2. Identification
When first discovered in 1959, the suggested name for the bacterium was
Flavobacterium meningosepticum, which was later recommended to be changed to
Chryseobacterium meningosepticum (in 1994) [
29]. In 2005, it was assigned to the genus
Elizabethkingia (named after the first scientist to report its’ discovery, Elizabeth King) under the Flavobacteriaceae family based on 16S rRNA phylogenetic studies [
30]. Recently, whole-genome sequence analysis along with optical mapping and MALDI-TOF mass spectrometry led to the revision of the genus
Elizabethkingia into eight species, namely
E. meningoseptica,
E. miricola,
E. anophelis,
E. bruuniana,
E. ursingii,
E. occulta [
31],
E. argenteiflava sp. nov. [
32], and the latest
E. umeracha [
33].
Since correct identification of
Elizabethkingia is difficult using traditional microbiological methods and misidentification of
E. anophelis with
E. meningoseptica has been found to be common (Lau et al., 2016), it is therefore highly likely for this pathogen to be underreported. Correct identification of the organism is crucial for the diagnosis and management strategies, as
E. anophelis is a nososcomial pathogen [
34]. Hence, differentiation between
E. anophelis and
E. meningoseptica requires accurate microbial identification, but the phenotypic similarities between
E. anophelis and
E. meningoseptica present a challenge to accurate identification, particularly for clinically derived isolates; 16S rRNA gene analysis had identified a 98.6% similarity between
E. meningoseptica and
E. anophelis, which has often led to the misidentification of these bacteria [
34].
The four automated bacterial identification systems that are commonly used in diagnostic laboratories are: (1) API/ID32 Phenotyping Kits (bioMérieux, Marcy l’Etoile, France); (2) Phoenix 100 ID/AST Automated Microbiology System (Becton Dickinson Co., Sparks, MD, USA); (3) VITEK 2 Automated Identification System [
35]; and (4) MALDI-TOF MS System (bioMérieux, Marcy l’Etoile, France) [
36]. At the time of writing this research, the four microbial identification systems that are listed above do not, however, contain all eight species of
Elizabethkingia in their reference spectra database. Studies have also shown that misidentification of
Elizabethkingia was rife using these automated identification systems, with
E. anophelis commonly misidentified as
E. meningoseptica [
1,
35,
37]. When the accuracy of the API/ID32, Phoenix 100 ID/AST, Vitek 2, and Vitek MS
Elizabethkingia, clinical isolate identifications were compared with 16S rRNA gene sequencing; it was reported that species identification concordance between these identification systems and 16S rRNA gene sequencing was low at only 24.5–26.5% [
35]. Nevertheless, MALDI-TOF MS systems with amended databases (labeled as “research-use only” system) either in the Vitek MS Knowledge Base v3.2 and Bruker MALDI Biotyper Library (Bruker Daltonics GmbH, Bremen, Germany) are now able to reliably differentiate
E. meningoseptica from
E. anophelis, but not the remaining species of the genus
Elizabethkingia [
35]. In a recent report of 22 clinical and 6 environmental hospital isolates from Queensland, Australia, Burnard et al. (2020) showed that the VITEK MS Knowledge Base v3.2 had a 96.2% accuracy in identifying
Elizabethkingia, with a solitary isolate of
E. bruuniana being the only species that was misidentified. Whole-genome sequencing confirmed that the majority of the isolates were
E. anophelis (
n = 22), with the rest being
E. miricola (
n = 3),
E. meningoseptica (
n = 2), and
E. bruuniana (
n = 1) [
38].
In the near future, the inclusion of novel Elizabethkingia species spectra into the databases should ensure highly accurate identification using MALDI-TOF MS systems, making it a reliable identification tool in lieu of whole-genome sequencing.
