PON1, PON2, and PON3 belong to a family of lactone hydrolyzing enzymes endowed with various substrate specificities. Among PONs, PON2 shows the highest hydrolytic activity toward many acyl-homoserine lactones (acyl-HL) involved in bacterial quorum-sensing signaling.
- PON2
- catalytic activity
- lactonase
- antioxidant
- bacterial infections
- inflammation
- cancer
- isoforms
- SNPs
- post-translational modifications
- PON2 regulation
1. Introduction
Paraoxonase 2 (PON2) is the oldest member and the most potent quorum quencher of the paraoxonase family, nevertheless it is less studied than PON1. Its intracellular localization, in contrast to PON1 and PON3 secreted extracellularly, makes PON2 studies more challenging. In fact in cells functional assays to measure its activity in different compartments are still not available and the lack of the 3D structure does not allow one to clarify reaction mechanisms. The most common PON2 polymorphisms are associated with its decreased lactonase activity and with a higher risk for coronary artery disease (CAD) and Alzheimer’s disease. From 2010 it was highlighted PON2’s ability to reduce oxidative stress in mitochondria and to prevent apoptosis in the endoplasmic reticulum [1][2][3][4] with a still unclarified mechanism suggested to be independent from the lactonase activity [1]. From here on scientists explored PON2 antioxidant effects, its role in preventing heart failure [5] and its involvement in any type of tumor [6].
Scientists are focusing on individual aspects of PON2. Of the 996 PON2 papers produced from 1998, 792 were published in the last 10 years. This huge amount of information needs to be collected and summarized allowing scientists to look at the whole picture of PON2 functions and roles while continuing to elucidate single mechanisms.
2. PON2 Structure and Function
2.1. Gene and Localization
Based on a phylogenetic analysis, PON2 emerges as the oldest member of this family, with PON1 and PON3 evolving from PON2 [2][8]. Their genes, which reside on the same cluster on chromosome 7 [2], share about 70% sequence identity. It is worth noting a structural similarity of PONs members with the endoplasmic reticulum (ER)-resident molecular chaperone MEC-6 [9]. At the genomic structure level, PON2 is arranged in nine exons encoding a protein of 355 amino acids, approximately 40–43 kDa in mass. PON2 displays a 66% sequence identity and 81% similarity with PON1, at least considering the most abundant form of PON2 [10]. PON1 and PON3 are extracellular proteins secreted in plasma and bound to high-density lipoproteins (HDLs). Their expression is predominant in the liver and kidneys [11][12][13][14]. PON2, in contrast, is a ubiquitously expressed intracellular protein [15]. It localizes in the perinuclear region, the endoplasmic reticulum (ER), mitochondria [3], and associates with plasma membrane fractions [16]. In summary, although PONs are very similar in their amino acid sequences, they have different functions and are found at different locations. For a recent review more focused on PON1 structure and mechanism see [7].
2.2. PON2 Model
PON1 was the first HDL-associated protein and the only PON family member for which the structure has been elucidated [17] (PDB ID: 1V04). The first crystallized PON1 is a recombinant variant from rabbit, highly similar in sequence to human PON1 (Figure 1).
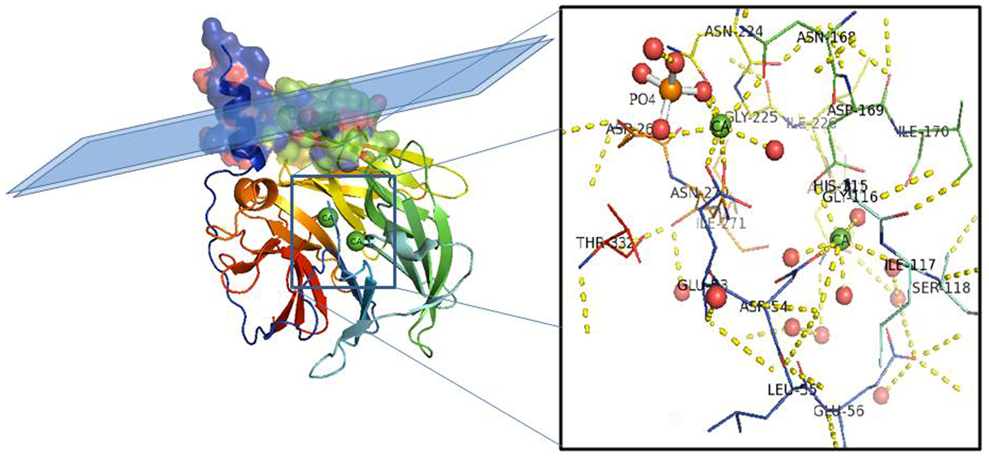
Figure 1. PON1 protein structure. Side view of the six-bladed propeller-like structure of PON1. The top left side of the structure (molecular surface colored by atoms and ribbon view in transparency) shows the retained N-terminal helix A and helix B taking contact with a schematic lipid surface (phospholipids of a high-density lipoprotein (HDL) particle; membrane in the case of PON2). The N and C termini and the two calcium atoms (green balls) localize in the central tunnel of the propeller. The ribbon of PON1 molecule is colored by chain progression. Residues interacting with the two calcium ions and phosphate are shown with line representation (inset). Red balls are water molecules. Picture was drawn by the Pymol program (The PyMOL Molecular Graphics System, Version 2.0. Schrödinger, LLC.).
