3.1. Genomic Changes through Domestication
Generally, domestication is a selection process that provides the increased adaptation and economic viability of the plants to be cultivated in a particular environmental condition [
27]. It is assumed that the first domestication of crop species started by humans 10,000 years ago [
22,
28]. The initial selection of wild plants as potential crops was the first step in the foundation of agriculture. However, plant selection under domestication is being continued since the Neolithic period through to plant breeding of today [
29]. Through the domestication process, plants go through a suite of complex morphological, physiological, and genetic changes [
30].
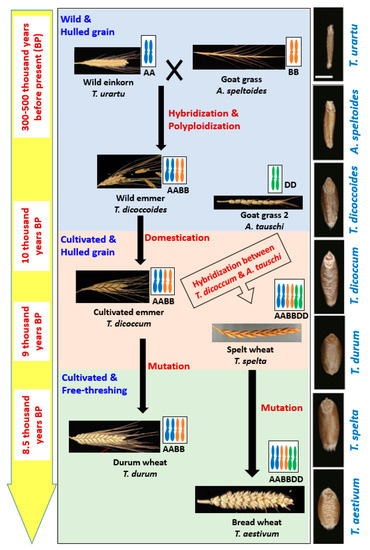
According to the history of wheat evolution, only wild einkorn and WEM wheats went through the early domestication selection [
1]. Einkorn,
T. monococcum is a diploid wheat, which was domesticated from the wild progenitor species
T. boeoticum in the Fertile Crescent. Later on, it was gradually replaced by tetraploid and hexaploid wheats during the last 5000 years, approximately. Einkorn has never been involved in the evolution of hexaploid bread wheat or tetraploid durum wheat. The wild diploid
Triticum species, which was the progenitor of hexaploid wheat and played an essential role in wheat evolution in
T. urartu (AA) [
28]. The tetraploid wheat species
T. dicoccoides known as WEM naturally had been grown all over the Fertile Crescent. The early wheat growers domesticated WEM, and thus, cultivated emmer (
T. dicoccum, AABB) was introduced. For a millennium or more since its domestication, emmer wheat was still growing with WEM in a complex cropping system in many Levantine sites. Thus, the genes (for example, non-brittleness gene) were transferred through spontaneous and uncontrolled hybridizations. As a result, the domesticated emmer wheat has appeared as polymorphic populations [
22].
Generally, domestication aims with the elimination of undesired or even deleterious alleles, but almost in every case it also reflects an erosion of alleles valuable for plant improvement and future demands of producers and consumers [
31]. It has been well documented that substantial genetic erosion occurred through the domestication process of wheat and that erosion was further reinforced during modern breeding processes [
32,
33,
34]. Consequently, loss of diversity, selective sweeps and adaptive diversification have occurred that caused considerable genetic modification [
27]. The development in molecular marker and quantitative trait loci (QTL) analysis techniques enabled to characterize those genetic losses or modifications. For example, nucleotide diversity at 21 gene loci was analyzed in wild, domesticated, cultivated durum and bread wheats, and revealed that diversity was reduced in cultivated forms during domestication by 69% in bread wheat and 84% in durum wheat [
35].
The most significant effect of domestication is that genetic diversity has been reduced and is being continued through the modern plant breeding. The occurrence of genetic narrow down essentially has reduced the efficiency of crop improvement [
36]. Wheat domestication by the early farmers eventually resulted in landrace cultivars (LCs) adapted to specific conditions of their habitats. At the advancement of modern plant breeding during the last century, most of the traditional LCs were continually replaced by modern wheat cultivars (MWCs) [
37]. As the MWCs were bred from a few LCs they contain less genetic diversity than traditional LCs [
36]. Growing wheat of such a narrow genetic diversity accelerates the risk of genetic vulnerability to the adverse condition. The risk has been further raised up due to the spontaneous mutations of a number of major insect and pathogens and the impulsive changes in environmental conditions. These might bring stresses in a new dimension that the present wheat cultivars could not cope with, and therefore, could lead to severe crop losses. During the second half of the last century number of such kind of severe crop loss had been evident. For examples, severe epidemics of shoot fly (
Atherigona spp.) and kernel bunt (
Tilletia indica); the outbreak of the southern corn leaf blight in the 1970s, and more recently, the outbreak of wheat blast in Bangladesh and northern India [
36,
38].