3. Antibiotic Resistance
Elizabethkingia are intrinsically resistant to most β-lactams, β-lactam/lactamase inhibitors, and carbapenems due to the presence of two unique class B metallo-β-lactamases (MBLs), namely
blaBlaB and
blaGOB, along with a class A extended-spectrum β-lactamase (ESBL),
blaCME [
39,
40,
41].
Elizabethkingia are the only known bacteria thus far with multiple chromosomally encoded MBLs [
42]. Reports of subclasses of MBL genes such as
blaBlaB-1 [
40],
blaBlaB, and
blaGOB in both
E. meningoseptica and
E. anophelis [
41], as well as
blaBlaB-16 and
blaGOB-19 in
E. miricola isolated from a black-spotted frog in China [
43], make
Elizabethkingia spp. a possible environmental reservoir for β-lactam resistance.
Elizabethkingia isolates are frequently resistant to aminoglycosides, macrolides, tetracycline, and vancomycin but show variable susceptibility to piperacillin, piperacillin-tazobactam, fluoroquinolones, minocycline, tigecycline, and trimethoprim-sulfamethoxazole [
3,
38,
41,
44,
45,
46]; cephalosporins, monobactams, and moderate susceptibilities to piperacillin [
47,
48,
49], ceftazidime, colistin, and meropenem [
50]; and levofloxacin [
51]. There are currently no established MIC breakpoints for
Elizabethkingia, and susceptibilities are largely reported based on
Enterobacteriaceae breakpoints of the Clinical and Laboratory Standards Institute (CLSI) M100 guidelines and/or the European Committee on Antimicrobial Susceptibility Testing (EUCAST) pharmacokinetic–pharmacodynamic (PK–PD) “non-species” breakpoints [
37,
38]. It has been pointed out that susceptibilities, especially for vancomycin and piperacillin-tazobactam as determined by disk diffusion and E-test, are deemed unreliable and inaccurate for
Elizabethkingia, and broth microdilution is instead recommended for susceptibility determination [
45,
52]. Although successful therapy has been attributed to rifampicin, there has been a report of bacterial resistance after three days of starting treatment [
53]. A similar case was reported for an
E. meningoseptica isolate in the Kuala Lumpur General Hospital, which developed resistance during treatment to cefepime, a cephalosporin antibiotic that is normally highly active against both Gram-positive and Gram-negative organisms [
54].
Using disk diffusion, Lau, Chow [
1] reported 21
Elizabethkingia isolates from Hong Kong as susceptible to vancomycin. However, studies using broth microdilution tests on isolates from Taiwan [
45,
55] and Australia [
38] indicated that the isolates are likely non-susceptible based on the high MICs obtained (that ranged between 8 and 256 µg/mL). Similar ranges of vancomycin MICs were obtained by Han et al. [
37], who investigated
Elizabethkingia isolates from South Korea using the agar dilution method and concluded that all isolates were non-susceptible based on the interpretive criteria used for
Staphylococcus spp. The vancomycin resistance gene,
vanW, was reported in the majority of
Elizabethkingia genomes, although the exact function of
vanW is currently unknown [
38,
46]. However, mutations in
vanW have been identified in microorganisms with VanB-type glycopeptide resistance [
46,
56]. In view of these facts and despite some anecdotal reports of success in using intravenous vancomycin alone to treat
Elizabethkingia infections [
57,
58], it was recommended that even if intravenous vancomycin is the favored therapy for
Elizabethkingia meningitis, ciprofloxacin, linezolid, or rifampicin should also be included until future clinical studies could be carried out to conclusively determine the clinical efficacy of these vancomycin-combination regiments for treatment [
52].
