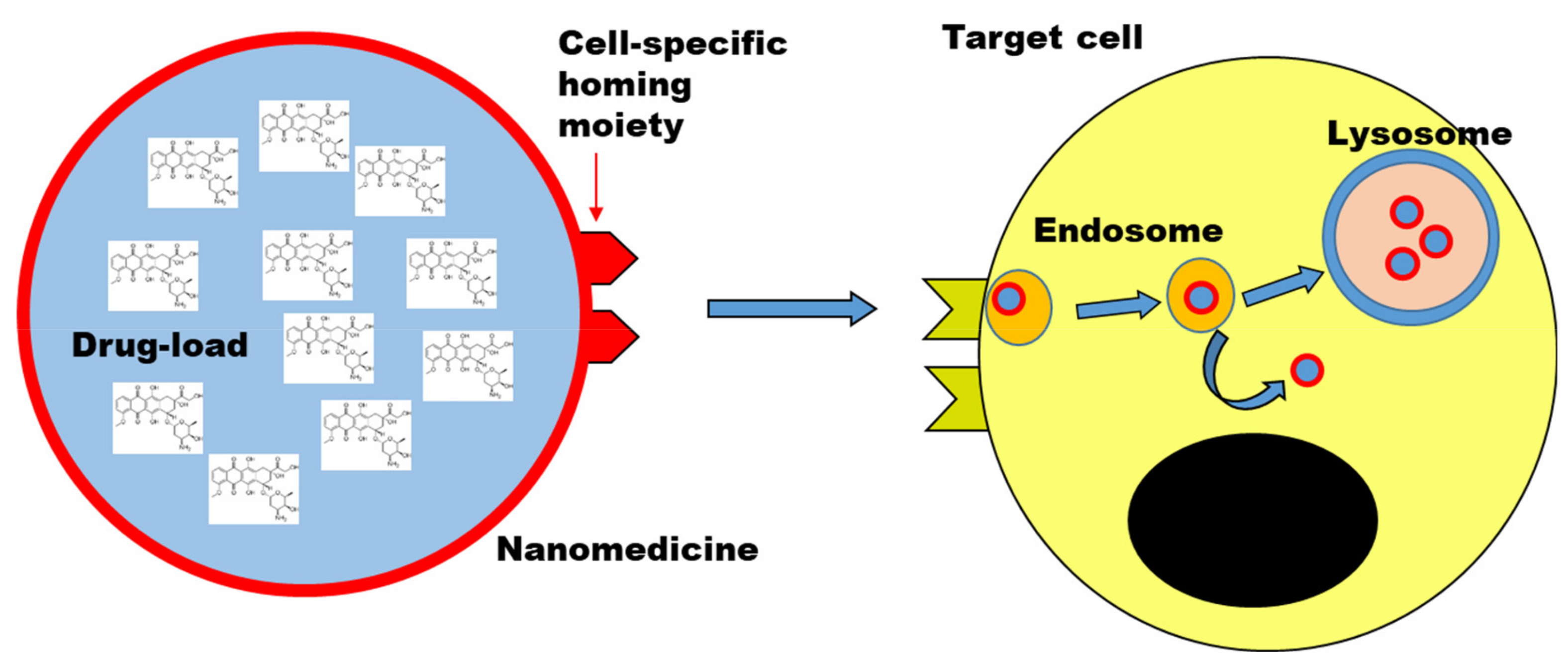
3.2. Interactions with Cells and Intracellular Fate
After administration and modifications in biological fluids, the next steps into the journey of nanomedicines and targeted drugs are the interactions at the cell membrane both of the targeted cells and all other cells, and the following uptake mechanism and intracellular trafficking.
While we discussed above examples of drug targeting strategies and effects of corona formation on targeting and interactions with cell receptors, there are other key aspects in the interactions at the cell membrane of nanomedicines and targeted drugs which need to be better understood and controlled in order to improve their efficacy.
One important aspect that has emerged is that even when specific receptors are targeted (via active targeting ligands or via the corona adsorbing on nanomedicines), the targeted drug or nanomedicine may be internalized and processed by cells in different ways comparing to what usually observed for the same receptors and their natural ligands [
161]. For instance, we found that even when the LDLR mediates nanoparticle uptake, the mechanisms of internalization is not clathrin mediated, as usually observed for this receptor [
148]. Similarly, nanoparticles targeting the transferrin receptor were not recycled and exported by cells as transferrin, but were trafficked by cells towards the lysosomes [
139]. More efforts are needed to understand how after interactions with certain receptors, cells internalize and process these materials. Without this knowledge, the development of nanomedicines and targeted drugs risks to remain mainly limited to screening of multiple materials, as opposed to the tailored design of materials with the properties required to achieve the desired outcomes at cell level (as well as in vivo).
The mechanisms cells use for the internalization of nanomedicine and targeted constructs also affect uptake efficiency and intracellular trafficking kinetics [
134,
161]. These factors affect the time required to reach the therapeutic dose intracellularly, and the time required to reach specific locations inside cells, which all together contribute to the final therapeutic efficacy.
