We analyze further extension of G-quadruplexes by additional structural elements and investigate whether junction of G-quadruplex with duplex, hairpin, triplex or second G-quadruplex motif is favorable for aptamers stability and biological activity. Furthermore, we indicate the specific and pivotal role of G-quadruplex domain and the additional structural elements in the interactions with target molecules. Finally, we consider the potency of G-quadruplex junctions in the future applications and indicate the emerging research area that is still waiting for development to obtain highly specific and effective nucleic acid-based molecular tools.
- G-quadruplex
- G-quadruplex junction
- duplex
- hairpin
- triplex
- aptamer
- antithrombin aptamer
- anticoagulants
- TBA
1. Introduction
Aptamers are single-stranded oligonucleotides with specified sequences which determine the adoption of a strictly defined spatial structure. This enables highly specific binding to target molecules and modulation of their activity [1]. The origin of the application of nucleic acids as high-affinity ligands comes from the independent works of L. Gold and J. Szostak, where systematic evolution of ligands by exponential enrichment (SELEX) has been discussed for the first time [2][3]. This main method of aptamers development relies on the selection of oligonucleotides with the greatest affinity towards a target compound from the combinatorial library as a result of repeatable cycles of binding reactions and washing off unbound molecules [4]. The discovery of aptamers has revolutionized the world of science due to their interesting properties such as high specificity, favorable dissociation constant, and relatively facile chemical synthesis [1][4][5][6].
A significant proportion of aptamers constitute G-quadruplexes presumably due to a greater negative charge density, which is twice as for double-stranded DNA, and which could influence the enhancement of interactions with proteins [7]. What is more, G-quadruplexes constitute a structurally diverse group of compounds, what results in almost unlimited possibilities of selecting aptamers towards a great repertoire of targets. All structural arrangements of G-quadruplexes possess common feature i.e., the presence of G-tetrads formed by Hoogsteen hydrogen-bonded guanines and stabilized by cations such as K+ [8] or Na+ [9]. Two or more G-tetrads stack upon each other forming a G-quadruplexes core [9][10] connected by several types of loops, which can be divided into propeller, lateral, and diagonal [11]. Depending on the arrangement of strands within the G-quadruplexes core, three different categories of folding topologies can be distinguished: parallel (all strands have the same orientation), antiparallel (two pairs of contrary oriented strands), and hybrid (three strands have the same orientation, one strand is aligned oppositely) [10][12]. Furthermore, based on the number of strands, G-quadruplexes can be further divided into intramolecular (formed by one strand) or intermolecular (multiple strands) [10]. In order to improve aptamers biological and physicochemical properties, a great number of modifications have been introduced. These alterations concerned the substitution of nucleotides with modified residues [13][14][15][16][17][18][19][20][21][22][23], changes to the phosphodiester bond [24][25][26][27], or structural variation [28][29][30][31][32][33].
Herein, structural engineering of G-rich aptamers by the junction with various structural motifs will be discussed. This interesting approach could provide improvements in aptamers affinity towards target compounds, multivalency of binding [34], or changes in the pharmacodynamic and pharmacokinetic properties [35]. The literature data collected in this review constitute a compendium of knowledge that might be potentially useful for the development of novel molecular tools or improvements of existing aptamers utility for therapeutic applications.
2. Quadruplex–Duplex Junction
The strategic positioning of both structural elements within quadruplex–duplex junctions was adopted to improve the properties of many G-quadruplex-based aptamers with a particular emphasis on anticoagulants.
One of the pioneering work in the field of quadruplex-duplex aptamers was described in 1995 by Macaya et al. [52]. Although they confirmed that TBA can be selected from their experiment, the vast majority of identified molecules contained additional 4-7 base pairs spanning both ends of G-quadruplex structure and showing higher affinity to target protein than the G-quadruplex motif alone. In 2011 Mazurov et al. characterized new antithrombin aptamer named RE31 [30]. The molecule consists of 15 nt long TBA G-quadruplex motif linked by two 2 nt spacers with 6 bp duplex domain. It shows improved affinity towards thrombin and significantly prolonged clotting time in comparison to TBA. The attempts of Kotkowiak et al. to characterize chemically modified variants of RE31 indicate no strict correlation between aptamers thermal stability and inhibitory activity of fibrin clotting [23]. Although authors noticed that the most thermally stable variants containing simultaneously LNAs in the duplex motif and single UNA within G-quadruplex domain show 2-fold improved inhibitory activity of fibrin clotting, the aptamer modified exclusively with LNAs possesses anticoagulant potential comparable to unmodified RE31 regardless almost 11°C higher melting temperature.
