1. Introduction
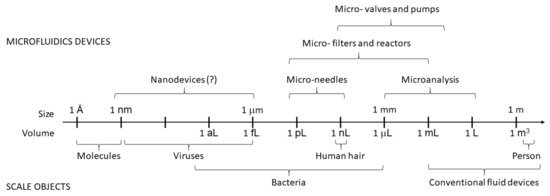
Microfluidics (MF) is a relatively new branch of science and microengineering that deals with manipulating fluids in microchannels with at least one dimension of 1 to 100 micrometers [1][2], as shown in . Scientists consider it a new discipline not only because of the recent emergence of microfluidic (MF) devices that can implement rapid solutions to complex analytical problems at the microscale but also because the physical principles of fluid flow at such small length scales differ from those in macrosystems [3]. MF spans several disciplines—physical and chemical sciences, micromechanics, electronics, and mechanical engineering. It has wide applicability in many fields, with particular emphasis on biology, biochemistry, biotechnology, medicine, pharmacology, and food and environmental analysis. It has made great progress in the last 15 to 20 years [4]. Intensively developing research areas in MF are lab-on-chip (LOC) devices [5][6] and microanalytical systems (μTAS) [7]. They can be considered synonymous with integrated circuits in electronics. The MF technology is based on micropumps, mixers, filters and valves to realize chemical and biological laboratory processes on a single chip. MF requires tiny samples and reagents for analysis, which makes it environmentally friendly and minimally invasive [8].
Figure 1. Scales and volumes in microfluidics.
MF is perceived as a new platform for highly efficient separation and highly sensitive analysis of (bio)molecules and (bio)particles in biochemistry, molecular biology and biotechnology [8] and relies on scale reduction to reduce material consumption and cost [1]. MF exploits the potential of flowing liquids at the microscale to generate “quantitative assays”. In peptidomics and proteomics, MF is most commonly used in combination with MS analyses. Hydrophobic membranes were originally used to adsorb native peptides or peptides generated by enzymatic digestion of proteins, followed by their desalting and elution in a controlled environment for MS analysis [1][2]. Peptides and proteins are analyzed at MS mainly by two ionization techniques, electrospray ionization (ESI) and matrix-assisted laser desorption ionization (MALDI). ESI–MS and MALDI–MS are used not only to determine the relative molecular masses of peptides and proteins but also to elucidate their amino acid sequences and post-translational modifications [9][10][11][12]). Rapid and highly sensitive analysis was achieved by using ultralow sample volumes and amounts (picomoles to femtomoles of compounds in nanoliters to picoliters of volume per single analysis), high separation efficiency, and short analysis times. The widespread use of MF can be attributed to the inherent advantages of MF instruments: Wide applicability, compactness and the need for extremely low sample volumes of the analyzed compounds or particles, as well as low reagent consumption [2][12][13]. By tailoring the geometry of pores in porous materials, fluid flow can be controlled in microfluidic devices. [14]
2. Applications of Microfluidics
3.1. Analysis of Peptides and Proteins; Peptidomics and Proteomics
Peptides and proteins are extremely important biological molecules. As hormones, hormone or drug receptors, enzymes, coenzymes, enzyme substrates or inhibitors, antigens, antibodies, immunomodulators, antibiotics, structural elements and transport molecules play a crucial role in all living organisms. They ensure the basic operations of the cellular machinery. In addition, there are many peptide and protein-based drugs and prodrugs, and some peptides and proteins are used as biomarkers [15]. Moreover, a complete analysis of all peptides (peptidomes) and proteins (proteomes) of a cell, tissue, biofluid, organ or organism is important to understand normal and pathological processes. This is the subject of peptidomics and proteomics—comprehensive and large-scale studies of complex mixtures of peptides and proteins. In this context, the relevance of peptides is increasing because the structure and function of proteins are often identified by their enzymatically generated peptide fragments. This bottom-up or shot-gun approach is one of the main directions in current proteomics research.
The separation and study of peptides and proteins in complex biological matrices is a challenging process that requires advanced and accurate methods that can provide relevant information about their structural and functional properties.