The overall architecture of PON1 is a β-propeller with six blades and a central tunnel; each blade consists of four β-sheets. A disulfide bridge between Cys42 and Cys353 forms a covalent closure between the N and C termini; the two Cys are conserved throughout the PON family [10][17]. Two calcium ions, one at the top of the structure (Ca1) and one in the central tunnel (Ca2), are present at a distance of 7.4 Å. The calcium at the top, considered to be the catalytic calcium, interacts with the side chain oxygens of Asn224, Asn270, Asn168, Asp269, and Glu53 (line representation in Figure 1 inset). The central calcium ion may contribute to the protein’s structural stability [17][18]. All three PONs diverged as independent genes during evolution and expansion, but they maintained a common active site and catalytic machinery. The PON1 glycosylated sites (Asn253 and Asn324) important for structure and high catalytic activity are highly conserved throughout the whole gene cluster. In PON2 four putative N-linked glycosylation sites (Asn 226, Asn 254, Asn 269, and Asn 323) have been predicted to be present, but only two out of them have been validated experimentally: Asn 254 [19][20] and Asn 323 [1]. Furthermore, overexpression in human umbilical vein endothelial 926 cells (EA.hy) of the full-length PON2 protein, and its mutants at positions 254 and 323, demonstrated them as glycosylation targets and that modifications seem requested for the enzyme hydrolytic activity [1]. PON2 is a type II transmembrane protein, with its N-terminal region identified as a single transmembrane domain, whereas the catalytic domain corresponds to the C-terminus, located extracellularly in the case of plasma membrane localization. Its role here should be to counteract lipid peroxidation [21] as PON1 does in other districts. PON2 3D models have been built based on the 3D structure of the homologue PON1 (PDB codes: 1v04 [22] and 4Q1U [23]). The structure of PON2 (as inferred from PON1) is characterized by the first α-helix (H1) at the N-terminus protruding from the globular structure and the hydrophobic H2 likely interacting with the lipid bilayer (Figure 2). The loop between the strand D6 and H1 α-helix is involved in the structural stabilization of PON1 by several interactions [17]. Looking at the PON2 model (Figure 2), the regions 18–31 and 92–109 were quite dissimilar in sequence from PON1. The residue ASN105 of PON1 seems missing in PON2.
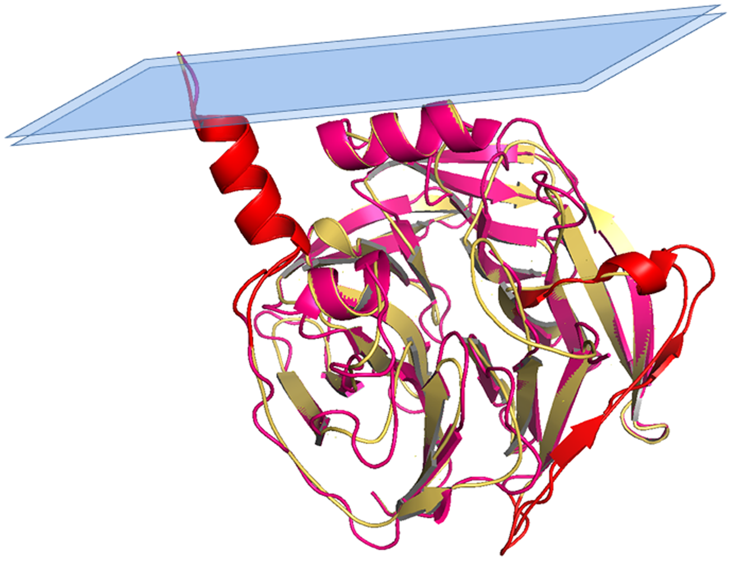
Figure 2. Structural superposition between PON1 and PON2 model. Superposition between PON1 structure (residues 16–355; cyan) and PON2 3D model (residues 16–354; yellow), with highlighted (in red) regions 18–31 and 92–109 mentioned in the text.
