
| Version | Summary | Created by | Modification | Content Size | Created at | Operation |
|---|---|---|---|---|---|---|
| 1 | Fabiana Passaro | + 911 word(s) | 911 | 2021-05-14 11:14:27 | | | |
| 2 | Rita Xu | + 242 word(s) | 1153 | 2021-05-28 08:50:36 | | |
Video Upload Options
Cell reprogramming can either refer to a direct conversion of a specialized cell into another or to a reversal of a somatic cell into an induced pluripotent stem cell (iPSC). It implies a peculiar modification of the epigenetic asset and gene regulatory networks needed for a new cell, to better fit the new phenotype of the incoming cell type. Cellular reprogramming also implies a metabolic rearrangement, similar to that observed upon tumorigenesis, with a transition from oxidative phosphorylation to aerobic glycolysis. The induction of a reprogramming process requires a nexus of signaling pathways, mixing a range of local and systemic information, and accumulating evidence points to the crucial role exerted by the Hippo pathway components Yes-Associated Protein (YAP) and Transcriptional Co-activator with PDZ-binding Motif (TAZ).
1. Introduction
Cell reprogramming is the process of converting a somatic, specialized, cell into a different cell type. The term “reprogramming” can refer either to a direct conversion of a specialized cell into another or to a reversal of a somatic cell into an induced pluripotent stem cell (iPSCs) [1]. In any case, cell reprogramming implies a dramatic re-set of the whole cell, starting from the erasure and re-establishment of epigenetic marks acquired during development [2]. Reprogramming modifies the gene regulatory network of the resident cell type, to better fit the characteristics of the incoming target cell type, and this ultimately leads to changes in cell behavior, proliferation and metabolic rate, allowing the acquisition of a new phenotype [1]. The same changes in cell fate specifications can also be associated with some pathological cellular states. Indeed, tumorigenesis requires the reprogramming of a specific cell type into a new one with different characteristics in terms of gene expression, proliferation and metabolism and several types of tumors further dedifferentiate to acquire features different from those of the initial tumor mass [3][4]. As such, the somatic state is plastic, and can be modified by the right set of extracellular and intracellular signals.
Cellular reprogramming also implies an essential metabolic rearrangement, especially upon transition to induced pluripotency [5][6]. Somatic cells are often quiescent and their metabolism relies on oxidative phosphorylation (OXPHOS), whereas embryonic stem cells (ESCs) change their cell metabolism in diverse ways to meet the requirements for proliferation, gaining energy by increasing glucose conversion into lactate, even in oxygen-rich conditions [5][6]. This “aerobic glycolysis”, also named the Warburg effect from its discoverer Otto Warburg [7], depends on an increase in glucose uptake and highly activated glycolysis, and was originally employed to describe the metabolic switch occurring upon tumor progression. In the last decade, several common characteristics between cancer cell transformation and reprogramming have been established, with particular emphasis on the connections between Epithelial-to-Mesenchymal Transition (EMT) and its reversal Mesenchymal-to-Epithelial Transition (MET) [8], energy metabolism [9], DNA methylation and histone modification [10]. Very recently, a multi-layered analysis of mouse iPSC generation revealed stage-specific interdependences between EMT and metabolism, controlling an intermediary pluripotent state that can be converted to naïve and primed states [11].
Thus, the induction of a reprogramming process requires a nexus of signaling pathways, integrating a range of local and systemic information to control a series of cellular features, and accumulating evidence points to the crucial role exerted by the Hippo pathway components Yes-Associated Protein (YAP) and Transcriptional Co-activator with PDZ-binding Motif (TAZ). As downstream effectors of the Hippo signaling cascade [12], YAP and TAZ are crucial players of various cellular processes regulating organ growth, cellular plasticity, cell proliferation and stem cell biology [13][14][15]. Their activity is tightly regulated by a variety of upstream inputs, including the energy status of the cell, the nutrient availability, the composition of the tissue microenvironment and architecture, and extrinsic biochemical signals [16].
Over the last decade, many studies have highlighted the importance of metabolic remodeling during the reprogramming [17].
It is well known that metabolism has an active role in the regulation of cell identity and behavior during embryonic development, and recent studies demonstrate that metabolic remodeling is not only associated with reprogramming, but also appears to mediate the process [17].
All evidence supporting the involvement of YAP/TAZ in the metabolic remodeling of cancer cells [18] suggest a possible role for these proteins also in metabolic changes observed upon cellular reprogramming.
