
| Version | Summary | Created by | Modification | Content Size | Created at | Operation |
|---|---|---|---|---|---|---|
| 1 | Quang-Minh Nguyen | + 3761 word(s) | 3761 | 2021-05-25 08:24:23 |
Video Upload Options
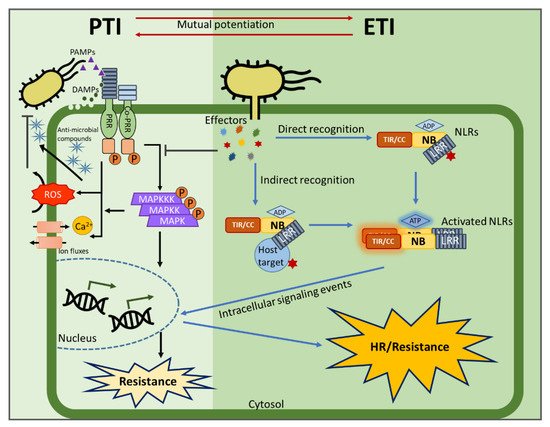
Plants rely on multiple immune systems to protect themselves from pathogens. When pattern-triggered immunity (PTI)—the first layer of the immune response—is no longer effective as a result of pathogenic effectors, effector-triggered immunity (ETI) often provides resistance. In ETI, host plants directly or indirectly perceive pathogen effectors via resistance proteins and launch a more robust and rapid defense response. Resistance proteins are typically found in the form of nucleotide-binding and leucine-rich-repeat-containing receptors (NLRs). Upon effector recognition, an NLR undergoes structural change and associates with other NLRs. The dimerization or oligomerization of NLRs signals to downstream components, activates “helper” NLRs, and culminates in the ETI response. Originally, PTI was thought to contribute little to ETI. However, most recent studies revealed crosstalk and cooperation between ETI and PTI.
1. Introduction
2. The Evolution of Pathogen Perception by NLRs
3. NLR Activation and Signaling Events Following Pathogen Recognition
3.1. Multi-Domain NLRs Act as Molecular Switches
3.2. Homo/Hetero-Complex Formation Is Necessary for NLR Signaling
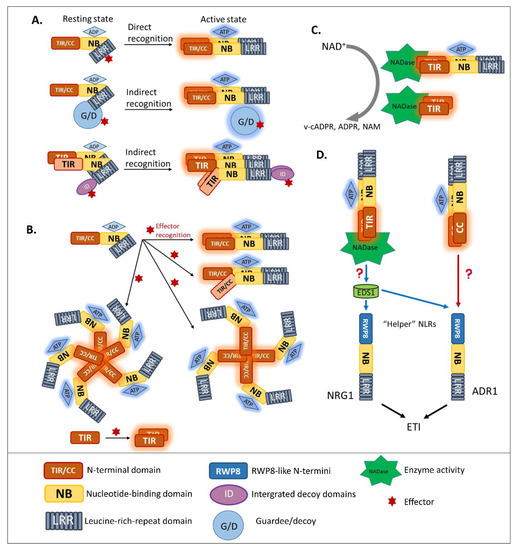
Previous studies reported that disruption of the Mildew A 10 (MLA10) CC dimerization abolished the activation of immunity [42][62][63], suggesting that CNLs require dimerization of the CC domain for signal transduction. Moreover, pentameric oligomerization of the CNL Hrp-dependent outer protein (Hop) Z-Activated Resistance 1, termed the “HopZ-Activated Resistance 1 resistosome”, is important for the formation of putative membrane pores and the immune response [64]. Similarly, several well-studied plant NLRs containing TIR domains, such as RECOGNITION OF PERONOSPORA PARASITICA 1 (RPP1), the flax resistance protein L6, RRS1, and RPS4, were found to require oligomerization by two distinct interfaces, for both self-association and defense signaling [65][66][67][68][69]. Similar to the case of MLA10, disrupting the homo-dimerization of L6 TIRs interferes with downstream signaling (Figure 2B). To effectively recognize the effector Xanthomonas outer protein Q (XopQ), the TNL Recognition of XopQ 1 resistosome requires tetramerization [70]. In addition, two asymmetric TIR homodimers that form an RPP1 tetrameric resistosome activate downstream signaling, in response to effector Arabidopsis thaliana Recognized 1(Figure 2B) [66].
Hetero-associations in addition to homo-dimerization were proven to be an indispensable aspect in NLR-mediated signaling. Indeed, genetically-linked paired NLRs were characterized as functioning together in conferring pathogen resistance [42]. RGA4/RGA5 is one of the functionally paired CNLs for Magnaporthe oryzae AVR-Pia/AVR-Pik-mediated resistance [41]. In addition, genetically-linked, paired TNLs, such as RPP2A/RPP2B, were found to provide resistance against Hpa race Cala 2 [71], along with previously discussed RRS1/RPS4 recognize AvrRps4 and PopP2 [55][72]. In the paired cases listed above, one NLR, the “sensor NLR”, usually contains an evolutionarily incorporated integrated domain, and acts as an effector receptor, while the second NLR, the “executor NLR”, induces downstream signaling [41].
3.3. Intramolecular Regulation of Guardee/Decoy Contributes to NLR-Mediated Resistance
It is now clear that R proteins can guard plant functions by monitoring different post-translational modifications of effector targets (guardee/decoy), and that different modifications can compete with or support each other. RIN4 was proposed to act as a phosphoswitch to detect the effector AvrRpm1. Targeting of RIN4 by AvrRpm1 causes the phosphorylation of threonine 166 within its C-terminal nitrate-induced domain; which leads to RPM1 activation and resistance [73]. A recent study revealed that the ADP-ribosylation of RIN4 at aspartate 153 by AvrRpm1, leads to threonine 166 phosphorylation and promotes RPM1 activation [74]. The addition of ADP-ribose supports the complete phosphorylation of threonine 166 in RIN4 [74]. Taken together, these reports indicate that several additive modifications can occur in a single guardee protein.