It has been known for more than a century that proteins can move or even “fly” under the influence of an electric field [16]. Therefore, electromigration methods represent powerful tools for their separation and analysis. Polyacrylamide gel electrophoresis in the presence of sodium dodecyl sulfate (SDS–PAGE), isoelectric gel focusing (IEF), and two-dimensional gel electrophoresis (2-DE), which combines the orthogonal principles of narrow tube gel IEF in the first dimension and plate gel SDS–PAGE in the second dimension, have been widely used in the past for protein and polypeptide analysis [17][18]. Currently, HPLC and UHPLC combined with high-resolution MS detection are the leading techniques for peptide and protein analysis [19]. Capillary and microchip electromigration methods (CE/MCE) are also very powerful and useful methods for the analysis and characterization of peptides and proteins [20][21]) and for applications in peptidomics and proteomics [22]. CE and MCE methods have several advantages, such as high separation efficiency, short analysis time, low sample and reagent consumption, and different separation modes (ZE, ITP, IEF, AE, EKC, and EC). In the field of peptidomics and proteomics, CE and MCE are usually combined with MS detection, as MS can identify and quantify the separated peptides and proteins. In addition to MS detection, UV absorption or LIF detection are also commonly used in CE and MCE analyses of peptides and proteins [21][23].
Biological samples (e.g., body fluids, tissues, and food extracts) contain complex mixtures of variable compounds with low and high molecular mass. Therefore, the target peptides and proteins usually need to be extracted or pre-isolated and/or preconcentrated from the sample matrix before analysis [24][25]. The method of sample preconcentration is based on (i) the principle of electrophoresis [26][27][28] (on-site field-assisted sample stacking, micelle stacking, ITP and IEF) or (ii) selective adsorption/extraction method (elution) [29][30]. Usually, a combination of different types of preconcentrates is used. Another important issue in CE and MCE analyses of peptides and proteins is the prevention of protein adsorption on the inner wall of the capillary.
CE/MCE technology was introduced to improve the efficiency of biomarker protein analysis. This technology can find wide applications in hospitals and other immediate medical facilities. This has been proved by several studies of protein processing related to biomedical research and application. Štěpánová and Kašička [21] and Dawod et al. [6][31] described the developments in protein analysis using various CE and MCE methods in 2011–2017. They showed that sample preparation, preconcentration, inhibition of adsorption and control of EOF, separation by a specific CE/MCE method and improvement of the detection scheme have greatly improved the ability of CE/MCE methods for protein analysis. The innovative application of CE and MCE methods in biopharmaceutical protein quality control, protein determination in complex biometrics, peptide-protein mapping, and determination of physical and chemical parameters of proteins are important achievements in this field.
However, a faster and more sensitive analysis than existing analytical methods is needed.
The application of MF technology in this field is an intensively developed concept to create integrated and fully automated analytical devices that can detect and quantify one or more peptides and proteins from a complex matrix. In this miniaturized MF form of CE, all operations (including sample preparation, derivatization, injection, separation, and detection) are integrated into a micrototal analytical system (uTAS) or lab on chip (LOC) platform [32].
Sonker et al. [33] reported the possibility of using a specific separation system for integrated immunoaffinity extraction to study human serum matrix biomarkers in preterm infants. They used a reactive polymer to immobilize the antibody as a whole and selectively extracted targeted preterm infant markers. For effective separation, they also optimized the MF immunoaffinity extraction protocol and combined it with MCE. The low nanomolar concentration of the two enriched preterm markers in the human serum matrix was studied for 30 min. Their observations may help to develop automated and integrated birth risk assessment tools.
Peptide identification by MS implicitly provides information about post-translational modification (PTM) sites. It can be used to identify the threshold of a particular enzyme under physiologically appropriate conditions. Noach-Hirsh et al. [34] presented a modular integrated MF platform to analyze multiple post-translational modifications of newly synthesized protein arrays. This method can also be used to clarify PTM fingerprints on single cells or tissues. Although the technology is comprehensive, it is limited in size and requires relatively small amounts of biological materials and reagents for research. It is suitable for basic and translational research [34].
More attention has been paid to tools used to identify target proteins or disease biomarkers. The most difficult challenge remains the ability to detect low protein abundances in a single cell. Recent advances in high-resolution/high-quality high-precision mass spectrometers have enabled identifying more than 5000 proteins from less than 100 ng of protein extracts in a short analysis time of 15 min LC–MS [35].
The crude protein content of mammalian cells is only about two orders of magnitude lower than the value that gives a detailed cell extract profile. There is also a need to improve the detection limit of MS, scan rate and intelligent data acquisition technology so that MF proteomics and single-cell or relatively small protein proteomics provide comparable results. Therefore, some researchers believe that MS detection strategies using existing data are most likely to achieve the required performance [35].