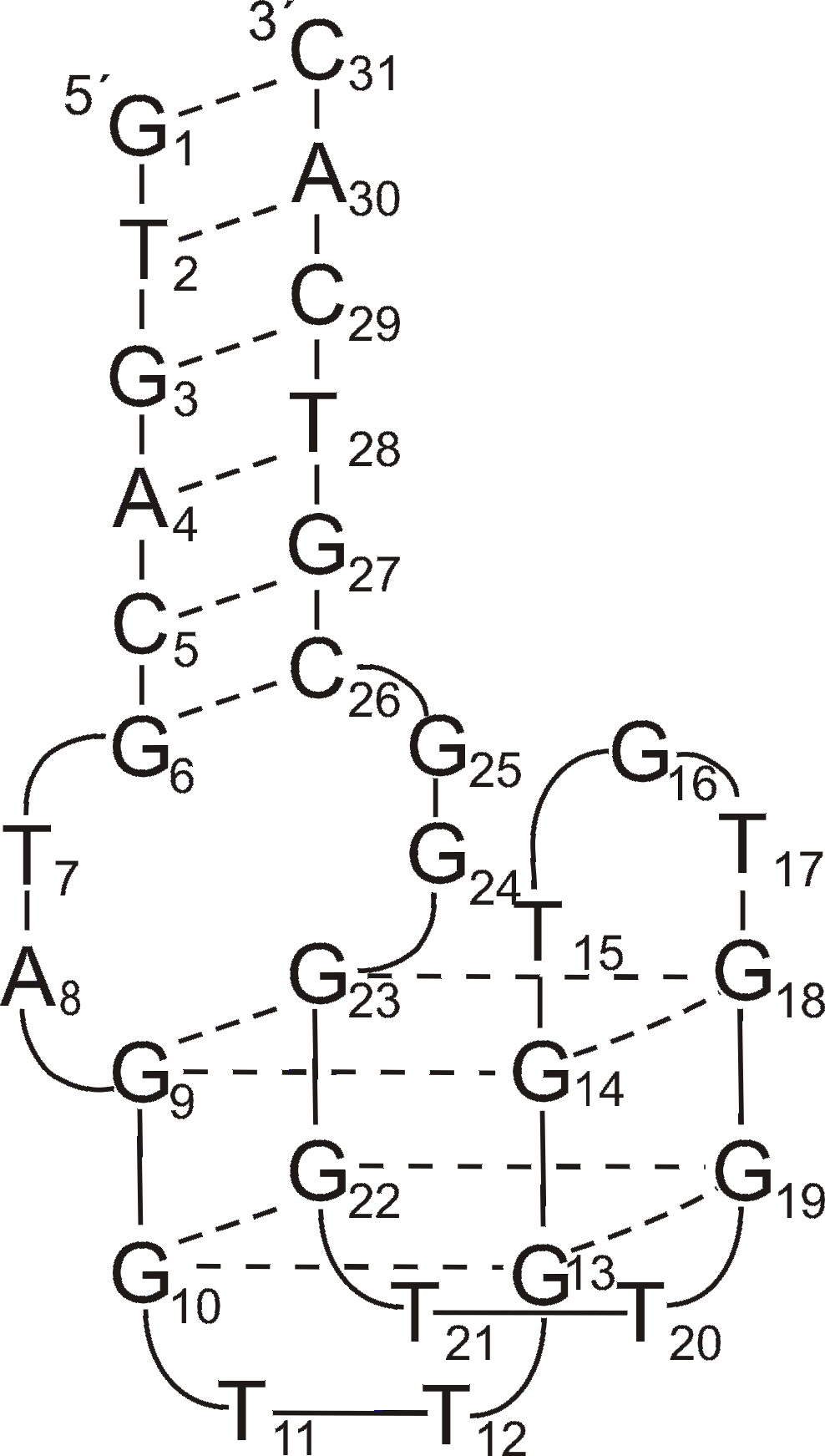
Figure 1. The structure of RE31 quadruplex-duplex junction [23].