2.3. PON2 Activities
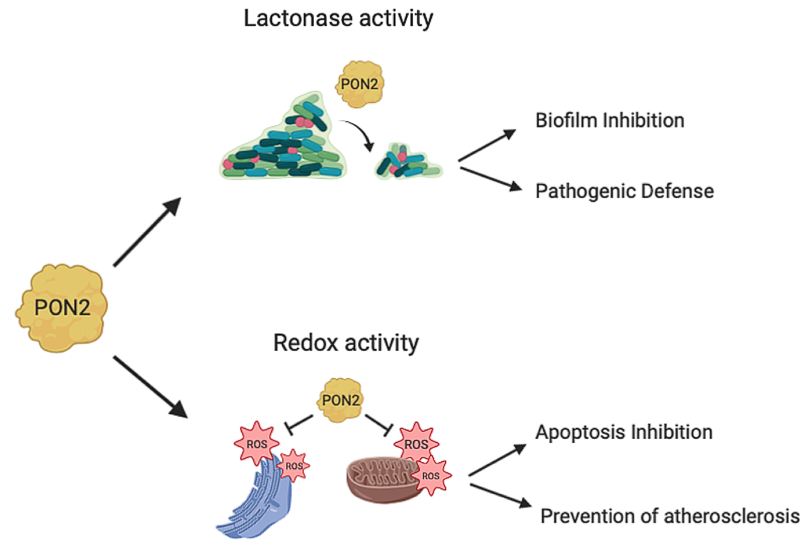
PON2 has a calcium-dependent hydrolytic activity on lactones, esters, and aryl esters [24] and in addition it functions as an antioxidant enzyme (Figure 3). PON2 over-expression is capable of lowering the oxidative state of cells, to prevent and to reverse the cell-mediated oxidative modification of low-density lipoprotein (LDL) and therefore blocks the ability of mildly oxidized LDL (MM-LDL) to induce monocyte chemotaxis [14]. The redox function reduces the levels of reactive oxygen species (ROS) thus displaying an antiapoptotic effect [3]. However, none has so far demonstrated that the anti-ROS activity is truly catalytic. This is an aspect that deserves to be explored in the next future. In contrast with PON1 and PON3, PON2 does not show hydrolytic activity toward phosphotriesters [24][25] albeit faint PTE activity has been recently reported for a mutated recombinant PON2 version [22].
Figure 3. PON2 activities. PON2 with its lactonase activity is able to hydrolyze the quorum sensing signaling molecules used by bacteria during infection. This catalytic activity results in biofilm inhibition and defense from pathogenic infections. PON2 also has antioxidant activity reducing oxidative stress in mitochondria and in the endoplasmatic reticulum with different mechanisms as described in this review (Section 4; 4.1; 4.2). The PON2 redox activity inhibits apoptosis and prevents the formation of atherosclerotic lesions.