The genetic basis of the domestication syndrome in wheat has been extensively studied which revealed that the loss of genetic diversity in spring bread wheat occurred during (i) its domestication, (ii) the change from traditional landrace cultivars (LCs) to modern breeding varieties, and (iii) 50 years of international breeding [
36]. Considerable loss of genetic diversity was observed at the early periodic domestication, and during the time of LCs to the elite breeding germplasm. It has also been evident that wheat’s genetic diversity was narrowed down more robustly during the time between 1950 and 1989. However, genetic diversity showed an uprising trend starting from 1990 indicating that breeders have experienced the consequence of narrowing down genetic diversity in the modern breeding and subsequently started to increase the genetic diversity through the introgression of novel materials. The LCs and
T. tauschii contain numerous unique alleles that were absent in modern spring bread wheat cultivars [
36].
It has been considered that, at the very beginning of the domestication process, the major domestication trait was the seed dispersal mode [
8]. Certainly plants with reduced spikelet shattering at maturity had been domesticated, which was considered as a key feature in preventing natural yield losses [
8]. In addition to the yield, the other major domestication-related traits include glume reduction (easier threshing), plant architecture (plant height, tiller numbers etc.), ear and kernel size, seed dormancy [
39]. Later on, with the improvement of grain analytical process, the grain protein and mineral concentrations, as well as carbohydrate content, also became major selection attributes (). Domestication has genetically not only transformed the brittle rachis, tenacious glume and non-free threshability, but also modified numbers of other traits [
8]. Meanwhile, breeding and selection had a different impact on different wheat genomes. For example, a greater number of genes related to those domestication traits are found on the A and B genomes [
40]. Differential loss has been found that supports greater gene loss in the A and B genomes compared with the D [
41].
Table 1. Important traits were considered for domestication and the responsible genes or QTLs with their location.
Reduction in diversity caused by intensive selection can be counterbalanced by introgression of novel germplasm. The best strategy for wheat improvement is to utilize the adaptive genetic resources of the wild progenitors, wild emmer (WEM,
T. dicoccoides) and other wheat relatives [
15,
34,
49,
50].
3.2. Genomic Changes through Polyploidization
During the polyploidization of wheat, rapid alteration and several genomic changes occurred in nature. Such phenomena can be divided into two groups: (i) Revolutionary changes, and (ii) evolutionary changes. Revolutionary changes took place rapidly, during or just immediate after allopolyploidization and within a few generations, whereas evolutionary changes happened throughout the evolutionary lane for hundreds to thousands of generations and accelerated by polyploidy [
10]. These changes can be of various types, including the elimination of both low-copy and high-copy DNA sequences, intergenomic disruption of DNA sequences, DNA methylation, deletion of rRNA, gene loss, suppression or activation of gene, chromatin modification and remodeling, heterochromatinization, sub-functionalization, and neo-functionalization [
51]. These changes are directly or indirectly influenced by allopolyploidization. Besides, hybridizations that occurred during the evolution of wheat also resulted in some significant genetic changes as this is a very common outcome of the process. For examples, in the crossing product (hybrid) of
Aegilops sharonensis and
Aegilops umbellulate, 14% loci from
Ae. Sharonensis and 0.5% loci from
Ae. Umbellulate were lost; whereas, in the case of a cross between
Ae. Sharonensis and
T. monococcum, many sequences from
T. monococcum were doubled in hybrid compared to another parent [
52]. However, it is evident that evolution results in several genomic changes. Some examples are given below in more details.