Despite a large number of articles concerning anticoagulant quadruplex–duplex junctions, some G-quadruplexes structural functionalization with a helix domain has been described to obtain highly selective aptamers towards targets other than thrombin. Recently, a characteristics of novel quadruplex-duplex junction based on AS1411 aptamer was published [28]. The AS1411 is the most clinically advanced antiproliferative and pro-apoptotic aptamer, which forms highly polymorphic 26 nt long DNA G-quadruplex structure with the majority of topologies in parallel strand orientation [63, 64]. Interestingly, one of TBA junction (31TBA was used in development of novel aptasensors with dye-forming cavity between two domains [65]. The presence of duplex domain appeared favorable also for improvement of the properties of G‑quadruplex aptamers targeted towards hemin resulting in development of a new type of DNAzymes with peroxidase-like activity. The application of quadruplex-duplex junction in diagnostic platform for detecting 17β‑estradiol is also worth discussing.
Based on extensive literature data it can be concluded that vast majority of quadruplex-duplex junctions is of DNA type and mainly concerns aptamers targeted toward thrombin. The duplex domain undoubtedly plays important role in folding of G-quadruplex motif and in its activity. In general, the presence of helix very often decreases thermal stability of G-quadruplex, however simultaneously improves contact with surface of target protein. Furthermore, the length and sequence of duplex motif and linkers, which span both structural motifs, should be taken into account as essential for activity when designing this type of junctions.
3. Quadruplex–Quadruplex Junction
An interesting example of chemical alteration of G-quadruplex structures is the development of conjugates composed of two G-quadruplex particles, termed bivalent aptamers. The components can have different targets, but more frequently it is a combination of the compounds targeted towards different binding sites of the same molecule. This approach enhances the bivalency of conjugates what ensures higher binding affinity in comparison to parental compounds, which is a result of multiple binding events, a higher rebinding rate, and, above all, better target selectivity [34].
One of the first attempts of combining two G-quadruplexes was performed for thrombin binding aptamers [36]. In this research, a group of conjugates formed by 15 nt-long TBA covalently connected via linkers containing 5 to 10 units of spacer18 with a 29 nt-long compound, named 60-18(29) (HD22(29)) [37], was synthesized and characterized. As discussed in a previous paragraph, the 60-18(29) adopts a quadruplex–duplex structure and in contrast to TBA, binds to thrombin heparin-binding exosite [38]. The different binding sites of two G-quadruplex domains within conjugates resulted in the cooperativity of their action, which was reflected in the increased affinity towards thrombin [36]. The best variant, THR7, which is composed of TBA and 60-18(29) connected by a 10-unit-long linker, was the most efficient competitor to wild-type TBA and entirely inhibited its binding to thrombin at a 1:1 ratio. Based on the above, it was estimated that the Kd value of THR7 is below 97 pM, and its binding affinity is 65-fold higher than TBA. The detailed analysis of the data also revealed that the appliance of a longer linker (10 vs. 5 units) and specific order of components (TBA aptamer localized at the 5′-end of the conjugate) resulted in a more favorable affinity towards thrombin.
Further continuation of the research trend concerning complexation of HD1 with HD22(29) was performed by the Veedu group, who developed conjugates composed of the above aptamers linked by triethylene glycol or four 2′-deoxythymidines and with an inverted polarity of 2′-deoxythymidine at the 3′-end [39]. Both of them were characterized by improved anticoagulant properties in comparison to wild-type TBA, but RNV220 possessed a superior ability to inhibit thrombin activity. The presence of inverted pyrimidine nucleoside residue at the 3′-end of the oligonucleotide has a protective effect, what found reflection in the extended biostability of analyzed bivalent aptamers. What is more, the anticoagulant effect of RNV220 and RNV220T could be reversed entirely due to the addition of oligonucleotide with a sequence complementary to conjugates.
Apart from the covalent junction, the bivalency in aptamers constructs could also be obtained by non-covalent assembly [35][40]. In this arrangement, two HD1 particles were connected via locking of the G-quadruplex or helix lock, formed between 3′-terminal fragments of each component and stabilized by Hoogsteen and Watson–Crick base pairing, respectively. The data analysis revealed that although both conjugates showed reduced stability in thermodynamic studies in comparison to wild-type HD1 (T M for HD1, HD1-ds, T4T4 was 38.8 °C, 33 °C, and 35.6 °C, respectively), they were characterized by superior affinity toward thrombin (Kd for HD1, HD1-ds , T4T4 was 2.6, 1.5, and 0.62 nM, respectively). Interestingly, all tested molecules displayed a similar activity level in functional activity tests (thrombin time test), which was slightly lower than for HD1. Noteworthy, the comparison of anticoagulant properties of HD1-ds with RA-36 revealed the differences in their mode of action. RA-36 was characterized by a lower inhibitory activity than HD1-ds, therefore, it could be concluded that HD1 motifs in this conjugate act as one unit and that the presence of covalent linkage causes a decrease in anticoagulant properties. On the contrary, only a small variation in thrombin inhibition in comparison to HD1 was observed for HD1-ds. Thus, it could be assumed that its components functioned as two independent domains and their activity depends only on the accessibility of exosite I. The CD analysis indicated that both conjugates exhibited features of antiparallel G-quadruplexes. Additionally, the spectrum of HD1-ds possessed also a minimum at 245 nm, which is typical for DNA duplexes.