One of the earliest reports of the whole-genome sequences of
Elizabethkingia spp. strains from Southeast Asia was from Singapore, whereby sputum isolates obtained from three patients (NUHP1, NUHP2, and NUHP3) and four from the hospital’s sink (NUH1, NUH4, NUH6, and NUH11) at the National University Hospital, Singapore, were compared against five previously sequenced
E. anophelis strains Ag1 (PRJNA80705) and R26 (PRJNA178189),
E. meningoseptica ATCC 12535 (NITE) (PRJNA199489),
E. meningoseptica ATCC 12535 (OSU) (PRJNA198814), and
E. meningoseptica 502 (PRJNA176121). This led to the discovery of 16 antibiotic resistance genes from the core genomes and 19 antibiotic resistance genes from the accessory genomes of
Elizabethkingia spp., and this included genes that confer resistance to aminoglycosides, β-lactams, fluoroquinolones, glycopeptides, macrolide-lincosamide-streptogramins, tetracyclines, trimethoprim, and rifampicin [
40]. A later study on two African isolates, E27017 and E18064, that compared their genomes with the genomes of 18 strains belonging to the genus
Elizabethkingia from many different regions, including Malaysian and Singaporean genomic sequences that were available at that time, identified that all
Elizabethkingia genomes contained at least 17 antimicrobial resistance genes [
41].
A whole-genome sequencing study on three isolates of
E. meningoseptica collected from an outbreak from three separate patients living in different counties in the Midwest regions of Michigan led to the identification of 22 resistance genes and 18 multidrug resistance efflux pump-encoded genes in all samples [
59]. While
Elizabethkingia spp. genomes shared many antibiotic-resistance genes with each other, minor differences have been reported [
3,
60]. Hence, genomic investigations of
Elizabethkingia spp. offers invaluable novel information on the species, but unfortunately, there have not been any reports of the whole-genome sequence of
Elizabethkingia spp. isolates from Southeast Asia besides those from Singapore.
4. Virulence Factors
The mechanisms of pathogenesis of
Elizabethkingia spp. are still being studied [
59]. When the virulence factor database (VFDB,
http://www.mgc.ac.cn/VFs/, accessed on 12 December 2021) was used to predict their presence from the genome sequences of various
Elizabethkingia spp., this led to the prediction of a total of 270 putative virulence factor genes. More than fourteen virulence factor classes for
Elizabethkingia spp. were identified with the following defined virulence-associated functions: adherence, antimicrobial activity, biofilm, cellular metabolism, effector delivery system, exoenzyme, exotoxin, immune modulation, invasion, motility, nutritional/metabolic factor, post-translational modification, regulation, stress survival, and others. Different species of
Elizabethkingia shared the same virulence factors (
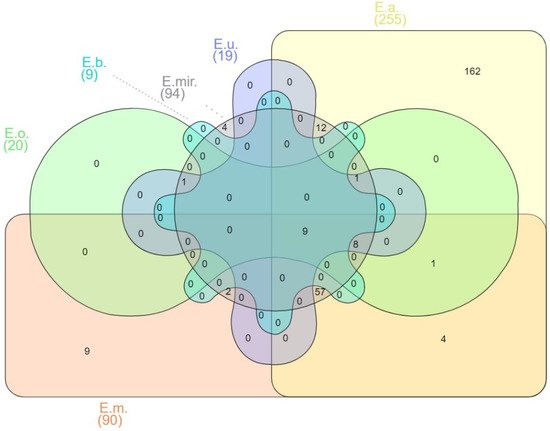
Figure 1).
Figure 1. Venn diagram of shared virulence factor genes of
Elizabethkingia spp. E.m.—
E. meningoseptica; E.a.—
E. anophelis; E.mir.
—E. miricola; E.o.
—E. occulta; E.u.—
E. ursingii; E.b.—
E. bruuniana. Edwards mode was used to process virulence factor gene outputs for Venn diagram visualization with InteractiVenn [
74].
