1000/1000
Hot
Most Recent

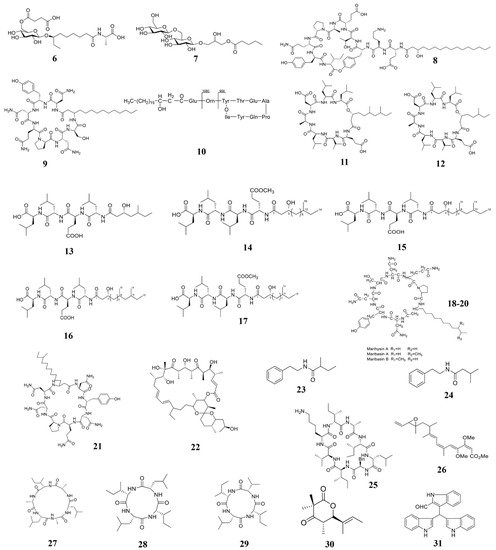
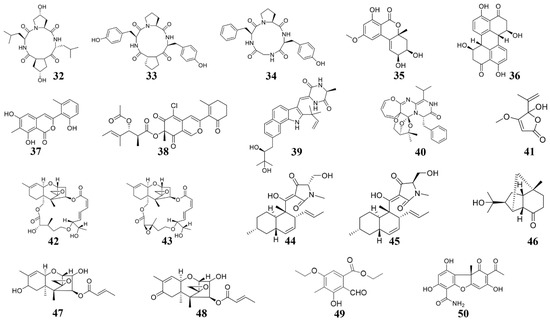
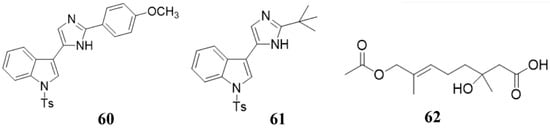
Marine natural compounds suppress or kill plant pathogenic pathogens through different mechanisms, including affecting microbial cell wall synthesis, cell membrane permeability, fatty acid metabolism, respiratory system, cytoskeleton, bacterial quorum sensing (QS), as well as inducing plant immune system for inhibition.