On the other hand, a post-translational modification of one effector target can antagonize another. The newest report of RRS1/RPS4-mediated immunity revealed that phosphorylation regulates the activation of paired RRS1/RPS4 [68]. In the absence of effector AvrRps4 or PopP2, phosphorylation at threonine 1214 in the integrated decoy WRKY domain keeps RRS1 from the resistant ecotype Wassilewskija, in a resting state. Dephosphorylation at that residue leads to the autoactivation of RRS1. Interestingly, PopP2 induces O-acetylation in the WRKY domain of RRS1, which competes with its phosphorylation and results in the dephosphorylated activated RRS1-mediated resistance to Ralstonia Solanacearum [68]. Other phosphorylation sites at the C terminus of RRS1 are required for PopP2 recognition, which enhances the interaction of the TIR domain with the WRKY domain. This study also proved that wild-type Columbia RRS1 lacks the C-terminal 83 amino acids that include the target phosphorylation sites, fails to recognize PopP2, and is thus susceptible to Ralstonia Solanacearum.
However, RRS1-mediated resistance to the Pseudomonas syringae effector AvrRps4 is determined by the association of the RRS1 C-terminus with its TIR, not by its phosphorylation status [68]. The C terminus and TIR of RRS1 interact with each other only in the presence of AvrRps4 [68]. During recognition of AvrRps4 or PopP2, the interaction of the RRS1 TIR domain with its C terminus is enhanced. This enhanced interaction releases the RPS4 TIR from the inhibition by the RRS1 TIR. Thus, the RPS4 TIR is activated, resulting in resistance to Pseudomonas syringae. The regulation of guardee/decoy monitoring is likely much more complex than is presently known.
3.4. News-Breaking: Enzyme Activity of Plant TIR in ETI Signaling
In animal immunity, an important function of Toll-like receptors is specifically recognizing their cognate pathogen-associated molecular patterns or synthetic compounds. Most animal Toll-like receptors contain two domains, one of which—the LRR domain—is necessary for PAMP recognition, while the other—the TIR domain—functions in signaling scaffolds. Some studies of animal-TIR domain crystallization showed that animal TIR associates during PAMP recognition. Animal TIR oligomerization is required for immune signaling, leading to the inflammatory cytokine response [75][76][77]. Unlike most Toll-like receptors, Sterile Alpha and TIR Motif Containing 1 (SARM1) was shown to have a surprisingly novel function [78][79]. Specifically, the nicotinamide adenine dinucleotide (NAD) hydrolase activity of its TIR domain contributes to axon degradation. This unique function raised the hypothesis that SARM1 probably arose from other domains in the animal system, through an evolutionary transfer event [80].
In plants, after NLR activation, the subsequent signal transduction cascade leading to the hypersensitive response and expression of plant immunity is at present unresolved. Although the signaling pathway of CNLs remains unclear, a piece of TNL downstream signaling was discovered [77][81]. As TIR domains are found in both plant intracellular TNLs and the animal cell surface Toll-like receptors, researchers compared the characteristics of plant TIR and animal TIR. Wan et al. and Horsefield et al. demonstrated that the TIR domains of plant TNLs are structurally similar to the TIR domain of mammalian SARM1 and that their enzymatic activity could degrade oxidized nicotinamide adenine dinucleotide (NAD+) (Figure 2C) [77][81]. Cell death activation and NAD+ catalytic activity of plant TIRs are self-association interface-dependent, placing the TIR enzyme activity downstream of TIR oligomerization. A conserved glutamic acid was found in plant TIR NAD+-cleaving enzymes and the human SARM1 NADase [81]. Although the putative catalytic glutamic acid does not affect the TIR association, it is the key residue for TIR-NADase activation. The accumulation of enzymatic products, such as variant-cyclic ADP-Ribose (v-cADPR), ADP-Ribose, and nicotinamide, which are necessary for immune signaling, are proposed to be downstream of TIR-enzyme activation. The NADase activity of the plant TIR domain is solely required for plant immunity, since the fusion of plant TIR (not animal or bacterial TIR) to the mammalian NLR Family CARD Domain Containing 4 activates immune signaling in plants [82]. Interestingly, in both enhanced disease susceptibility 1 (eds1) and n requirement gene 1 (nrg1) mutants, the activation of RBA1 accumulates v-cADPR but fails to induce a cell-death response [81], indicating that the accumulation of enzymatic products happens upstream of EDS1-NRG1. However, from catalytic product accumulation to EDS1-NRG1 downstream signaling, an undefined gap remains.
4. Helper NLR Cooperation beyond Genetically Linked Pairs
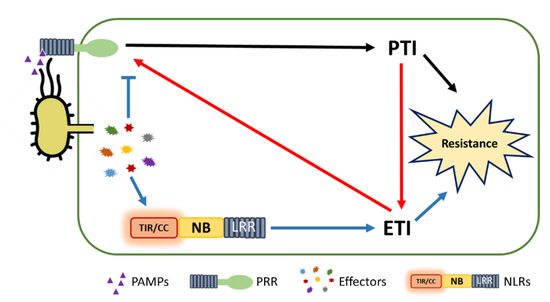
5. PTI/ETI Unity Produces Full Plant Immunity
References
- Jeffery L. Dangl; Diana M. Horvath; Brian J. Staskawicz; Pivoting the Plant Immune System from Dissection to Deployment. Science 2013, 341, 746-751, 10.1126/science.1236011.
