K.; Nojima, K.; Araki, K.; Shinohara, N.; Matsumoto, C.; Satake, M.; Takasaki, T.; Saijo, M.; et al. Development of a novel dengue virus serotype-specific multiplex real-time reverse transcription-polymerase chain reaction assay for blood screening. Transfusion 2016, 56, 3094–3100.
- Tsai, J.-J.; Liu, W.-L.; Lin, P.-C.; Huang, B.-Y.; Tsai, C.-Y.; Lee, P.-Y.A.; Tsai, Y.-L.; Chou, P.-H.; Chung, S.; Liu, L.-T.; et al. A fully automated sample-to-answer PCR system for easy and sensitive detection of dengue virus in human serum and mosquitos. PLoS ONE 2019, 14, e0218139.
- WHO; WE.C.o.B. Standardization, Collaborative Study to Establish a World Health Organization International Reference Panel for Dengue Virus Types 1 to 4 RNA for Nucleic Acid Amplification Technology (NAT)-Based Assays; World Health Organization: Geneva, Switzerland, 2016.
- Santiago, G.A.; Vázquez, J.; Courtney, S.; Matías, K.Y.; Andersen, L.E.; Colón, C.; Butler, A.E.; Roulo, R.; Bowzard, J.; Villanueva, J.M.; et al. Performance of the Trioplex real-time RT-PCR assay for detection of Zika, dengue, and chikungunya viruses. Nat. Commun. 2018, 9, 1391.
- Colombo, T.E.; Versiani, A.F.; Dutra, K.R.; Rubiato, J.G.D.; Galvão, T.M.; Reis, A.F.N.; Nogueira, M.L. Performance of CDC Trioplex qPCR during a dengue outbreak in Brazil. J. Clin. Virol. 2019, 121, 104208.
- Antiago, G.A.; Vergne, E.; Quiles, Y.; Cosme, J.; Vazquez, J.; Medina, J.F.; Medina, F.; Colón, C.; Margolis, H.; Muñoz-Jordán, J.L. Analytical and clinical performance of the CDC real time RT-PCR assay for detection and typing of dengue virus. PLoS Negl. Trop. Dis. 2013, 7, e2311.
- Teoh, B.T.; Sam, S.S.; Tan, K.K.; Johari, J.; Danlami, M.B.; Hooi, P.S.; Md-Esa, R.; AbuBakar, S. Detection of dengue viruses using reverse transcription-loop-mediated isothermal amplification. BMC Infect. Dis. 2013, 13, 387.
- Wong, Y.P.; Othman, S.; Lau, Y.L.; Radu, S.; Chee, H.Y. Loop-mediated isothermal amplification (LAMP): A versatile technique for detection of micro-organisms. J. Appl. Microbiol. 2018, 124, 626–643.
- Sekaran, S.D.; Soe, H.J. Issues in contemporary and potential future molecular diagnostics for dengue. Expert Rev. Mol. Diagn. 2017, 17, 217–223.
- Darwish, N.T.; Sekaran, S.D.; Alias, Y.; Khor, S.M. Immunofluorescence-based biosensor for the determination of dengue virus NS1 in clinical samples. J. Pharm. Biomed. Anal. 2018, 149, 591–602.
- Abd El Wahed, A.; Patel, P.; Faye, O.; Thaloengsok, S.; Heidenreich, D.; Matangkasombut, P.; Manopwisedjaroen, K.; Sakuntabhai, A.; Sall, A.A.; Hufert, F.T.; et al. Recombinase Polymerase Amplification Assay for Rapid Diagnostics of Dengue Infection. PLoS ONE 2015, 10, e0129682.
- Kim, J.G.; Baek, S.H.; Kim, S.; Kim, H.I.; Lee, S.W.; Phan, L.M.T.; Kailasa, S.K.; Park, T.J. Rapid discriminative detection of dengue viruses via loop mediated isothermal amplification. Talanta 2018, 190, 391–396.
- Omar, N.A.S.; Fen, Y.W.; Abdullah, J.; Chik, C.E.N.C.E.; Mahdi, M.A. Development of an optical sensor based on surface plasmon resonance phenomenon for diagnosis of dengue virus E-protein. Sens. Biosens. Res. 2018, 20, 16–21.
- Austin Suthanthiraraj, P.P.; Sen, A.K. Localized surface plasmon resonance (LSPR) biosensor based on thermally annealed silver nanostructures with on-chip blood-plasma separation for the detection of dengue non-structural protein NS1 antigen. Biosens. Bioelectron. 2019, 132, 38–46.
- Singh, P. SPR biosensors: Historical perspectives and current challenges. Sensors and actuators B: Chemical 2016, 229, 110–130.
- Chowdhury, A.D.; Takemura, K.; Khorish, I.M.; Nasrin, F.; Ngwe Tun, M.M.; Morita, K.; Park, E.Y. The detection and identification of dengue virus serotypes with quantum dot and AuNP regulated localized surface plasmon resonance. Nanoscale Adv. 2020, 2, 699–709.