There are many important advances in the MF platform that can quantify unicellular proteins. Absolute quantification is the key criterion for the determination of unicellular proteins. Without absolute quantification, it is impossible to accurately compare protein amounts determined by different methods. Research to improve the precision of single-cell protein quantification also has great potential. Fluorescent NPs with higher fluorescence intensity can be used to replace conventional fluorescent probes. Nucleic acid-labeled antibodies can be further amplified and quantified in automated PCR [36].
3.2. Separation and Analysis of Cells
3.2.1. Cell Sorting and Single-Cell Analysis
Sorting cells based on their size, type, or density is important for their study and analysis, especially for distinguishing diseased from normal cells in the diagnosis and treatment of various diseases
[37]. Cell separation methods are evaluated based on their separation efficiency, enrichment, and throughput
[38]. Separation efficiency refers to the homogeneity of the separated cell (sub)populations, enrichment refers to the increase in cell concentrations compared to the initial concentration, and sample throughput refers to the speed of cell separation. Fluorescent markers and labeling techniques have been an important aspect of conventional approaches. This problem was addressed by relying on the physical properties of the particles that acted as targets for separation.
One of the most important features of MF devices is their ability to separate and sort cells for automated processing in the diagnosis and treatment of disease
[39][40]. Passive MF devices are preferable to active devices for several reasons. Active devices use variable forces, including magnetic, electrical, and optical forces, for cell movement and operate with higher efficiency
[39]. However, they are more expensive and can damage the cells. On the other hand, passive systems work with inertial forces, sediment, gravity, and filters for cell enrichment. The design factors of passive devices include the elements of viscosity, capillary forces, and surface forces. Since they do not require extensive equipment, improvements in their efficiency can lead to higher reliability
[41][42]. In this context, flow cytometry is an important strategy for sorting cells based on their characteristics, shape and size in heterogeneous cell populations. Separation by membrane filtration and centrifugation has inherent problems related to preparation complexity, time and skill requirements. In contrast, methods from MF offer advantages in terms of high speed of analysis and low cost. They are also less invasive. Among the three different groups of separation techniques (active, passive and combined), the passive method is the simplest. Cell separation in MF requires some kind of fractionation method or force
[43]. The different types of filters used for this purpose include microscale filters, which sort cells based on their deformability and size, and hydrodynamic filtration techniques, which take into account the size and shape of the cells. Other separation techniques include pinched-flow fractionation, which separates particles based on their size, and the deterministic lateral shift method, which changes the trajectories of cells based on their size
[44]. Gravity and sedimentation fractionation methods are based on the density of particles. Biomimetic devices separate particles based on their actual behavior in the human system. Finally, inertial devices can be used for particle migration
[41][44][45].
For cell separation systems, the separation capacity takes into account the amount of sample available for analysis, the distinguishing characteristics of the cell types, the purity required, and the desired characteristics of the isolated population
[46]. The total number of cells lost during the separation process, the viability of the cells after separation, and the physical pressure on the cells. Finally, the choice of effective sample preparation procedures, the time required for the entire cell separation process, and the cost-effectiveness of the technology are also important
[47]. It is advantageous to integrate the technology for the separation of cells and other particles into the lab-on-chip equipment because these unlabeled processes are continuous, the separation process can be continuously monitored, and the sample components can be moved laterally so that each part can be collected independently
[48]. It is expected that the specific cells in the biopsy sample will be treated individually. Further analysis should focus on system expansion, which obviously requires thorough optimization
[33].
MF design considerations were related to the design and generation of forces and determining how the forces move the particles. A discrete-phase approach was used to determine particle motion and to study the effects of forces, such as buoyancy force, virtual mass force, pressure gradient force, Saffman force, Basset force, body force, and Brownian force
[49]. High throughput methods MF included “controlled incremental filtration”, “continuous particle separation in a spiral microchannel,” and “shear modulated inertial migration”. Data analysis was performed using MATLAB software
[50][51]. In the study, three different passive MF methods were demonstrated for stepwise filtering of fluid from the main channel into the side channels to achieve single-cell isolation. The method that proved most successful had low recovery
[41].