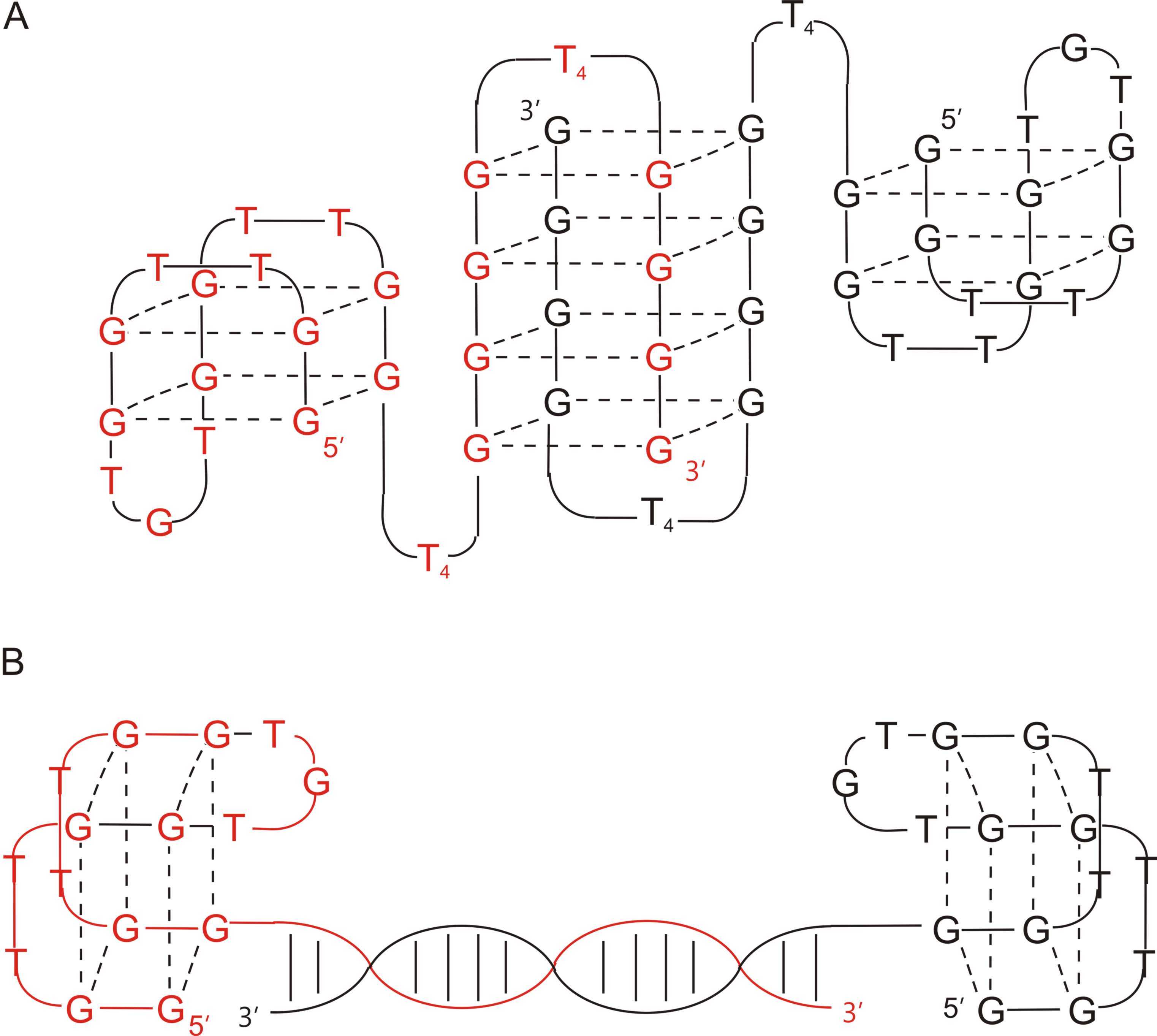
Figure 2. The structure of T4T4 (A) and HD1-ds (B) [35].
