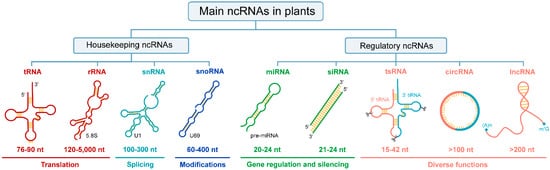
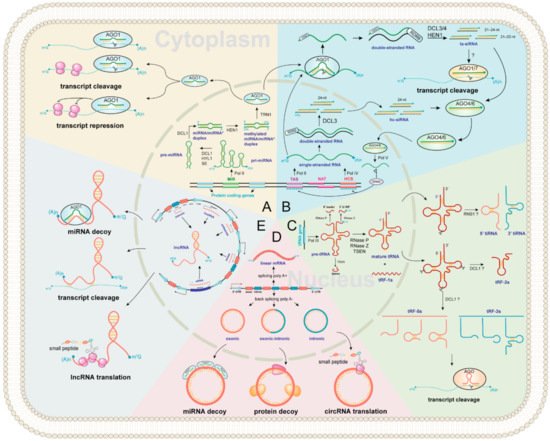
With a broad size range of 15–42 nt, tsRNAs are a new category of regulatory ncRNAs, which are classified into five categories according to the cleavage sites, namely tRF-1s, tRF-2s, tRF-3s, tRF-5s, and tiRNA (
Figure 2C). However, the study in plants has just started, and many questions remain to be answered. For example, the biogenesis pathway of tsRNAs in plants is still unclear, and the physiological function of certain tsRNA in plants is currently very limited [
68]. In this review, we propose a hypothesis of tsRNAs biogenesis in plants based on previous studies. First, RNA poly III transcribes tRNA gene as precursor tRNA (pre-tRNA) [
69], which includes a 5′ leader, a mature tRNA backbone, a 3′ U trailer, and sometimes an intron [
70]. Then, the 5′ leader, 3′ U trailer, and intronic sequences are cleaved by RNase P, RNase Z, and tRNA-splicing endonucleases (TSEN) to produce mature tRNA and tRF-1s (tRF-1s could be derived from 3′-end of pre-tRNA) [
71,
72,
73]. The mature tRNA (73–90 nt) forms a secondary cloverleaf structure with a D-loop (left), a T-loop (right), anticodon loop (bottom), a variable loop, and an acceptor stem (
Figure 2C). Finally, the mature tRNAs could be cleavaged by Arabidopsis S-like Ribonuclease 1 (RNS1) and/or DCL1 to form tRF-2s, tRF-3s, tRF-5s, and tiRNA (
Figure 2C) [
30,
31]. In mammals, tsRNAs incorporate into silencing AGO and trigger RNA interference [
74]. Likewise, AGO-associated tsRNAs have been predicted in
Arabidopsis and rice [
30,
75]. An in vitro assay has shown that certain tsRNAs regulate gene expression by translation inhibition, and tsRNA-AGO1 complex tends to target transposable element transcripts and probably maintains genome stability [
31,
76,
77].