Among the 270 predicted genes for virulence factors, 162 have been reported as unique in E. anophelis. E. meningoseptica carried six unique genes involved in adherence that encode curli nucleator protein (csgB), curli assembly proteins (curEm1, curEm2, curEm3, curEm4), a curli production assembly protein (csgG), and two genes involved in immune modulation encoding a capsular polysaccharide synthesis enzyme (cap8O), a gene encoding Rab2-interacting conserved protein A (ricA) and a putative carbonic anhydrase-encoded gene (mig-5). Four of the E. miricola unique virulence genes were predicted to be involved with urease accessory protein (ureE), urease alpha subunit (ureA), twitching motility protein (pilG), and sphingomyelinase-c (smcL).
Identification of 6880 gene families in
E. anophelis highlighted the genomic heterogeneity of
Elizabethkingia species [
41]. Genes homologous to heme iron acquisition, oxidative stress resistance proteins, and hemolysins were reported in earlier studies [
34,
61,
62]. Extensive variations of capsular polysaccharide synthesis genes in
E. anopheles were first reported by Breurec, Criscuolo [
41], with variable
cps clusters observed amongst the different lineages suggesting virulence heterogeneity among
Elizabethkingia strains [
41]. Identification of the capsule biosynthesis gene,
capD [
59], and the
adeG gene for the AdeFGH efflux pump [
20] in all
Elizabethkingia species leads to possible biofilm formation [
44,
63], which empowers the bacteria with the ability to persist on various surfaces [
59,
64]. Thirty clinical isolates from Malaysia, which comprised
E. anophelis,
E. meningoseptica, and
E. miricola, were recently shown to produce biofilms on polystyrene microtiter plates [
65].
Nine virulence factor genes were shared between six of the
Elizabethkingia spp., including the
E. argenteiflava-encoded
adeFGH efflux pump, isocitrate lyase (
icl), catalase/(hydro)peroxidase (
katA), 60K heat shock protein (
htpB), phospholipase C (
plc), phosphopyruvate hydratase (
eno), translation elongation factor (
tuf), catalase/peroxidase HPI (
katG), and aspartate 1-decarboxylase precursor (
panD), which is involved with adherence, biofilm formation, cellular metabolism, exotoxin production, and stress survival. Isocitrate lyase (
icl) plays an important role in the glyoxylate cycle [
66], and its presence in
Elizabethkingia spp. can predict its essential role in stationary-phase survival. An early report had shown that the presence of
icl in
Mycobacterium tuberculosis promoted the tenacity of infection by helping the pathogen to survive inside macrophages [
67].
However, the specific role of bacterial enzymes in pathogenesis varies with infection. The presence of phospholipases C (
plc) in all
Elizabethkingia spp. [
46] suggest its crucial role in downregulating host immunity [
68]. In
L. monocytogenes,
plc aided bacterial escape toward the cytosol and cell-to-cell propagation, whereas, in
C. perfringens, it helped bacteria induce endothelial damage and platelet aggregation, and in
P. aeruginosa, it led to the triggering of signaling pathways that lead to inflammation [
69].
The catalase-peroxidase genes,
katA and
katG (encoding hydroperoxidase I), are crucial against oxidative stress [
70]. An earlier report showed that strains with
katA were resistant to dodecyl sulfate, proteinase K, pepsin, trypsin, chymotrypsin, and the neutrophil protease cathepsin G, and they could survive for a long period once released from lysed cells [
71]. Presence of
katA [
40,
41,
44,
46,
60,
61,
72,
73] and
katG [
44,
46,
72,
73] could also support
Elizabethkingia species’ resistance to aminoglycosides.
5. Sources of Isolation and Transmission
The genera
Elizabethkingia are aerobic, non-fermenting, non-motile, catalase-positive, oxidase-positive, indole-positive, and Gram-negative bacilli widely distributed in soil, mosquitoes, plants, fresh and marine fish [
30,
75], food products [
76], hospital settings [
77], stagnant water, inland wetlands, and rivers [
33]. Due to their biofilm-forming ability [
63], they have been isolated from sinks and taps where they colonize the most, leading to nosocomial and community infections [
78] (
Table 1).