- Basso, C.R.; Tozato, C.C.; Crulhas, B.P.; Castro, G.R.; Junior, J.P.A.; Pedrosa, V.A. An easy way to detect dengue virus using nanoparticle-antibody conjugates. Virology 2018, 513, 85–90.
- Omar, N.A.S.; Fen, Y.W.; Abdullah, J.; Sadrolhosseini, A.R.; Mustapha Kamil, Y.; Fauzi, N.M.; Hashim, H.S.; Mahdi, M.A. Quantitative and Selective Surface Plasmon Resonance Response Based on a Reduced Graphene Oxide-Polyamidoamine Nanocomposite for Detection of Dengue Virus E-Proteins. Nanomaterials 2020, 10, 569.
- Kabir, M.A.; Zilouchian, H.; Caputi, M.; Asghar, W. Advances in HIV diagnosis and monitoring. Crit. Rev. Biotechnol. 2020, 40, 623–638.
- Shafiee, H.; Asghar, W.; Inci, F.; Yuksekkaya, M.; Jahangir, M.; Zhang, M.H.; Durmus, N.G.; Gurkan, U.A.; Kuritzkes, D.R.; Demirci, U. Paper and Flexible Substrates as Materials for Biosensing Platforms to Detect Multiple Biotargets. Sci. Rep. 2015, 5, 8719.
- Sher, M.; Asghar, W. Development of a multiplex fully automated assay for rapid quantification of CD4(+) T cells from whole blood. Biosens. Bioelectron. 2019, 142, 111490.
- Chowdhury, A.D.; Takemura, K.; Li, T.-C.; Suzuki, T.; Park, E.Y. Electrical pulse-induced electrochemical biosensor for hepatitis E virus detection. Nat. Commun. 2019, 10, 3737.
- Tancharoen, C.; Sukjee, W.; Thepparit, C.; Jaimipuk, T.; Auewarakul, P.; Thitithanyanont, A.; Sangma, C. Electrochemical Biosensor Based on Surface Imprinting for Zika Virus Detection in Serum. ACS Sens. 2019, 4, 69–75.
- Ilkhani, H.; Farhad, S. A novel electrochemical DNA biosensor for Ebola virus detection. Anal. Biochem. 2018, 557, 151–155.
- Arshad, R.; Rhouati, A.; Hayat, A.; Nawaz, M.H.; Yameen, M.A.; Mujahid, A.; Latif, U. MIP-Based Impedimetric Sensor for Detecting Dengue Fever Biomarker. Appl. Biochem. Biotechnol. 2020, 191, 1384–1394.
- Lim, J.M.; Kim, J.H.; Ryu, M.Y.; Cho, C.H.; Park, T.J.; Park, J.P. An electrochemical peptide sensor for detection of dengue fever biomarker NS1. Anal. Chim. Acta 2018, 1026, 109–116.
- Oliveira, M.D.; Correia, M.T.; Diniz, F.B. A novel approach to classify serum glycoproteins from patients infected by dengue using electrochemical impedance spectroscopy analysis. Synth. Met. 2009, 159, 2162–2164.
- Pimenta, T.C.; Santos, C.D.C.; Thomasini, R.L.; Ferreira, L.F. Impedimetric immunosensor for dengue diagnosis using graphite screen-printed electrodes coated with poly(4-aminophenylacetic acid). Biomed. Microdevices 2018, 20, 78.
- Lu, C.H.; Yang, H.H.; Zhu, C.L.; Chen, X.; Chen, G.N. A graphene platform for sensing biomolecules. Angew. Chem. 2009, 121, 4879–4881.
- De Oliveira, O., Jr.; Ferreira, L.; Marystela, G.; de Lima Leite, F.; Da Róz, A.L. Nanoscience and its Applications; William Andrew: Norwich, NY, USA, 2016.
- Palomar, Q.; Xu, X.; Gondran, C.; Holzinger, M.; Cosnier, S.; Zhang, Z. Voltammetric sensing of recombinant viral dengue virus 2 NS1 based on Au nanoparticle–decorated multiwalled carbon nanotube composites. Microchim. Acta 2020, 187, 1–10.
- Dutta Chowdhury, A.; Ganganboina, A.B.; Nasrin, F.; Takemura, K.; Doong, R.-a.; Utomo, D.I.S.; Lee, J.; Khoris, I.M.; Park, E.Y. Femtomolar detection of dengue virus DNA with serotype identification ability. Anal. Chem. 2018, 90, 12464–12474.
- Marks, H.; Schechinger, M.; Garza, J.; Locke, A.; Coté, G. Surface enhanced Raman spectroscopy (SERS) for in vitro diagnostic testing at the point of care. Nanophotonics 2017, 6, 681–701.
- Kamińska, A.; Witkowska, E.; Winkler, K.; Dzięcielewski, I.; Weyher, J.L.; Waluk, J. Detection of Hepatitis B virus antigen from human blood: SERS immunoassay in a microfluidic system. Biosens. Bioelectron. 2015, 66, 461–467.
- Fu, X.; Cheng, Z.; Yu, J.; Choo, P.; Chen, L.; Choo, J. A SERS-based lateral flow assay biosensor for highly sensitive detection of HIV-1 DNA. Biosens. Bioelectron. 2016, 78, 530–537.