2. The Hippo Kinase Cascade
Activation of the Hippo pathway prevents the nuclear translocation of YAP and TAZ (Figure 1). The kinase cascade begins with the dimerization of the Mammalian Sterile 20-like 1/2 (MST1/2, also named as STK4/3), which phosphorylates Salvador (SAV1) and MOB kinase activator 1A/B (MOB1A/B) to recruit Large Tumor Suppressor homolog 1/2 (LATS1/2). The subsequent phosphorylation of LATS1 on T1079 and LATS2 on T1041 is assisted by NF2, which interacts with LATS1/2 to bring it to the plasma membrane [19][20]. NF2, in its complexed form with KIBRA / WWC1 or Angiomotin (AMOT), can also induce LATS1/2 activation [21][22][23]. Subsequently, LATS1/2 undergoes autophosphorylation and activation [24]. In the absence of MST1/2, LATS1/2 can be activated through phosphorylation by proteins belonging to the MAPK4 family [25] and by proteins of the TAO kinase family [26]. The latter are also capable of stimulating indirect activation of LATS1/2, through the phosphorylation and activation of MST1/2 [27][28].
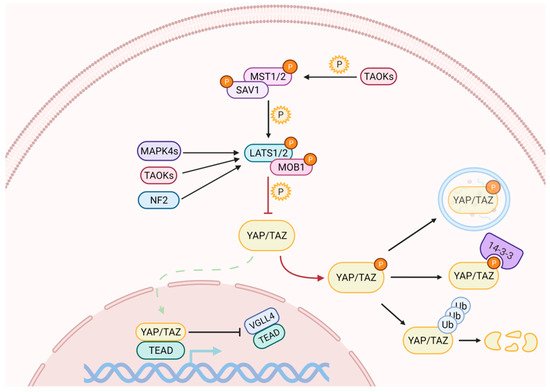
Figure 1. The Hippo signaling pathway comprises a cascade of phosphorylation and activation of kinases, which ultimately regulate YAP and TAZ subcellular localization. When the pathway is on, the activation of LATS1/2 triggers the inhibitory phosphorylation of YAP and TAZ. Phosphorylation on different residues may cause their cytoplasmic retention and/or their degradation through the ubiquitin-proteasome pathway. YAP can also be degraded by autophagy. When the pathway is off, YAP/TAZ dephosphorylation allows their nuclear import, where they compete with VGLL4 for binding to TEAD transcription factors to induce the expression of target genes.
3. Conclusion
YAP/TAZ activation may regulate the switch from OXPHOS to glycolysis not only in tumor cells, but also in cell reprogramming to sustain the increasing in the proliferation rate of somatic cells undergoing reprogramming. Along with aerobic glycolysis, another hallmark of metabolic reprogramming is the enhanced glutaminolysis, which serves to directly fuel the TCA cycle and produce key metabolites for cell growth as well as ATP. Many glutamine-metabolizing enzymes are transcriptionally governed by YAP [29] as well as main transcription factors involved in the acquisition of the transient hyper-energetic state, like c-MYC, HIF-1α, NRF2 and OCT4 [13]. Furthermore, YAP and TAZ regulate mitochondrial biogenesis and size [29]. Very recently, a novel role of YAP/TAZ in modulation of cell plasticity has been discovered. Totaro et al., found that YAP/TAZ transcriptionally control autophagy by regulating autophagosomal degradation into autolysosomes, and linked this process to the YAP/TAZ-mediated dedifferentiation and acquisition of self-renewing properties of normal cells. The authors, indeed, propose that YAP/TAZ-driven autophagy represents a form of “cytoplasmic” reprogramming, a checkpoint ensuring that YAP/TAZ transcriptional rewiring in the nucleus goes hand in hand with the need of cytoplasmic and whole-cell renovation and restructuring [30].
Finally, YAP/TAZ can potentially exert profound effects on cell plasticity and metabolic reprogramming by affecting other processes like epigenetic modifications to chromatin structure and post-transcriptional microRNAs (miRNAs) processing [16][30].
References
- Yin, F.; Yu, J.; Zheng, Y.; Chen, Q.; Zhang, N.; Pan, D. Spatial organization of Hippo signaling at the plasma membrane mediated by the tumor suppressor Merlin/NF2. Cell 2013, 154, 1342–1355.
- Glantschnig, H.; Rodan, G.A.; Reszka, A.A. Mapping of MST1 kinase sites of phosphorylation. Activation and autophosphorylation. J. Biol. Chem. 2002, 277, 42987–42996.
- Genevet, A.; Wehr, M.C.; Brain, R.; Thompson, B.J.; Tapon, N. Kibra is a regulator of the Salvador/Warts/Hippo signaling network. Dev. Cell 2010, 18, 300–308.