Table 1. Various sources of isolation of Elizabethkingia spp. in Southeast Asia.
Vector-borne transmission of the bacterial pathogen via mosquito bites has been suggested ever since the discovery of
E. anophelis in the midgut of the
Anopheles gambiae mosquito [
79,
80] and, more recently, in the salivary glands and saliva of
Aedes albopictus [
81]. The microbiome of
Anopheles mosquitoes has evidently revealed the strong symbiotic nature of
E. meningoseptica [
82], which has been isolated from various independent sources, including
Anopheles stephensi, the vector for the malarial parasite
Plasmodium vivax [
78,
83,
84], semi-field
Anopheles gambiae females [
82,
84,
85,
86], field sampled mosquitoes in Cameroon [
87,
88], and laboratory-reared mosquitoes where
Anopheles were the predominant species [
87,
89]. Another comparative study on bacterial microbiota isolated from the midgut of various
Anopheles spp., which were obtained in the same region of Mae Sot District and Sop Moei District in Thailand, reported on the findings of
Elizabethkingia spp. from
Anopheles minimus,
Anopheles dirus, Anopheles maculatus, Anopheles sawadwongporni, and
Anopheles dravidicus mosquitoes [
90]. However, sequences associated with the genus
Elizabethkingia could not be definitively assigned to either
E. anophelis or
E. meningoseptica as the V3–V4 region of the 16S rRNA gene used for microbiome profiling could not differentiate between the two species [
90]. Despite these multiple discoveries of
Elizabethkingia spp. in the midgut and salivary glands of various mosquito species, there is currently a lack of strong direct evidence that supports
Elizabethkingia infection, particularly
E. anophelis, as a mosquito-borne disease [
45], although this should not be ruled out with the current level of knowledge. A comparative genomics study of three cases of
E. anophelis also provided evidence of vertical transmission from mother to her baby [
62].
Zainuri et al. (2013) reported on the isolation of
E. meningoseptica from American bullfrogs (
Lithobates catesbeianus or
Rana catesbeiana) suffering from red leg syndrome and cataract in Sabah, Malaysia [
106]. Isolation of
E. meningoseptica from bullfrogs was also described in an earlier study, in which the isolates obtained were found to be resistant to multiple antibiotics [
107].
E. miricola, which had been implicated in acute infections in humans, caused a disease outbreak associated involving the internal organs of different anuran species, including northern leopard frogs (
Lithobates pipiens), Chapa bug-eyed frogs (
Theloderma bicolor), and Vietnamese warty toads (
Bombina microdeladigitora) captured in Vietnam. The presence of β-lactamases and putative virulence genes in the
E. miricola isolates were detected in silico [
27].
E. miricola was also reportedly isolated from Tra catfish (
Pangasius hypophthalmus) fillets in the industrial processing lines in Vietnam [
109]. Tra catfish is a type of freshwater fish, which is one of the major fish species in the Mekong River, and its processed fillets are exported to more than 80 different countries worldwide [
28]. Other scientists have also reported the isolation of
E. meningoseptica from retail sausages in Kampar, Malaysia, although the identification was performed by traditional biochemical methods and identified as
Chryseobacterium meningosepticum [
76].
Furthermore, 454 pyrosequencing of the 16S rRNA gene from the bacterial community of the root of the gnetalean gymnosperm
Gnetum gnemon and nearby bulk soils of a tropical forest arboretum at the Forest Research Institute of Malaysia (FRIM) at Kepong, near Kuala Lumpur, identified the mutualistic presence of
E. meningoseptica and
E. miricola [
110].
Elizabethkingia spp. was surprisingly found in relative abundance (4.9%) on the leaves of
Gnetum gnemon in comparison with rhizoplane (1.4%) [
111]. These reports indicate the ubiquity of
Elizabethkingia spp. in the environment and, thus, the difficulty in tracing an outbreak should one occur in the community and outside of hospital settings.