- Kukushkin, V.I.; Ivanov, N.M.; Novoseltseva, A.A.; Gambaryan, A.S.; Yaminsky, I.V.; Kopylov, A.M.; Zavyalova, E.G. Highly sensitive detection of influenza virus with SERS aptasensor. PLoS ONE 2019, 14, e0216247.
- Wang, C.; Wang, C.; Wang, X.; Wang, K.; Zhu, Y.; Rong, Z.; Wang, W.; Xiao, R.; Wang, S. Magnetic SERS strip for sensitive and simultaneous detection of respiratory viruses. ACS Appl. Mater. Interfaces 2019, 11, 19495–19505.
- Chen, H.; Park, S.-G.; Choi, N.; Kwon, H.-J.; Kang, T.; Lee, M.-K.; Choo, J. Sensitive Detection of SARS-CoV-2 Using a SERS-Based Aptasensor. ACS Sens. 2021.
- Sánchez-Purrà, M.; Carré-Camps, M.; de Puig, H.; Bosch, I.; Gehrke, L.; Hamad-Schifferli, K. Surface-enhanced Raman spectroscopy-based sandwich immunoassays for multiplexed detection of Zika and Dengue viral biomarkers. ACS Infect. Dis. 2017, 3, 767–776.
- Gahlaut, S.K.; Savargaonkar, D.; Sharan, C.; Yadav, S.; Mishra, P.; Singh, J.P. SERS Platform for Dengue Diagnosis from Clinical Samples Employing a Hand Held Raman Spectrometer. Anal. Chem. 2020, 92, 2527–2534.
- Langer, J.; Jimenez de Aberasturi, D.; Aizpurua, J.; Alvarez-Puebla, R.A.; Auguié, B.; Baumberg, J.J.; Bazan, G.C.; Bell, S.E.; Boisen, A.; Brolo, A.G. Present and future of surface-enhanced Raman scattering. ACS Nano 2019, 14, 28–117.
- Omar, N.A.S.; Fen, Y.W.; Abdullah, J.; Mustapha Kamil, Y.; Daniyal, W.M.E.M.M.; Sadrolhosseini, A.R.; Mahdi, M.A. Sensitive Detection of Dengue Virus Type 2 E-Proteins Signals Using Self-Assembled Monolayers/Reduced Graphene Oxide-PAMAM Dendrimer Thin Film-SPR Optical Sensor. Sci. Rep. 2020, 10, 2374.
- Coarsey, C.; Coleman, B.; Kabir, M.A.; Sher, M.; Asghar, W. Development of a flow-free magnetic actuation platform for an automated microfluidic ELISA. RSC Adv. 2019, 9, 8159–8168.
- Sher, M.; Zhuang, R.; Demirci, U.; Asghar, W. based analytical devices for clinical diagnosis: Recent advances in the fabrication techniques and sensing mechanisms. Expert Rev. Mol. Diagn. 2017, 17, 351–366.
- Ilyas, S.; Sher, M.; Du, E.; Asghar, W. Smartphone-based sickle cell disease detection and monitoring for point-of-care settings. Biosens. Bioelectron. 2020, 165, 112417.
- Park, J.; Sunkara, V.; Kim, T.-H.; Hwang, H.; Cho, Y.-K. Lab-on-a-disc for fully integrated multiplex immunoassays. Anal. Chem. 2012, 84, 2133–2140.
- Henderson, B.; Kinahan, D.J.; Ducrée, J. The Centrifugal Microfluidic: Lab-on-a-Disc Platform. In Microfluidics for Biologists; Springer: Berlin/Heidelberg, Germany, 2016; pp. 115–144.
- Zhang, Y.; Bai, J.; Ying, J.Y. A stacking flow immunoassay for the detection of dengue-specific immunoglobulins in salivary fluid. Lab Chip 2015, 15, 1465–1471.
- Priye, A.; Bird, S.W.; Light, Y.K.; Ball, C.S.; Negrete, O.A.; Meagher, R.J. A smartphone-based diagnostic platform for rapid detection of Zika, chikungunya, and dengue viruses. Sci. Rep. 2017, 7, 1–11.
- Thao, T.T.N.; de Bruin, E.; Phuong, H.T.; Thao Vy, N.H.; van den Ham, H.-J.; Wills, B.A.; Tien, N.T.H.; Le Duyen, H.T.; Whitehead, S.S.; Boni, M.F. Using NS1 flavivirus protein microarray to infer past infecting dengue virus serotype and number of past dengue virus infections in Vietnamese individuals. J. Infect. Dis. 2020.
- Li, X.; Tian, J.; Shen, W. Thread as a versatile material for low-cost microfluidic diagnostics. ACS Appl. Mater. Interfaces 2010, 2, 1–6.
- Tomimuro, K.; Tenda, K.; Ni, Y.; Hiruta, Y.; Merkx, M.; Citterio, D. Thread-Based bioluminescent sensor for detecting multiple antibodies in a single drop of whole blood. ACS Sens. 2020, 5, 1786–1794.