- Hirate, Y.; Hirahara, S.; Inoue, K.; Suzuki, A.; Alarcon, V.B.; Akimoto, K.; Hirai, T.; Hara, T.; Adachi, M.; Chida, K.; et al. Polarity-dependent distribution of angiomotin localizes Hippo signaling in preimplantation embryos. Curr. Biol. 2013, 23, 1181–1194.
- Li, Y.; Zhou, H.; Li, F.; Chan, S.W.; Lin, Z.; Wei, Z.; Yang, Z.; Guo, F.; Lim, W.J.; Xing, W.; et al. Angiomotin binding-induced activation of Merlin/NF2 in the Hippo pathway. Cell Res. 2015, 25, 801–817.
- Chan, E.H.; Nousiainen, M.; Chalamalasetty, R.B.; Schäfer, A.; Nigg, E.A.; Sillje, H.H.W. The Ste20-like kinase Mst2 activates the human large tumor suppressor kinase Lats1. Oncogene 2005, 24, 2076–2086.
- Meng, Z.; Moroishi, T.; Mottier-Pavie, V.; Plouffe, S.W.; Hansen, C.G.; Hong, A.W.; Park, H.W.; Mo, J.S.; Lu, W.; Lu, S.; et al. MAP4K family kinases act in parallel to MST1/2 to activate LATS1/2 in the Hippo pathway. Nat. Commun. 2015, 6, 8357.
- Plouffe, S.W.; Meng, Z.; Lin, K.C.; Lin, B.; Hong, A.W.; Chun, J.V.; Guan, K.L. Characterization of Hippo Pathway Components by Gene Inactivation. Mol. Cell 2016, 64, 993–1008.
- Boggiano, J.C.; Vanderzalm, P.J.; Fehon, R.G. Tao-1 phosphorylates Hippo/MST kinases to regulate the Hippo-Salvador-Warts tumor suppressor pathway. Dev. Cell 2011, 21, 888–895.
- Poon, C.L.; Lin, J.I.; Zhang, X.; Harvey, K.F. The sterile 20-like kinase Tao-1 controls tissue growth by regulating the Salvador-Warts-Hippo pathway. Dev. Cell 2011, 21, 896–906.
- Sun, H.; Yang, X.; Liang, L.; Zhang, M.; Li, Y.; Chen, J.; Wang, F.; Yang, T.; Meng, F.; Lai, X.; et al. Metabolic switch and epithelial-mesenchymal transition cooperate to regulate pluripotency. EMBO J. 2020, 39, e102961.
- Yu, F.X.; Zhao, B.; Guan, K.L. Hippo Pathway in Organ Size Control, Tissue Homeostasis, and Cancer. Cell 2015, 163, 811–828.
- Heng, B.C.; Zhang, X.; Aubel, D.; Bai, Y.; Li, X.; Li, X.; Wei, Y.; Fussenegger, M.; Deng, X. Role of YAP/TAZ in Cell Lineage Fate Determination and Related Signaling Pathways. Front. Cell Dev. Biol. 2020, 8, 735.
- Moya, I.M.; Halder, G. Hippo–YAP/TAZ signalling in organ regeneration and regenerative medicine. Nat. Rev. Mol. Cell Biol. 2019, 20, 211–226.
- Davis, J.R.; Tapon, N. Hippo signalling during development. Development 2019, 146, dev167106.
- Pocaterra, A.; Romani, P.; Dupont, S. YAP/TAZ functions and their regulation at a glance. J. Cell Sci. 2020, 133, jcs230425.
- 114. Folmes, C. D.; Nelson, T. J.; Martinez-Fernandez, A.; Arrell, D. K.; Lindor, J. Z.; et al. Somatic oxidative bioenergetics transitions into pluripotency‐dependent glycolysis to facilitate nuclear reprogramming. Cell Metab. (2011) 14(2):264-271. doi:10.1016/j.cmet.2011.06.011
- Yamaguchi, H.; Taouk, G.M. A potential role of YAP/TAZ in the interplay between metastasis and metabolic alterations. Front Oncol (2020) 10:928. Doi:10.3389/fonc.2020.00928
- Koo, J.H.; Guan, K.L. Interplay between YAP/TAZ and Metabolism. Cell Metab. (2018) 28(2):196-206. doi:10.1016/j.cmet.2018.07.010
- Totaro, A.; Zhuang, Q.; Panciera, T.; Battilana, G.; Azzolin, L.; et al. Cell phenotypic plasticity requires autophagic flux driven by YAP/TAZ mechanotransduction. Proc Natl Acad Sci U S A. (2019) 116(36):17848-17857. doi:10.1073/pnas.1908228116