- Alberto Pascale; Silvia Proietti; Iakovos S. Pantelides; Ioannis A. Stringlis; Modulation of the Root Microbiome by Plant Molecules: The Basis for Targeted Disease Suppression and Plant Growth Promotion. Frontiers in Plant Science 2020, 10, 1741, 10.3389/fpls.2019.01741.
- Dmitry Lapin; Guido Van Den Ackerveken; Susceptibility to plant disease: more than a failure of host immunity. Trends in Plant Science 2013, 18, 546-554, 10.1016/j.tplants.2013.05.005.
- Brian J. Staskawicz; Genetics of Plant-Pathogen Interactions Specifying Plant Disease Resistance. Plant Physiology 2001, 125, 73-76, 10.1104/pp.125.1.73.
- Jonathan D. G. Jones; Jeffery L. Dangl; The plant immune system. Nature 2006, 444, 323-329, 10.1038/nature05286.
- Yusuke Saijo; Eliza Loo; Shigetaka Yasuda; Pattern recognition receptors and signaling in plant-microbe interactions. The Plant Journal 2018, 93, 592-613, 10.1111/tpj.13808.
- Yunxia He; Jinggeng Zhou; Libo Shan; Xiangzong Meng; Plant cell surface receptor-mediated signaling – a common theme amid diversity. Journal of Cell Science 2018, 131, jcs209353, 10.1242/jcs.209353.
- Ying Wu; Jian-Min Zhou; Receptor-Like Kinases in Plant Innate Immunity. Journal of Integrative Plant Biology 2013, 55, 1271-1286, 10.1111/jipb.12123.
- Cyril Zipfel; Pattern-recognition receptors in plant innate immunity. Current Opinion in Immunology 2008, 20, 10-16, 10.1016/j.coi.2007.11.003.
- Jean Bigeard; Jean Colcombet; Heribert Hirt; Signaling Mechanisms in Pattern-Triggered Immunity (PTI). Molecular Plant 2015, 8, 521-539, 10.1016/j.molp.2014.12.022.
- Elena Jeworutzki; Rob Roelfsema; Uta Anschütz; Elzbieta Krol; J. Theo M. Elzenga; Georg Felix; Thomas Boller; Rainer Hedrich; Dirk Becker; Early signaling through the Arabidopsis pattern recognition receptors FLS2 and EFR involves Ca2+-associated opening of plasma membrane anion channels. The Plant Journal 2010, 62, 367-378, 10.1111/j.1365-313x.2010.04155.x.
- Stefanie Ranf; Lennart Eschen-Lippold; Pascal Pecher; Justin Lee; Dierk Scheel; Interplay between calcium signalling and early signalling elements during defence responses to microbe- or damage-associated molecular patterns. The Plant Journal 2011, 68, 100-113, 10.1111/j.1365-313x.2011.04671.x.
- Hironari Nomura; Toshihisa Komori; Shuhei Uemura; Yoshinobu Kanda; Koji Shimotani; Kunihisa Nakai; Tatsuya Furuichi; Kosuke Takebayashi; Takanori Sugimoto; Satoshi Sano; et al.I Nengah SuwastikaEiichiro FukusakiHirofumi YoshiokaYoichi NakahiraTakashi Shiina Chloroplast-mediated activation of plant immune signalling in Arabidopsis. Nature Communications 2012, 3, 926, 10.1038/ncomms1926.
- Jie Zhang; Feng Shao; Yan Li; Haitao Cui; Linjie Chen; Hongtao Li; Yan Zou; Chengzu Long; Lefu Lan; Jijie Chai; et al.She ChenXiaoyan TangJian-Min Zhou A Pseudomonas syringae Effector Inactivates MAPKs to Suppress PAMP-Induced Immunity in Plants. Cell Host & Microbe 2007, 1, 175-185, 10.1016/j.chom.2007.03.006.
- Zhibin Zhang; Yaling Wu; Minghui Gao; Jie Zhang; Qing Kong; Yanan Liu; Hongping Ba; Jianmin Zhou; Yuelin Zhang; Disruption of PAMP-Induced MAP Kinase Cascade by a Pseudomonas syringae Effector Activates Plant Immunity Mediated by the NB-LRR Protein SUMM2. Cell Host & Microbe 2012, 11, 253-263, 10.1016/j.chom.2012.01.015.
- Delphine Chinchilla; Cyril Zipfel; Silke Robatzek; Birgit Kemmerling; Thorsten Nürnberger; Jonathan D. G. Jones; Georg Felix; Thomas Boller; A flagellin-induced complex of the receptor FLS2 and BAK1 initiates plant defence. Nature 2007, 448, 497-500, 10.1038/nature05999.
- Byung-Wook Yun; Angela Feechan; Minghui Yin; Noor Baity Saidi; Thierry Le Bihan; Manda Yu; John W. Moore; Jeong-Gu Kang; Eunjung Kwon; Steven H. Spoel; et al.Jacqueline PallasGary Loake S-nitrosylation of NADPH oxidase regulates cell death in plant immunity. Nature 2011, 478, 264-268, 10.1038/nature10427.
- Claudia Scheler; Jörg Durner; Jeremy Astier; Nitric oxide and reactive oxygen species in plant biotic interactions. Current Opinion in Plant Biology 2013, 16, 534-539, 10.1016/j.pbi.2013.06.020.
- Yozo Okazaki; Kazuki Saito; Roles of lipids as signaling molecules and mitigators during stress response in plants. The Plant Journal 2014, 79, 584-596, 10.1111/tpj.12556.