4. Quadruplex–Hairpin Junction
Another interesting type of combination of G-quadruplexes with different structural motifs is their junction with a hairpin. One of the first reports on this topic concerns a group of DNA aptamers, which are inhibitors of HIV-1 reverse transcriptase (RT), called RT5, RT6, and RT47 [41]. All conjugates have a similar structural pattern with a stem–loop structure at the 5′-end connected by a linker (two thymidines) to the G-quadruplex motif, built up with three G-tetrads. It was proven that both of the above structures, connected covalently, are required to obtain the inhibitory activity of the aptamer. Moreover, it was pointed out that not the sequence of the stem fragment but rather its length could have an influence on conjugate activity. Interestingly, the types of junctions with G-tetrads also did not affect the aptamer inhibitory properties, since the presence of closed-loop and unlinked 5′- or 3′-ends was equally efficient.
The second example of a G-quadruplex-hairpin junction constitutes the aptamer targeted towards hemin, being a DNAzyme with peroxidase-like activity [42]. The conjugate is formed by a G-quadruplex joined by unpaired 5 nt regions with a hairpin with a five base-paired stem and a 3 nt loop. This quadruplex–hairpin combination also shows improved affinity in comparison to the parental G-quadruplex and higher DNAzyme activity, which seems to be due not only to the presence of the stem–loop structure but also to the type of residues joining both domains.
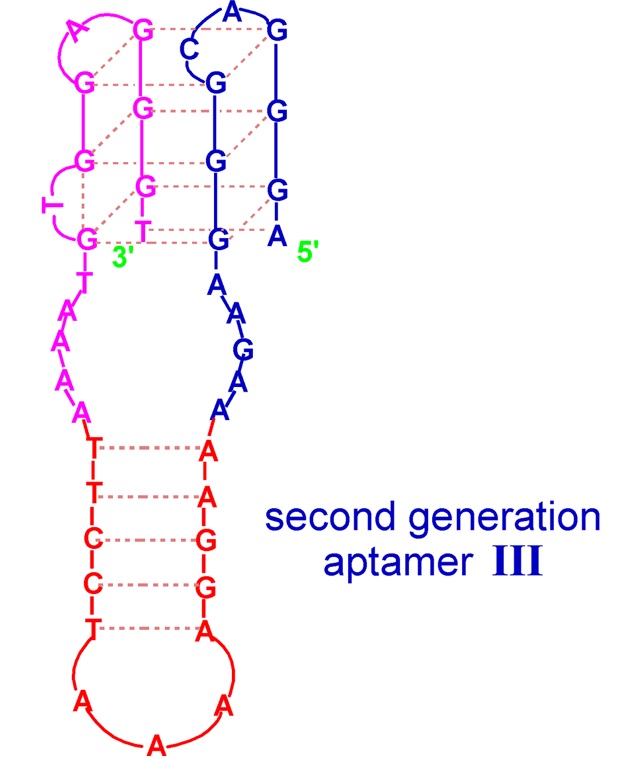
Figure 3. Schematic structure of quadruplex-hairpin junction with peroxidase-like activity [66]. Reproduced under the Creative Commons Attribution License.
A great supplement for the reports referring to aptamers with quadruplex–hairpin junctions is research concerning the structural analysis of model fragments representing genome sequences, which could provide some general information about the influence of a hairpin on the G-quadruplex structure. The Weisz group created quadruplex–hairpin conjugates based on the promoter region of c-Myc oncogene and a hairpin fragment localized at its 3′-termini or 5′-termini [43]. The CD and thermodynamic data analysis showed no influence of the presence of a stem–loop structure on general G-quadruplex folding topology and stability. Interestingly, it was observed that the stem–loop structure presented at the 5′-end of G-quadruplex was more thermodynamically stable (56.1 °C) than the same sequence localized at the 3′-end (45.1 °C). It was also proved that a quadruplex–hairpin junction can constitute a good binding site for ligands with G-quadruplex specificity, sometimes even more efficient than the G-tetrads itself like in case the of the indoloquinoline ligand.