Ahmad et al. proposed a tapering MF device for separating multiple particles (cells and microbeads) in biological samples (removing contaminants from heterogeneous mixtures) in a passive process
[52][53]. Their microfabrication research focused on heterogeneous samples and addressed the problems associated with the isolation of cells and microbeads. The present research involved developing a microfluidic device capable of using passive separation technology for microparticles and cells. The device consists of a tapered microchannel with an outlet, an inlet, and tapered angles. Sample solutions are introduced into the inlet and move along the centerline of the narrow microchannel, which serves as the focal region. The cells and microbeads move along different trajectories as they leave the focal area. These trajectories are affected by the density, deformability, and size of the cells. Particles and cells migrate laterally depending on their flow velocity and radius. The widening of the microchannel causes the average velocity of the particles and cells in the center to slow down. This change causes the flow to form a downward angle near the lateral exits. This creates streamlines that help the particles and cells travel to the collection outlets. Inside the microchannel, the particles and cells experience hydrodynamic drag that is introduced into the fluid flow. Particles and cells entering the microchannel cause a change in flow velocity at the outlet. The widening of the microchannel results in varying degrees of hydrodynamic drag at different outlets. The hydrodynamic drag at an outlet causes trapping. The design of the microchannel is optimized using simulation finite element analysis (FEA). The system MF created with hydrodynamic principles is valid for many different applications. Its parameters and operating conditions can be further optimized. Calibration of parameters, such as flow rate and sample concentration, helps to increase the throughput significantly
[53].
The major challenges for MF separation devices are the throughput of the separation process, complexity in design and purity of the sample. The solution proposed in the paper achieves the desired results by using multiple cone angles to separate samples with high purity. The “hydrodynamic separations” are supported by a coupling mechanism and sedimentation, which help to achieve the required sample purity. The process is supported by technology in the form of computer-aided design (CAD)
[54][55] and finite element analysis (FEA)
[56][57]. The device is fabricated using soft lithography.
3.2.2. Secretome Analysis and Single-Cell Omics Analysis
The cell secretome is a complex proteome secreted by cells. It is the basic mechanism for communication between cells in vivo and in vitro. Analysis of proteins secreted in body fluids can determine biomarkers for important pathophysiological conditions
[58]. However, due to the complexity of protein content in body fluids, there is a great need for a better understanding of the proteins secreted by different cell types. This can be more easily explored in vitro (). To this end, MF tissue culture systems may be particularly important as they can accumulate endogenous and exogenous signals at the microliter scale, thus better preserving the self-regulation that occurs in small interstitial spaces in the body
[59] ().
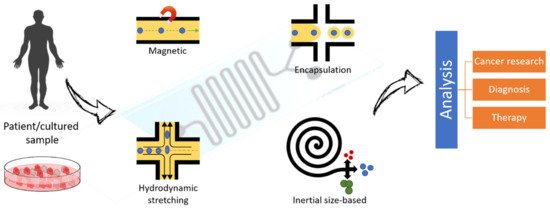
Figure 6. Schematic representation of the most commonly used techniques for the isolation, trapping and manipulation of single cells in a microfluidic device. Potential scope of applications is highlighted.
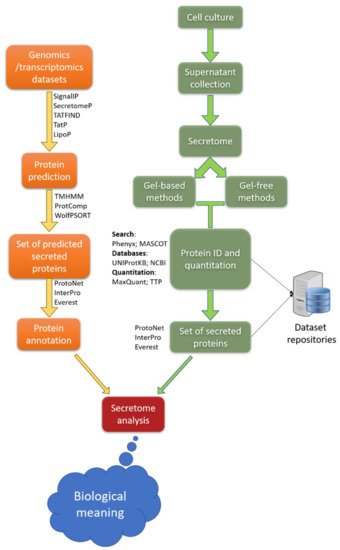
Figure 7. Schematic representation of the workflow of secretome analysis for the detailed characterization proteins secreted by different cells, tissues and organs. The two main approaches described, as well as the most commonly used bioinformatics tools used in each of the shown steps. Adapted from
https://www.sciencedirect.com/science/article/pii/S1570963913000502, 26 November 2013.
Hu et al.
[58] quantitatively analyzed the proteins secreted by human foreskin fibroblasts grown in multi-well plates or in MF systems. This comparison showed the general accumulation of secreted proteins in MF systems. However, not all proteins accumulate equally. This suggests that the culture microenvironment plays a feedback role in cell regulation of protein secretion. Therefore, it is important to study the cell secretion group in a culture system that is more similar to the conditions of the microenvironment in vivo, and the MF volume is small, which better mimics the small interstitial spaces in the tissue
[59].