- Zhe Wang; Xifeng Li; Xiaoting Wang; Nana Liu; Binjie Xu; Qi Peng; Zhifu Guo; Baofang Fan; Cheng Zhu; Zhixiang Chen; et al. Arabidopsis Endoplasmic Reticulum-Localized UBAC2 Proteins Interact with PAMP-INDUCED COILED-COIL to Regulate Pathogen-Induced Callose Deposition and Plant Immunity. The Plant Cell 2019, 31, 153-171, 10.1105/tpc.18.00334.
- Akinori Kiba; Masahito Nakano; Miki Hosokawa; Ivan Galis; Hiroko Nakatani; Tomonori Shinya; Kouhei Ohnishi; Yasufumi Hikichi; Phosphatidylinositol-phospholipase C2 regulates pattern-triggered immunity in Nicotiana benthamiana. Journal of Experimental Botany 2020, 71, 5027-5038, 10.1093/jxb/eraa233.
- Lin Jin; David M. Mackey; Measuring Callose Deposition, an Indicator of Cell Wall Reinforcement, During Bacterial Infection in Arabidopsis. Methods in Molecular Biology 2017, 1578, 195-205, 10.1007/978-1-4939-6859-6_16.
- Masahito Nakano; Masahiro Nishihara; Hirofumi Yoshioka; Hirotaka Takahashi; Tatsuya Sawasaki; Kouhei Ohnishi; Yasufumi Hikichi; Akinori Kiba; Suppression of DS1 Phosphatidic Acid Phosphatase Confirms Resistance to Ralstonia solanacearum in Nicotiana benthamiana. PLOS ONE 2013, 8, e75124, 10.1371/journal.pone.0075124.
- Weijie Huang; Yiran Wang; Xin Li; Yuelin Zhang; Biosynthesis and Regulation of Salicylic Acid and N-Hydroxypipecolic Acid in Plant Immunity. Molecular Plant 2020, 13, 31-41, 10.1016/j.molp.2019.12.008.
- Wei Zhang; Feng Zhao; Lihui Jiang; Cun Chen; Lintao Wu; Zhibin Liu; Different Pathogen Defense Strategies in Arabidopsis: More than Pathogen Recognition. Cells 2018, 7, 252, 10.3390/cells7120252.
- Magdaléna Bryksová; Siarhei Dabravolski; Zuzana Kučerová; Filip Zavadil Kokáš; Martina Špundová; Lucie Plíhalová; Tomáš Takáč; Jiří Grúz; Martin Hudeček; Veronika Hloušková; et al.Radoslav KoprnaOndřej NovákMiroslav StrnadOndřej PlíhalKarel Doležal Aromatic Cytokinin Arabinosides Promote PAMP-like Responses and Positively Regulate Leaf Longevity. ACS Chemical Biology 2020, 15, 1949-1963, 10.1021/acschembio.0c00306.
- Mai Jarad; Kiruthiga Mariappan; Marilia Almeida-Trapp; Michael Florian Mette; Axel Mithöfer; Naganand Rayapuram; Heribert Hirt; The Lamin-Like LITTLE NUCLEI 1 (LINC1) Regulates Pattern-Triggered Immunity and Jasmonic Acid Signaling. Frontiers in Plant Science 2020, 10, 1639, 10.3389/fpls.2019.01639.
- Raul Zavaliev; Rajinikanth Mohan; Tianyuan Chen; Xinnian Dong; Formation of NPR1 Condensates Promotes Cell Survival during the Plant Immune Response. Cell 2020, 182, 1093-1108.e18, 10.1016/j.cell.2020.07.016.
- Yasuomi Tada; Steven H. Spoel; Karolina Pajerowska-Mukhtar; Zhonglin Mou; Junqi Song; Chun Wang; Jianru Zuo; Xinnian Dong; Plant Immunity Requires Conformational Charges of NPR1 via S-Nitrosylation and Thioredoxins. Science 2008, 321, 952-956, 10.1126/science.1156970.
- Yujun Peng; Rowan Van Wersch; Yuelin Zhang; Convergent and Divergent Signaling in PAMP-Triggered Immunity and Effector-Triggered Immunity. Molecular Plant-Microbe Interactions® 2018, 31, 403-409, 10.1094/mpmi-06-17-0145-cr.
- Hanna Alhoraibi; Jean Bigeard; Naganand Rayapuram; Jean Colcombet; Heribert Hirt; Plant Immunity: The MTI-ETI Model and Beyond. Current Issues in Molecular Biology 2019, 30, 39-58, 10.21775/cimb.030.039.
- Manon Richard; Ariane Gratias; Blake C. Meyers; Valérie Geffroy; Molecular mechanisms that limit the costs of NLR-mediated resistance in plants. Molecular Plant Pathology 2018, 19, 2516-2523, 10.1111/mpp.12723.
- Solveig van Wersch; Xin Li; Stronger When Together: Clustering of Plant NLR Disease resistance Genes. Trends in Plant Science 2019, 24, 688-699, 10.1016/j.tplants.2019.05.005.
- Jeffery L. Dangl; Jonathan D. G. Jones; Plant pathogens and integrated defence responses to infection. Nature 2001, 411, 826-833, 10.1038/35081161.
- Yang Gao; Yujun Wu; Junbo Du; Yanyan Zhan; Doudou Sun; Jianxin Zhao; Shasha Zhang; Jia Li; Kai He; Both Light-Induced SA Accumulation and ETI Mediators Contribute to the Cell Death Regulated by BAK1 and BKK1. Frontiers in Plant Science 2017, 8, 622, 10.3389/fpls.2017.00622.