5. Quadruplex-Triplex Junction
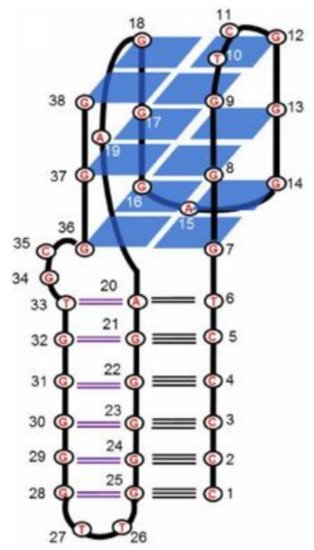
6. Conclusions
The promising physicochemical and biological characteristics of G-quadruplex junctions with other structural motifs studied so far provides preliminary evidences for favorable modification of G-rich structures. Undoubtedly, connection of G-quadruplexes with helices, hairpins, triplexes or other G-quadruplexes appeared favorable from the therapeutic point of view thus increasing repertoire of molecules potentially useful in therapy of human diseases. The doors for the development of new molecular tools based on nucleic acids are already opened. However, the area of research discussed herein still requires detailed studies of the requirements needed to design G‑quadruplex junctions with superior properties. Importantly, combination of structural and chemical modifications of G-quadruplex based junctions and their properties are still mostly a blank slate, thus leaving a great scope of activity for the future investigations.
This entry is adapted from the peer-reviewed paper 10.3390/ijms22189948
References
- Santosh, B.; Yadava, P.K. Nucleic acid aptamers: Research tools in disease diagnostics and therapeutics. Biomed. Res. Int. 2014, 2014, 540451.
- Tuerk, C.; Gold, L. Systematic evolution of ligands by exponential enrichment: RNA ligands to bacteriophage T4 DNA polymerase. Science 1990, 249, 505–510.
- Ellington, A.D.; Szostak, J.W. In vitro selection of RNA molecules that bind specific ligands. Nature 1990, 346, 818–822.
- Darmostuk, M.; Rimpelova, S.; Gbelcova, H.; Ruml, T. Current approaches in SELEX: An update to aptamer selection technology. Biotechnol. Adv. 2015, 33, 1141–1161.
- Banerjee, J.; Nilsen-Hamilton, M. Aptamers: Multifunctional molecules for biomedical research. J. Mol. Med. 2013, 91, 1333–1342.
- Ni, X.; Castanares, M.; Mukherjee, A.; Lupold, S.E. Nucleic acid aptamers: Clinical applications and promising new horizons. Curr. Med. Chem. 2011, 18, 4206–4214.
- Tucker, W.O.; Shum, K.T.; Tanner, J.A. G-quadruplex DNA aptamers and their ligands: Structure, function and application. Curr. Pharm. Des. 2012, 18, 2014–2026.
- Robinson, J.; Raguseo, F.; Nuccio, S.P.; Liano, D.; Di Antonio, M. DNA G-quadruplex structures: More than simple roadblocks to transcription? Nucleic Acids Res. 2021, 49, 8419–8431.
- Bhattacharyya, D.; Mirihana Arachchilage, G.; Basu, S. Metal Cations in G-Quadruplex Folding and Stability. Front. Chem. 2016, 4, 38.
- Bochman, M.L.; Paeschke, K.; Zakian, V.A. DNA secondary structures: Stability and function of G-quadruplex structures. Nat. Rev. Genet. 2012, 13, 770–780.
- Ogloblina, A.M.; Bannikova, V.A.; Khristich, A.N.; Oretskaya, T.S.; Yakubovskaya, M.G.; Dolinnaya, N.G. Parallel G-Quadruplexes Formed by Guanine-Rich Microsatellite Repeats Inhibit Human Topoisomerase I. Biochemistry 2015, 80, 1026–1038.
- Yang, D. G-Quadruplex DNA and RNA. Methods Mol. Biol. 2019, 2035, 1–24.
- Cho, Y.; Lee, Y.B.; Lee, J.H.; Lee, D.H.; Cho, E.J.; Yu, S.J.; Kim, Y.J.; Kim, J.I.; Im, J.H.; Lee, J.H.; et al. Modified AS1411 Aptamer Suppresses Hepatocellular Carcinoma by Up-Regulating Galectin-14. PLoS ONE 2016, 11, e0160822.
- Fan, X.; Sun, L.; Wu, Y.; Zhang, L.; Yang, Z. Bioactivity of 2′-deoxyinosine-incorporated aptamer AS1411. Sci. Rep. 2016, 6, 25799.
- Virgilio, A.; Amato, T.; Petraccone, L.; Esposito, F.; Grandi, N.; Tramontano, E.; Romero, R.; Haider, S.; Gomez-Monterrey, I.; Novellino, E.; et al. Improvement of the activity of the anti-HIV-1 integrase aptamer T30175 by introducing a modified thymidine into the loops. Sci. Rep. 2018, 8, 7447.