Heterogeneity studies based on single-cell-omics analyses are important for identifying diverse cell populations, discovering new cell types, revealing informative cell properties, and discovering important relationships between cells. Recently, MF has become a powerful technique for single-cell-omics analyses due to the MF technology’s advantages of flow, sensitivity, and precision
[60]. Here, recent advances in MF single-cell oocyte analysis, including various designs of microfluidic platforms, lysis strategies, and oocyte analysis techniques, have become essential biomolecular tools
[60][61].
Compared to traditional test tube operations, MF offers excellent flow, sensitivity, precision, integration, and partial automation in omics research. However, the new research still faces challenges
[62]. First, in single-cell isolation, not all advantages, such as high throughput and effective cell isolation, simple chip design and fabrication, and integrated multistep operation, can be maintained because each isolation method has its advantages and limitations
[60][61]. Second, in omics analysis, barcoding technologies used for cell/molecule labeling can increase throughput and significantly reduce cost, but they are often biased (e.g., the 3-terminal cDNA ends) and can lead to loss of important signals. To address this challenge, mature third-generation sequencing technology can reverse this trend by increasing read length and single-molecule sequencing
[60][61].
3.2.3. Investigation of Stimulus-Driven Cell Behavior
Lazar and colleagues
[63] attempted to elucidate cell behavior by using MF in combination with MS analysis. Their research aimed to perform proteomic profiling for cell responses to stimuli. Their motivation was the complex nature of cell responses to chemical or physical stimuli: different time scales and rates, presence or absence of protein synthesis; transient or sustained signaling processes; involvement of neuronal, metabolic, and immune systems; involvement of signal transduction and protein phosphorylation; and the effects of cell proliferation, growth, transformation to a disease state, and differentiation
[64][65]. Their approach consisted of important features, such as including a substantial number of cells for plausible MS analysis and achieving rapid sampling of cell contents and uniform cell stimulation. Their protocol was designed to deliver the cell stimulant axially and transversely. Fluorescent dyes were used to identify fluid manipulation on the chips and determine the results of stimulation. Different biological processes were addressed by axial and transverse delivery. Processes that required sustained stimulation of cells were not affected by a small concentration gradient. Those that did not require a concentration gradient were amenable to axial delivery. In contrast, biological processes characterized by rapid responses to stimulation, those requiring steady and rapid delivery of the stimulant, or those requiring rapid lysis of cells were amenable to transverse delivery of the stimulant
[63][66].
3.2.4. Investigation of Biomolecular Coronas Nanoparticles
Nanoparticles (NPs) serve as potential vehicles for drug and nucleic acid research. However, their mechanism of action has not been fully elucidated. The implications of this limited understanding of in vitro data mean that the physiological response to NPs in vivo cannot be predicted. Biomolecular coronas (BCs) formed on NPs by their contact with biofluids are of great importance in molecular medicine
[67][68]. BCs are influenced by various factors, such as size, temperature, surface charge, hydrophobicity, incubation time and protein source. This premise is further explored to associate disease-specific protein changes with BC composition to gain deeper insight into personalized BCs. BCs may be able to distinguish cancer patients from normal patients and understand disease progression. The current research is focus on discovering new mechanisms to generate crowns through applying shear stress
[67][68]. Previously, the process of incubating NPs with human serum (HS) or human plasma (HP) was used to adsorb blood proteins. Recent research shows that the sheer stress of a “laminar fluid flow” compared to static incubation leads to the formation of more complex coronae
[69][70][71]. It is expected that this form of dynamic flow will provide a better understanding of actual physiological processes. MF provides a standardized environment for performing dynamic incubation protocols as the reproducibility is far greater than conventional silicone tubing and peristaltic pumps
[68].
Digiacomo et al.
[68] investigated the role of MF in this field by using gold nanoparticles (GNPs) to explore their plausibility in chemotherapy research and drug delivery in cancer. GNPs can activate multiple juxtaposed receptor sites and increase cellular uptake. They possess surface plasmon resonance and can efficiently convert light to heat, localize the effect of temperature, and induce rapid cell death. The analysis showed that the sheer stress of an MF environment leads to the formation of more negatively charged NPs and affects protein composition. Future meta-analysis will be possible by standardizing the composition and formation of BCs
[67][68]. Changing the size and shape of GNPs may lead to different results
[72]. It is important to note that the method chosen to analyze the particles MS/MS is not able to identify the exact protein structure on the surface of GNPs. It may be possible to understand the role of NPs in the physiological response by studying the exposed protein epitopes. In addition, a detailed understanding of the structure of the corona is essential to determine the nature of activation of processes at the cellular and subcellular levels
[68].