- Yi Tao; Zhiyi Xie; Wenqiong Chen; Jane Glazebrook; Hur-Song Chang; Bin Han; Tong Zhu; Guangzhou Zou; Fumiaki Katagiri; Quantitative Nature of Arabidopsis Responses during Compatible and Incompatible Interactions with the Bacterial Pathogen Pseudomonas syringae [W]. The Plant Cell 2003, 15, 317-330, 10.1105/tpc.007591.
- Lionel Navarro; Cyril Zipfel; Owen Rowland; Ingo Keller; Silke Robatzek; Thomas Boller; Jonathan D.G. Jones; The Transcriptional Innate Immune Response to flg22. Interplay and Overlap with Avr Gene-Dependent Defense Responses and Bacterial Pathogenesis. Plant Physiology 2004, 135, 1113-1128, 10.1104/pp.103.036749.
- Bruno Pok Man Ngou; Hee-Kyung Ahn; Pingtao Ding; Jonathan D. G. Jones; Mutual potentiation of plant immunity by cell-surface and intracellular receptors. Nature 2021, 592, 110-115, 10.1038/s41586-021-03315-7.
- Minhang Yuan; Zeyu Jiang; Guozhi Bi; Kinya Nomura; Menghui Liu; Yiping Wang; Boying Cai; Jian-Min Zhou; Sheng Yang He; Xiu-Fang Xin; et al. Pattern-recognition receptors are required for NLR-mediated plant immunity. Nature 2021, 592, 105-109, 10.1038/s41586-021-03316-6.
- Erik A. Van Der Biezen; Jonathan D.G. Jones; Plant disease-resistance proteins and the gene-for-gene concept. Trends in Biochemical Sciences 1998, 23, 454-456, 10.1016/s0968-0004(98)01311-5.
- Stella Césari; Hiroyuki Kanzaki; Tadashi Fujiwara; Maud Bernoux; Véronique Chalvon; Yoji Kawano; Ko Shimamoto; Peter Dodds; Ryohei Terauchi; Thomas Kroj; et al. The NB ‐ LRR proteins RGA 4 and RGA 5 interact functionally and physically to confer disease resistance. The EMBO Journal 2014, 33, 1941-1959, 10.15252/embj.201487923.
- Stella Cesari; Multiple strategies for pathogen perception by plant immune receptors. New Phytologist 2017, 219, 17-24, 10.1111/nph.14877.
- David Mackey; Youssef Belkhadir; Jose M. Alonso; Joseph R. Ecker; Jeffery L. Dangl; Arabidopsis RIN4 Is a Target of the Type III Virulence Effector AvrRpt2 and Modulates RPS2-Mediated Resistance. Cell 2003, 112, 379-389, 10.1016/s0092-8674(03)00040-0.
- Michael J. Axtell; Brian J. Staskawicz; Initiation of RPS2-Specified Disease Resistance in Arabidopsis Is Coupled to the AvrRpt2-Directed Elimination of RIN4. Cell 2003, 112, 369-377, 10.1016/s0092-8674(03)00036-9.
- Han-Suk Kim; Darrell Desveaux; Alex U. Singer; Priyesh Patel; John Sondek; Jeffery L. Dangl; The Pseudomonas syringae effector AvrRpt2 cleaves its C-terminally acylated target, RIN4, from Arabidopsis membranes to block RPM1 activation. Proceedings of the National Academy of Sciences 2005, 102, 6496-6501, 10.1073/pnas.0500792102.
- David Mackey; Ben F. Holt; Aaron Wiig; Jeffery L. Dangl; RIN4 Interacts with Pseudomonas syringae Type III Effector Molecules and Is Required for RPM1-Mediated Resistance in Arabidopsis. Cell 2002, 108, 743-754, 10.1016/s0092-8674(02)00661-x.
- Maxim Prokchorchik; Sera Choi; Eui‐Hwan Chung; Kyungho Won; Jeffery L. Dangl; Kee Hoon Sohn; A host target of a bacterial cysteine protease virulence effector plays a key role in convergent evolution of plant innate immune system receptors. New Phytologist 2019, 225, 1327-1342, 10.1111/nph.16218.
- Jianbin Su; Benjamin J. Spears; Sang Hee Kim; Walter Gassmann; Constant vigilance: plant functions guarded by resistance proteins. The Plant Journal 2018, 93, 637-650, 10.1111/tpj.13798.
- Jie Zhang; Wei Li; Tingting Xiang; Zixu Liu; Kristin Laluk; Xiaojun Ding; Yan Zou; Minghui Gao; Xiaojuan Zhang; She Chen; et al.Tesfaye MengisteYuelin ZhangJian-Min Zhou Receptor-like Cytoplasmic Kinases Integrate Signaling from Multiple Plant Immune Receptors and Are Targeted by a Pseudomonas syringae Effector. Cell Host & Microbe 2010, 7, 290-301, 10.1016/j.chom.2010.03.007.
- Brody J. Deyoung; Dong Qi; Sang-Hee Kim; Thomas P. Burke; Roger W. Innes; Activation of a plant nucleotide binding-leucine rich repeat disease resistance protein by a modified self protein. Cellular Microbiology 2012, 14, 1071-1084, 10.1111/j.1462-5822.2012.01779.x.
- Sang Hee Kim; Dong Qi; Tom Ashfield; Matthew Helm; Roger W. Innes; Using decoys to expand the recognition specificity of a plant disease resistance protein. Science 2016, 351, 684-687, 10.1126/science.aad3436.