Nucleolus organizing regions (NORs), also named as ribosomal DNA (rDNA) loci are present on different chromosomes (1A, 5A, 1B, 6B, and 5D) of diploid wheat [
53]. This gene is composed of long tandem repeats that clustered on the chromosome and translated into important components of the chromosome [
7]. Its activity is associated with the size of the intergenic regulatory region and the status of cytosine methylation [
54,
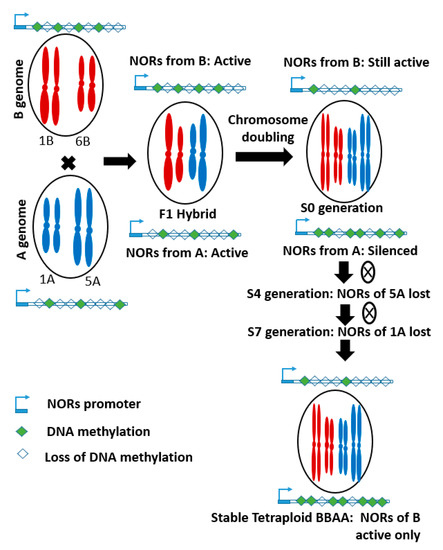
55]. However, NORs from the A genome are largely lost during the evolution of synthetic tetraploid wheat, due to asymmetric transcription and epigenetic modifications during polyploidization (). In hybrids, NORs from both parents were expressed. However, after chromosome doubling, it became silenced in one parent (A genome), due to increased DNA methylation. In this process, a pair of NOR on the 5A chromosome were deleted first, the gradual elimination of another pair from the 1A chromosome, resulted in complete loss of NORs from the A genome by S7 generation. Therefore, in stable synthetic tetraploid wheat, rDNA from only the B genome was present. In the case of bread wheat, the rDNA loci form both A and D genomes were largely eliminated during evolution [
7]. Additionally, genome wide transcription analysis revealed that gene expression in synthetic bread wheat is parentally dominant and only one of the parental genomes determines morphological traits and ecological adaptations [
46,
51,
54,
56].
Figure 2. Schematic diagram showing the loss of Nucleolus organizing regions (NORs) from A genome, due to increased DNA methylation during the evolution process. S4 and S7 mean fourth and seventh generation of selfing (adopted from Guo and Han (2014) [
7]).
Chromosome-specific sequences (CSSs) occur in only one homologous chromosome pair, i.e., 1A and 1A. These types of sequences were present in all diploid species. However, after polyploidization, CSSs from one genome were eliminated immediately or after some generations. As a consequence, in hexaploid or tetraploid, these sequences occur only in one homologous pair but absent from the homeologous chromosome [
57]. Meanwhile, allopolyploidization results in rapid non-random deletion of specific non-coding, low-copy and high-copy DNA [
58]. Again, sometimes some genes of the A and B genomes get suppressed upon adding of D genome. As results, they expressed in tetraploid AABB genome, but not in hexaploid AABBDD genome [
10]. This is called intergenomic suppression. For example, the rust resistance gene(s) present in the A or B genome was suppressed by a gene present in the long arm of chromosome 7D [
59].
3.3. Genomic Changes through Natural Mutation
Through the centuries, natural mutation resulted in significant changes in the genomic structures of wheat, which contributed substantially in the genetic evolutionary process of wheat. In general, mutation generated new alleles, while recombination created novel allele combinations. Accumulation of new mutations in older polyploid species, such as WEM, results in increased diversity and more uniform distribution across the genome [
60]. For example, Genetic studies revealed that two recessive alleles at two major loci (
Br-A1 and
Br-B1) controlling non-brittle rachis raised through mutation during domestication [
61]. One of the most important genomic changes is the evolution of free-threshing wheat as a result of several major and minor mutation events. A single major gene
Q on chromosome 5AL is responsible for free-threshing of modern bread and durum wheat, whereas the recessive
q allele is for non-free-threshing wild wheats [
1]. A recent study showed that the
Q allele arose from
q allele through a gain of function mutation [
62]. Free-threshabiliy is also related with tenacious glume (
Tg) gene, because
Tg inhibits the expression of
Q gene. QTL correspond to the
Tg gene is located on 2D and 2B chromosome. Free-threshing phenotype evolved when mutation transformed
Tg into
tg. Therefore free-threshing common bread wheat (
QQ5Atgtg2Btgtg2D) and free-threshing durum wheat (
QQ5Atgtg2B) have mutant alleles at each of the important threshability loci [
2,
17].