2.4. PON2 Isoforms
Seven PON2 mRNA isoforms have been described [8][10]. Some of them include small mRNA size variations that are produced by alternative splicing of the primary transcript, or by use of a second transcription start site. These transcripts predict significant alterations in the deduced proteins such as the premature truncation after 50 or 84 residues (transcript I.3A and I.3B, respectively), the lack of 86 N-terminal residues (transcript II), or the loss of the second putative Ca2+ binding loop (transcripts I.5A and II.5A) [10]. Which of these alternate PON2 transcripts are translated in vivo, and what the biological significance of such variations is, it remains to be established. At the protein level, three isoforms corresponding to alternative splicing are described [23]. The canonical full-length protein 354 aa (39,381 kDa), an isoform that differs from the canonical sequence as follows: 1-16: MGRLVAVGLLGIALAL → MGAWVGCGLAGDRAGF (transcript 1) [8]; and a third isoform missing the region: 123-134 (342 aa; 37,980 kDa) [26][27]. While the existence in vivo of the canonical PON2 isoform was obvious from several studies, the existence and function of the two non-canonical isoforms as expressed proteins is still a matter of debate. In our recent paper [23], peptides corresponding to these two isoforms were identified by mass spectrometry (MS) of endogenous PON2 immunoprecipitated in HeLa cells. This result defines the presence in vivo of the non-canonical isoforms as expressed proteins. The isoform with the deletion of twelve amino acids of exon V lacks one residue of the active site (His134) that helps to increase His 115 basicity in the homologous PON1 structure [17][22]. Therefore, it is highly likely that its deletion makes the protein substantially inactive [24]. The mutant 123-134delrPON2 harboring the deletion was produced in E. coli and a small-angle X-ray scattering (SAXS) method was applied for the structure reconstruction of this protein. The results showed a disordered protein suggesting that this isoform is unstructured and mostly inactive, as catalytic activity assays also demonstrated [23].
2.5. PON2 SNPs
PON2 possesses unique properties that distinguish it from PON1 and PON3. PON2 is ubiquitously expressed in many different tissue types and is highly expressed in the vital organs, such as heart brain and lungs. Early research revealed that PON2 is exclusively found intracellularly, wherein it functions as an antioxidative protein by reducing intracellular and local oxidative stress. Studies in the last five years have demonstrated that PON2 protects against atherosclerosis by preventing LDL oxidation, reversing the oxidation of mildly oxidized LDL, inhibiting monocyte chemotaxis, and increasing cholesterol efflux. Recently, emerging evidence has proposed that PON2 is antiatherosclerotic and may be associated with cardiovascular disease (CVD). The number of investigations concerning the relationship between two common PON2 polymorphisms and CVD among different ethnic groups and regions is rapidly growing. Here, we briefly review the developments in PON2 research by focusing on past and recent findings. In 1996 two polymorphisms were detected in PON2 coding sequences in the Pima Indians and Caucasians that predict an A148→G148 and an S311→C311 substitution in the protein deduced from the transcript I [8]. The clear concordance of the genotypes between both polymorphisms in the Pima Indians and Caucasians indicates a strong disequilibrium between these variants. Considering the potential importance of the conserved cysteines in PON1 [28], it is intriguing to speculate that the introduction of an additional cysteine by the polymorphism at codon 311 may affect the structure and/or function of PON2. However, from the PON2 model the putative Cys311 is far away from the other cysteines (e.g., Cys284), excluding interference with or formation of a new bond. Cys284 in PON1 has been reported to be important for activity [29], whereas cysteines in general are the target of S-glutathionylation. Rozemberg et al. (2006) [30] reported that S-glutathionylation regulates HDL-associated paraoxonase 1 (PON1) activity. It is possible that post-translational modification of Cys284 can affect the nearby His285 and their interaction with the catalytic calcium. However, we did not detect PON2 S-glutathionylation (unpublished). The two reported PON2 polymorphisms associate with alterations of plasma lipid levels in patients affected by human diseases related to oxidative stress such as coronary artery disease (CAD) [31][32][33], type 2 diabetes mellitus, Alzheimer’s disease [34][35], and reduced bone mass in postmenopausal women [36]. Almost total linkage disequilibrium exists between these two polymorphic sites in four different human populations, which indicate that the genotype at one location can be used as a surrogate for the genotype at the other one. In other words, the A148 and S311 variants form one common allelic haplotype, while the G148 and C311 ones form the second common