- Fan, X.; Sun, L.; Li, K.; Yang, X.; Cai, B.; Zhang, Y.; Zhu, Y.; Ma, Y.; Guan, Z.; Wu, Y.; et al. The Bioactivity of D-/L-Isonucleoside- and 2′-Deoxyinosine-Incorporated Aptamer AS1411s Including DNA Replication/MicroRNA Expression. Mol. Ther. Nucleic Acids 2017, 9, 218–229.
- Mukundan, V.T.; Do, N.Q.; Phan, A.T. HIV-1 integrase inhibitor T30177 forms a stacked dimeric G-quadruplex structure containing bulges. Nucleic Acids Res. 2011, 39, 8984–8991.
- Kovačič, M.; Podbevšek, P.; Tateishi-Karimata, H.; Takahashi, S.; Sugimoto, N.; Plavec, J. Thrombin binding aptamer G-quadruplex stabilized by pyrene-modified nucleotides. Nucleic Acids Res. 2020, 48, 3975–3986.
- Goji, S.; Matsui, J. Direct Detection of Thrombin Binding to 8-Bromodeoxyguanosine-Modified Aptamer: Effects of Modification on Affinity and Kinetics. J. Nucleic Acids 2011, 2011, 316079.
- Saccà, B.; Lacroix, L.; Mergny, J.-L. The effect of chemical modifications on the thermal stability of different G-quadruplex-forming oligonucleotides. Nucleic Acids Res. 2005, 33, 1182–1192.
- Zhao, X.; Liu, B.; Yan, J.; Yuan, Y.; An, L.; Guan, Y. Structure variations of TBA G-quadruplex induced by 2′-O-methyl nucleotide in K+ and Ca2+ environments. Acta Biochim. Biophys. Sin. 2014, 46, 837–850.
- Varada, M.; Aher, M.; Erande, N.; Kumar, V.A.; Fernandes, M. Methoxymethyl Threofuranosyl Thymidine (4′-MOM-TNA-T) at the T7 Position of the Thrombin-Binding Aptamer Boosts Anticoagulation Activity, Thermal Stability, and Nuclease Resistance. ACS Omega 2020, 5, 498–506.
- Kotkowiak, W.; Wengel, J.; Scotton, C.J.; Pasternak, A. Improved RE31 Analogues Containing Modified Nucleic Acid Monomers: Thermodynamic, Structural, and Biological Effects. J. Med. Chem. 2019, 62, 2499–2507.
- Zaitseva, M.; Kaluzhny, D.; Shchyolkina, A.; Borisova, O.; Smirnov, I.; Pozmogova, G. Conformation and thermostability of oligonucleotide d(GGTTGGTGTGGTTGG) containing thiophosphoryl internucleotide bonds at different positions. Biophys. Chem. 2010, 146, 1–6.
- Gunjal, A.D.; Fernandes, M.; Erande, N.; Rajamohanan, P.R.; Kumar, V.A. Functional isoDNA aptamers: Modified thrombin binding aptamers with a 2′-5′-linked sugar-phosphate backbone (isoTBA). Chem. Commun. 2014, 50, 605–607.
- Esposito, V.; Scuotto, M.; Capuozzo, A.; Santamaria, R.; Varra, M.; Mayol, L.; Virgilio, A.; Galeone, A. A straightforward modification in the thrombin binding aptamer improving the stability, affinity to thrombin and nuclease resistance. Org. Biomol. Chem. 2014, 12, 8840–8843.
- Kosman, J.; Juskowiak, B. Thrombin-Binding Aptamer with Inversion of Polarity Sites (IPS): Effect on DNAzyme Activity and Anticoagulant Properties. Int. J. Mol. Sci. 2021, 22, 7902.
- Miranda, A.; Santos, T.; Largy, E.; Cruz, C. Locking up the AS1411 Aptamer with a Flanking Duplex: Towards an Improved Nucleolin-Targeting. Pharmaceuticals 2021, 14, 121.
- Carvalho, J.; Lopes-Nunes, J.; Lopes, A.C.; Cabral Campello, M.P.; Paulo, A.; Queiroz, J.A.; Cruz, C. Aptamer-guided acridine derivatives for cervical cancer. Org. Biomol. Chem. 2019, 17, 2992–3002.