- Dong Qi; Brody J. DeYoung; Roger W. Innes; Structure-Function Analysis of the Coiled-Coil and Leucine-Rich Repeat Domains of the RPS5 Disease Resistance Protein. Plant Physiology 2012, 158, 1819-1832, 10.1104/pp.112.194035.
- Feng Shao; Catherine Golstein; Jules Ade; Mark Stoutemyer; Jack E. Dixon; Roger W. Innes; Cleavage of Arabidopsis PBS1 by a Bacterial Type III Effector. Science 2003, 301, 1230-1233, 10.1126/science.1085671.
- Doris Birker; Katharina Heidrich; Hiroyuki Takahara; Mari Narusaka; Laurent Deslandes; Yoshihiro Narusaka; Matthieu Reymond; Jane E. Parker; Richard O’Connell; A locus conferring resistance toColletotrichum higginsianumis shared by four geographically distinct Arabidopsis accessions. The Plant Journal 2009, 60, 602-613, 10.1111/j.1365-313x.2009.03984.x.
- Mari Narusaka; Ken Shirasu; Yoshiteru Noutoshi; Yasuyuki Kubo; Tomonori Shiraishi; Masaki Iwabuchi; Yoshihiro Narusaka; RRS1andRPS4provide a dualResistance-gene system against fungal and bacterial pathogens. The Plant Journal 2009, 60, 218-226, 10.1111/j.1365-313x.2009.03949.x.
- Clementine Le Roux; Gaëlle Huet; Alain Jauneau; Laurent Camborde; Dominique Trémousaygue; Alexandra Kraut; Binbin Zhou; Marie Levaillant; Hiroaki Adachi; Hirofumi Yoshioka; et al.Sylvain RaffaeleRichard BerthoméYohann CoutéJane E. ParkerLaurent Deslandes A Receptor Pair with an Integrated Decoy Converts Pathogen Disabling of Transcription Factors to Immunity. Cell 2015, 161, 1074-1088, 10.1016/j.cell.2015.04.025.
- Panagiotis F. Sarris; Zane Duxbury; Sung Un Huh; Yan Ma; Cécile Segonzac; Jan Sklenar; Paul Derbyshire; Volkan Cevik; Ghanasyam Rallapalli; Simon B. Saucet; et al.Lennart WirthmuellerFrank L.H. MenkeKee Hoon SohnJonathan D.G. Jones A Plant Immune Receptor Detects Pathogen Effectors that Target WRKY Transcription Factors. Cell 2015, 161, 1089-1100, 10.1016/j.cell.2015.04.024.
- Stella Cesari; Maud Bernoux; Philippe Moncuquet; Thomas Kroj; Peter N. Dodds; A novel conserved mechanism for plant NLR protein pairs: the “integrated decoy†hypothesis. Frontiers in Plant Science 2014, 5, 606, 10.3389/fpls.2014.00606.
- Thomas Griebel; Takaki Maekawa; Jane E. Parker; NOD-like receptor cooperativity in effector-triggered immunity. Trends in Immunology 2014, 35, 562-570, 10.1016/j.it.2014.09.005.
- Paul Kapos; Karen Thulasi Devendrakumar; Xin Li; Plant NLRs: From discovery to application. Plant Science 2019, 279, 3-18, 10.1016/j.plantsci.2018.03.010.
- Freddy Monteiro; Marc T. Nishimura; Structural, Functional, and Genomic Diversity of Plant NLR Proteins: An Evolved Resource for Rational Engineering of Plant Immunity. Annual Review of Phytopathology 2018, 56, 243-267, 10.1146/annurev-phyto-080417-045817.
- Lachlan Casey; Peter Lavrencic; Adam R. Bentham; Stella Cesari; Daniel Ericsson; Tristan Croll; Dušan Turk; Peter A. Anderson; Alan E. Mark; Peter N. Dodds; et al.Mehdi MobliBostjan KobeSimon J. Williams The CC domain structure from the wheat stem rust resistance protein Sr33 challenges paradigms for dimerization in plant NLR proteins. Proceedings of the National Academy of Sciences 2016, 113, 12856-12861, 10.1073/pnas.1609922113.
- Stella Cesari; John Moore; Chunhong Chen; Daryl Webb; Sambasivam Periyannan; Rohit Mago; Maud Bernoux; Evans S. Lagudah; Peter N. Dodds; Cytosolic activation of cell death and stem rust resistance by cereal MLA-family CC–NLR proteins. Proceedings of the National Academy of Sciences 2016, 113, 10204-10209, 10.1073/pnas.1605483113.
- Jizong Wang; Meijuan Hu; Jia Wang; Jinfeng Qi; Zhifu Han; Guoxun Wang; Yijun Qi; Hong-Wei Wang; Jian-Min Zhou; Jijie Chai; et al. Reconstitution and structure of a plant NLR resistosome conferring immunity. Science 2019, 364, eaav5870, 10.1126/science.aav5870.
- Xiaoxiao Zhang; Maud Bernoux; Adam R. Bentham; Toby Newman; Thomas Ve; Lachlan Casey; Tom M. Raaymakers; Jian Hu; Tristan I. Croll; Karl J. Schreiber; et al.Brian J. StaskawiczPeter A. AndersonKee Hoon SohnSimon J. WilliamsPeter N. DoddsBostjan Kobe Multiple functional self-association interfaces in plant TIR domains. Proceedings of the National Academy of Sciences 2017, 114, E2046-E2052, 10.1073/pnas.1621248114.
- Shoucai Ma; Dmitry Lapin; Li Liu; Yue Sun; Wen Song; Xiaoxiao Zhang; Elke Logemann; Dongli Yu; Jia Wang; Jan Jirschitzka; et al.Zhifu HanPaul Schulze-LefertJane E. ParkerJijie Chai Direct pathogen-induced assembly of an NLR immune receptor complex to form a holoenzyme. Science 2020, 370, eabe3069, 10.1126/science.abe3069.