Characterizing the mechanisms by which nanomedicines and targeted drugs are internalized by cells remains highly challenging [
134,
161,
162]. Many contrasting results are often reported and it is hard to draw conclusions on how nanoparticle properties affect the mechanism of uptake or how they vary among different cell types. This is also due to complicating factors such as corona formation, which are not always controlled and reproduced among different studies. Additionally, the methods usually applied to try to characterize these mechanisms are often limited [
134,
162]. We have shown examples of this for classic transport inhibitors commonly used for this purpose [
163]. For instance we found that when using high concentration of inhibitors such as chlorpromazine, a common drug to block clathrin-mediated endocytosis, reduced nanoparticle uptake was observed but this was due to strong toxicity on cells, as opposed to a specific inhibition of the pathway. Additionally, many of these compounds lost their efficacy when applied to cells in a medium with serum, probably due to protein binding [
163]. Given the effect of corona formation on targeting and nanomedicine interactions with cells, thus the need to include serum when testing nanomedicine and targeted drugs in vitro, other compounds whose efficacy is not affected by the presence of serum need to be used [
161,
163]. Thus, without controls for similar effects and to exclude toxicity, results obtained with these common transport inhibitors may lead to wrong conclusions. Given these limits, multiple methods need to be combined. Furthermore, we cannot exclude that nano-sized materials may be processed in different ways than natural ligands and alternative endocytic mechanisms not yet characterized may be triggered by these special cargoes. Thus, new methods need to be developed to identify potential novel targets not yet associated to cell uptake mechanisms. To this aim, we have applied genome-wide screening and developed proteomic based approaches to identify all genes and proteins involved in nanoparticle uptake (Montizaan et al., unpublished; Garcia-Romeu et al., unpublished). Indeed, many novel targets have been identified thanks to this kind of approaches and current efforts are focused on understanding their role.
Finally, it is commonly observed that after uptake most nanomedicines are trafficked by cells along the endo-lysosomal pathway towards the lysosomes [
133,
164]. While this can be exploited when these are the targeted intracellular compartments, all drugs which require accumulation into other locations inside cells require to escape the endo-lysosomal compartments or are otherwise degraded once they are transported in the lysosomes. Several strategies for endosomal escape have been investigated, and often are limited by toxicity and poor efficacy [
165,
166]. In the case of RNA delivery and recently approved lipid nanoparticles such as Onpattro or the RNA vaccines against COVID-19, this has been achieved utilizing ionizable lipids which become positively charged only once inside the slightly acidic environment of the endosome [
11,
12]. The acquired cationic charge can destabilize the endosomal membrane and promote escape of the nanomedicine into the cytosol. This could be developed by carefully tuning lipid design to change their pKa in order to promote this effect.
This is another example which beautifully illustrates the importance of understanding how nanomedicines and targeted drugs are processed by cells so that their design can be carefully tailored in order to achieve the desired effects.
In order to characterize the intracellular trafficking of nanomedicines and targeted drugs, methods to be able to follow them inside cells over time and determine their intracellular location as well as the time needed to reach certain location are needed. For instance, we have used live cell fluorescence microscopy imaging to measure nanoparticle location inside cells over time and in particular their colocalization with the lysosomes [
167]. Using nanoparticles of different sizes and theoretical modeling, the intracellular trafficking kinetics could be determined, thus how nanoparticle size affect the departure time from the cell membrane and arrival time to the lysosomes. With similar methods, we can then determine how nanoparticle properties affect these details of intracellular trafficking, which also affect nanomedicine efficacy. Our results also showed that within the same cells some nanoparticles arrive to lysosomes within short times, while others take much longer or get stuck somewhere else and never seem to arrive there. Additionally, we also found that not only uptake efficiency, but also intracellular trafficking kinetics strongly vary within individual cells in a cell population [
167]. Clearly, further studies are needed to understand similar observations and learn how to design nanomedicines and targeted drugs with controlled and uniform behavior at cell level. Within this context, in order to try to understand the sources of such heterogeneity in nanoparticle uptake by cells, we recently used repeated cell sorting to isolate cells with low and high nanoparticle uptake within a cell population and used transcriptomics to identify differences in gene expression in such sub-populations [
168].
Next to such imaging-based approaches, taking advantage of latest developments in high sensitivity flow cytometry, we have also developed a method based on organelle flow cytometry to gain temporal as well as spatial information on nanoparticle intracellular distribution and trafficking [
169]. The method allows to gain information on nanoparticle intracellular location as usually obtaining by imaging-based methods but with the high throughput and robust quantification enabled by flow cytometry. Thus, cells are exposed to nanoparticles, then all organelles are extracted and characterized by flow cytometry. In this way, intracellular trafficking kinetics can be easily obtained from thousands of organelles extracted from thousands of cells, without the complex analysis required to extract similar results from imaging-based methods. The two different methods present different advantages and limits, thus they can be used to complement each other in order to gain a better understanding on the details of intracellular trafficking of these object by cells. A better understanding of how cells process nanomedicines and targeted drugs is needed in order to optimize their design and efficacy.