- Mazurov, A.V.; Titaeva, E.V.; Khaspekova, S.G.; Storojilova, A.N.; Spiridonova, V.A.; Kopylov, A.M.; Dobrovolsky, A.B. Characteristics of a new DNA aptamer, direct inhibitor of thrombin. Bull. Exp. Biol. Med. 2011, 150, 422–425.
- Dolinnaya, N.G.; Yuminova, A.V.; Spiridonova, V.A.; Arutyunyan, A.M.; Kopylov, A.M. Coexistence of G-quadruplex and duplex domains within the secondary structure of 31-mer DNA thrombin-binding aptamer. J. Biomol. Struct. Dyn. 2012, 30, 524–531.
- Russo Krauss, I.; Spiridonova, V.; Pica, A.; Napolitano, V.; Sica, F. Different duplex/quadruplex junctions determine the properties of anti-thrombin aptamers with mixed folding. Nucleic Acids Res. 2015, 44, 983–991.
- Savchik, E.Y.; Kalinina, T.B.; Drozd, N.N.; Makarov, V.A.; Zav’yalova, E.G.; Lapsheva, E.N.; Mudrik, N.N.; Babij, A.V.; Pavlova, G.V.; Golovin, A.V.; et al. Aptamer RA36 inhibits of human, rabbit, and rat plasma coagulation activated with thrombin or snake venom coagulases. Bull. Exp. Biol. Med. 2013, 156, 44–48.
- Kim, Y.; Cao, Z.; Tan, W. Molecular assembly for high-performance bivalent nucleic acid inhibitor. Proc. Natl. Acad. Sci. USA 2008, 105, 5664–5669.
- Alieva, R.; Novikov, R.; Tashlitsky, V.; Arutyunyan, A.; Kopylov, A.; Zavyalova, E. Bimodular thrombin aptamers with two types of non-covalent locks. Nucleosides Nucleotides Nucleic Acids 2021, 40, 559–577.
- Heyduk, E.; Heyduk, T. Nucleic Acid-Based Fluorescence Sensors for Detecting Proteins. Anal. Chem. 2005, 77, 1147–1156.
- Müller, J.; Freitag, D.; Mayer, G.; Pötzsch, B. Anticoagulant characteristics of HD1-22, a bivalent aptamer that specifically inhibits thrombin and prothrombinase. J. Thromb. Haemost. 2008, 6, 2105–2112.
- Tasset, D.M.; Kubik, M.F.; Steiner, W. Oligonucleotide inhibitors of human thrombin that bind distinct epitopes11 Edited by R. Huber. J. Mol. Biol. 1997, 272, 688–698.
- Hughes, Q.W.; Le, B.T.; Gilmore, G.; Baker, R.I.; Veedu, R.N. Construction of a Bivalent Thrombin Binding Aptamer and Its Antidote with Improved Properties. Molecules 2017, 22, 1770.
- Zavyalova, E.; Kopylov, A. DNA aptamer-based molecular nanoconstructions and nanodevices for diagnostics and therapy. In Nanostructures for the Engineering of Cells, Tissues and Organs; Grumezescu, A.M., Ed.; Elsevier: Amsterdam, The Netherlands, 2018; pp. 249–290.
- Michalowski, D.; Chitima-Matsiga, R.; Held, D.M.; Burke, D.H. Novel bimodular DNA aptamers with guanosine quadruplexes inhibit phylogenetically diverse HIV-1 reverse transcriptases. Nucleic Acids Res. 2008, 36, 7124–7135.
- Li, T.; Wang, E.; Dong, S. A Grafting Strategy for the Design of Improved G-Quadruplex Aptamers and High-Activity DNAzymes. PLoS ONE 2009, 4, e5126.
- Vianney, Y.M.; Preckwinkel, P.; Mohr, S.; Weisz, K. Quadruplex-Duplex Junction: A High-Affinity Binding Site for Indoloquinoline Ligands. Chemistry 2020, 26, 16910–16922.
- Bing, T.; Zheng, W.; Zhang, X.; Shen, L.; Liu, X.; Wang, F.; Cui, J.; Cao, Z.; Shangguan, D. Triplex-quadruplex structural scaffold: A new binding structure of aptamer. Sci. Rep. 2017, 7, 15467.
- Kotkowiak, W.; Kotkowiak, M.; Kierzek, R.; Pasternak, A. Unlocked nucleic acids: Implications of increased conformational flexibility for RNA/DNA triplex formation. Biochem. J. 2014, 464, 203–211.