- Maud Bernoux; Hayden Burdett; Simon J. Williams; Xiaoxiao Zhang; Chunhong Chen; Kim Newell; Gregory J. Lawrence; Bostjan Kobe; Jeffrey G. Ellis; Peter A. Anderson; et al.Peter N. Dodds Comparative Analysis of the Flax Immune Receptors L6 and L7 Suggests an Equilibrium-Based Switch Activation Model. The Plant Cell 2016, 28, 146-159, 10.1105/tpc.15.00303.
- Hailong Guo; Hee-Kyung Ahn; Jan Sklenar; Jianhua Huang; Yan Ma; Pingtao Ding; Frank L.H. Menke; Jonathan D.G. Jones; Phosphorylation-Regulated Activation of the Arabidopsis RRS1-R/RPS4 Immune Receptor Complex Reveals Two Distinct Effector Recognition Mechanisms. Cell Host & Microbe 2020, 27, 769-781.e6, 10.1016/j.chom.2020.03.008.
- Sung Un Huh; Volkan Cevik; Pingtao Ding; Zane Duxbury; Yan Ma; Laurence Tomlinson; Panagiotis F. Sarris; Jonathan D. G. Jones; Protein-protein interactions in the RPS4/RRS1 immune receptor complex. PLOS Pathogens 2017, 13, e1006376, 10.1371/journal.ppat.1006376.
- Raoul Martin; Tiancong Qi; Haibo Zhang; Furong Liu; Miles King; Claire Toth; Eva Nogales; Brian J. Staskawicz; Structure of the activated ROQ1 resistosome directly recognizing the pathogen effector XopQ. Science 2020, 370, eabd9993, 10.1126/science.abd9993.
- Eva Sinapidou; Kevin Williams; Lucy Nott; Saleha Bahkt; Mahmut Tor; Ian Crute; Peter Bittner-Eddy; Jim Beynon; Two TIR:NB:LRR genes are required to specify resistance toPeronospora parasiticaisolate Cala2 inArabidopsis. The Plant Journal 2004, 38, 898-909, 10.1111/j.1365-313x.2004.02099.x.
- Simon B. Saucet; Yan Ma; Panagiotis F. Sarris; Oliver J. Furzer; Kee Hoon Sohn; Jonathan D.G. Jones; Two linked pairs of Arabidopsis TNL resistance genes independently confer recognition of bacterial effector AvrRps4. Nature Communications 2015, 6, 6338, 10.1038/ncomms7338.
- Eui-Hwan Chung; Farid El-Kasmi; Yijian He; Alex Loehr; Jeffery L. Dangl; A Plant Phosphoswitch Platform Repeatedly Targeted by Type III Effector Proteins Regulates the Output of Both Tiers of Plant Immune Receptors. Cell Host & Microbe 2014, 16, 484-494, 10.1016/j.chom.2014.09.004.
- Thomas J. Redditt; Eui-Hwan Chung; Hana Zand Karimi; Natalie Rodibaugh; Yixiang Zhang; Jonathan C. Trinidad; Jin Hee Kim; Qian Zhou; Mingzhe Shen; Jeffery L. Dangl; et al.David M. MackeyRoger W. Innes AvrRpm1 Functions as an ADP-Ribosyl Transferase to Modify NOI-domain Containing Proteins, Including Arabidopsis and Soybean RPM1-interacting Protein 4. The Plant Cell 2019, 31, 2664-2681, 10.1105/tpc.19.00020.
- Kiyoshi Takeda; Shizuo Akira; Toll‐Like Receptors. Current Protocols in Immunology 2015, 109, 14.12.1-14.12.10, 10.1002/0471142735.im1412s109.
- Himanshu Kumar; Taro Kawai; Shizuo Akira; Toll-like receptors and innate immunity. Biochemical and Biophysical Research Communications 2009, 388, 621-625, 10.1016/j.bbrc.2009.08.062.
- Shane Horsefield; Hayden Burdett; Xiaoxiao Zhang; Mohammad K. Manik; Yun Shi; Jian Chen; Tiancong Qi; Jonathan Gilley; Jhih-Siang Lai; Maxwell X. Rank; et al.Lachlan W. CaseyWeixi GuDaniel J. EricssonGabriel FoleyRobert O. HughesTodd BosanacMark Von ItzsteinJohn P. RathjenJeffrey D. NansonMikael BodenIan B. DrySimon J. WilliamsBrian J. StaskawiczMichael P. ColemanThomas VePeter N. DoddsBostjan Kobe NAD+ cleavage activity by animal and plant TIR domains in cell death pathways. Science 2019, 365, 793-799, 10.1126/science.aax1911.
- Josiah Gerdts; E. J. Brace; Y. Sasaki; Aaron DiAntonio; Jeffrey Milbrandt; SARM1 activation triggers axon degeneration locally via NAD+ destruction. Science 2015, 348, 453-457, 10.1126/science.1258366.
- Kow Essuman; Daniel W. Summers; Yo Sasaki; Xianrong Mao; Aldrin Kay Yuen Yim; Aaron DiAntonio; Jeffrey Milbrandt; TIR Domain Proteins Are an Ancient Family of NAD+-Consuming Enzymes. Current Biology 2018, 28, 421-430.e4, 10.1016/j.cub.2017.12.024.
- Adam M. Bayless; Marc T. Nishimura; Enzymatic Functions for Toll/Interleukin-1 Receptor Domain Proteins in the Plant Immune System. Frontiers in Genetics 2020, 11, 539, 10.3389/fgene.2020.00539.
- Li Wan; Kow Essuman; Ryan G. Anderson; Yo Sasaki; Freddy Monteiro; Eui-Hwan Chung; Erin Osborne Nishimura; Aaron DiAntonio; Jeffrey Milbrandt; Jeffery L. Dangl; et al.Marc T. Nishimura TIR domains of plant immune receptors are NAD+-cleaving enzymes that promote cell death. Science 2019, 365, 799-803, 10.1126/science.aax1771.
- Zane Duxbury; Shanshan Wang; Craig I. MacKenzie; Jeannette L. Tenthorey; Xiaoxiao Zhang; Sung Un Huh; Lanxi Hu; Lionel Hill; Pok Man Ngou; Pingtao Ding; et al.Jian ChenYan MaHailong GuoBaptiste CastelPanagiotis N. MoschouMaud BernouxPeter N. DoddsRussell E. VanceJonathan D. G. Jones Induced proximity of a TIR signaling domain on a plant-mammalian NLR chimera activates defense in plants. Proceedings of the National Academy of Sciences 2020, 117, 18832-18839, 10.1073/pnas.2001185117.
- Sarah Collier; Louis-Philippe Hamel; Peter Moffett; Cell Death Mediated by the N-Terminal Domains of a Unique and Highly Conserved Class of NB-LRR Protein. Molecular Plant-Microbe Interactions® 2011, 24, 918-931, 10.1094/mpmi-03-11-0050.
- Suzan H.E.J. Gabriëls; Jack H. Vossen; Sophia K. Ekengren; Gerben Van Ooijen; Ahmed M. Abd-El-Haliem; Grardy C.M. Van Den Berg; Daphne Y. Rainey; Gregory B. Martin; Frank Takken; Pierre J.G.M. De Wit; et al.Matthieu H.A.J. Joosten An NB-LRR protein required for HR signalling mediated by both extra- and intracellular resistance proteins. The Plant Journal 2007, 50, 14-28, 10.1111/j.1365-313x.2007.03027.x.
- Shunyuan Xiao; Ozer Calis; Elaine Patrick; Guangmin Zhang; Piyavadee Charoenwattana; Paul Muskett; Jane E. Parker; John G. Turner; The atypical resistance gene, RPW8, recruits components of basal defence for powdery mildew resistance in Arabidopsis. The Plant Journal 2005, 42, 95-110, 10.1111/j.1365-313x.2005.02356.x.
- Vera Bonardi; Saijun Tang; Anna Stallmann; Melinda Roberts; Karen Cherkis; Jeffery L. Dangl; Expanded functions for a family of plant intracellular immune receptors beyond specific recognition of pathogen effectors. Proceedings of the National Academy of Sciences 2011, 108, 16463-16468, 10.1073/pnas.1113726108.
- Oliver Xiaoou Dong; Meixuezi Tong; Vera Bonardi; Farid El Kasmi; Virginia Woloshen; Lisa K. Wünsch; Jeffery L. Dangl; Xin Li; TNL ‐mediated immunity in A rabidopsis requires complex regulation of the redundant ADR 1 gene family. New Phytologist 2016, 210, 960-973, 10.1111/nph.13821.
- Zhongshou Wu; Meng Li; Oliver Xiaoou Dong; Shitou Xia; Wanwan Liang; Yongkang Bao; Geoffrey Wasteneys; Xin Li; Differential regulation of TNL‐mediated immune signaling by redundant helper CNLs. New Phytologist 2018, 222, 938-953, 10.1111/nph.15665.
- Jack R. Peart; Pere Mestre; Rui Lu; Isabelle Malcuit; David C. Baulcombe; NRG1, a CC-NB-LRR Protein, together with N, a TIR-NB-LRR Protein, Mediates Resistance against Tobacco Mosaic Virus. Current Biology 2005, 15, 968-973, 10.1016/j.cub.2005.04.053.
- Tiancong Qi; Kyungyong Seong; Daniela P. T. Thomazella; Joonyoung Ryan Kim; Julie Pham; Eunyoung Seo; Myeong-Je Cho; Alex Schultink; Brian J. Staskawicz; NRG1 functions downstream of EDS1 to regulate TIR-NLR-mediated plant immunity in Nicotiana benthamiana. Proceedings of the National Academy of Sciences 2018, 115, E10979-E10987, 10.1073/pnas.1814856115.
- Baptiste Castel; Pok‐Man Ngou; Volkan Cevik; Amey Redkar; Dae‐Sung Kim; Ying Yang; Pingtao Ding; Jonathan D. G. Jones; Diverse NLR immune receptors activate defence via the RPW 8‐ NLR NRG 1. New Phytologist 2018, 222, 966-980, 10.1111/nph.15659.
- Rico A. Caldo; Dan Nettleton; Roger P. Wise; María Del Mar Castellano; María Beatrice Boniotti; Elena Caro; Arp Schnittger; Crisanto Gutierrez; Interaction-Dependent Gene Expression in Mla-Specified Response to Barley Powdery Mildew[W]. The Plant Cell 2004, 16, 2514-2528, 10.1105/tpc.104.023382.
- Noriyuki Hatsugai; Daisuke Igarashi; Keisuke Mase; You Lu; Yayoi Tsuda; Suma Chakravarthy; Hai-Lei Wei; Joseph Foley; Alan Collmer; Jane Glazebrook; et al.Fumiaki Katagiri A plant effector‐triggered immunity signaling sector is inhibited by pattern‐triggered immunity. The EMBO Journal 2017, 36, 2758-2769, 10.15252/embj.201796529.