allelic haplotype in white, South Asian, and African samples [37]. PON2 A148/S311 homozygotes exhibited significantly higher plasma total and LDL cholesterol and apoB than subjects with the other two genotypes [37], which was confirmed by an investigation of Shin et al. [38]. The PON2 S311C SNP affects lactonase activity [19], a finding not confirmed by a subsequent study [1]. Regarding its role in CAD, it is reported an association [33], while the PON2 codon 311 Cys/Cys genotypes are significantly associated with CAD severity in terms of the number of diseased vessels but not in terms of stenosis severity [39]. A meta-analysis performed in 2004 showed conflicting results with no significant association of PON2 Ser311Cys polymorphism [40], but a more recent research clarified that the PON2 Ser311Cys polymorphism is associated with CHD risk in Caucasians, but there is no association between this polymorphism and CHD in Asians or Hispanic populations [41]. The Ser311Cys polymorphism and low levels of HDL contribute to a higher mortality risk after acute myocardial infarction in elderly patients [42], and it is associated with a risk of large vessel disease stroke in a Polish population [43]. Besides, a higher risk for cardiovascular diseases and for Alzheimer’s disease is described in carriers of the 311C allele [44][45]. Significant evidence of an association between polymorphisms in the PON gene cluster (Ser311Cys in PON2) and AD in African Americans and Caucasians was observed [46]. Newborn babies with PON2 148GA/GG genotype were also found to be at high risk of low weight and short length at birth when exposed to di-n-butyl phthalate (MBP) and di-2-ethylhexyl phthalate (MEHP) [47]. A combined genotype analysis for PON2 polymorphisms revealed that the combination of G148 and S311 was more frequent in cataract subjects, whereas heterogeneous alleles at 311 and 148 were more likely to be present in subjects without lens opacity. It seems likely that the presence of mutations at 148 and 311 positions might be considered as a risk factor for the development of cataracts [48]. The PON2 gene polymorphisms A148G and S311C have been independently associated with diabetic nephropathy in type II diabetic patients. The susceptibility to diabetic nephropathy positively correlated with the degree of obesity [49]. PON2 expression and insulin resistance relationships were also observed [50]. In diabetes mellitus, the PON2-311 SNP associated with the presence of microvascular complications, with an over-representation of the C/C 311 genotype [51]. A recombinant PON2 was expressed in E. coli to test the role of these SNPs on PON2 activity [23]. Broad decrease of lactonase, esterase, and phosphotriesterase activities was observed for all mutants, particularly lactonase activity against 3OC12HSL, which should represent the physiological substrate for PON2. In our system, we confirmed the data of Stoltz et al. [19] that reported lower activity of the mutant expressed in CHO cells and decreased lactonase activity in primary airway epithelial cells characterized by the C/C genotype. CD spectra in the far and near UV showed similar structures in wt rPON2 and mutants, allowing one to rule out that the decrease in activity was due to a fault in the refolding of mutants [23].
2.6. PON2 Protein–Protein Interactions
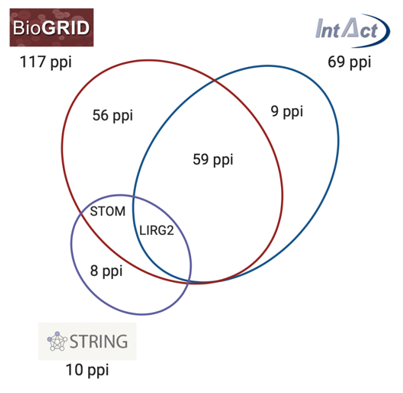
By comparing the main databases and repositories for protein interactions PON2 is reported to physically interact with 117 (BioGRID), 69 (IntAct), and 10 (STRING) human proteins, some of them overlapping and others being reported only by one database (Figure 4). BioGRID and IntAct share 60 protein–protein interactions (ppi), STOM ppi is reported by STRING and BIOGRID, while LIRG2 is common in all three databases.
Figure 4. Schematic representation of PON2 protein–protein interactions (ppi) network reported by BioGRID, IntAct, and STRING databases. The intersection between circles represents common ppi shared by the different databases.
PON2 also interacts with two HIV-1 proteins: the envelope glycoprotein gp160 (env) [52] and the protein Rev (rev), which its function is to escort unspliced or incompletely spliced viral pre-mRNAs (late transcripts) out of the nucleus of infected cells [53]. Due to the latest SARS-CoV-2 pandemic it is also interesting to highlight the SARS-CoV-2 proteins interacting with PON2: the envelope small membrane protein (E), the membrane glycoprotein (M), the non-structural protein 4 (Nsp4), the non-structural protein 6 (Nsp6), the open reading frame 7a (ORF7a), the non-structural Protein 7b (Nsp7b), the non-structural protein 8 (Nsp8), and the open reading frame 9b (ORF9b) [54][55] (see also graphical abstract).
Interactome mapping of the relevant protein is the main scope of current biological research, similar to the way “genome” projects were a driving force of molecular biology 20 years ago. PON2 interactome characterization would allow us to understand the intricate physicochemical dynamic connections of PON2 biological functions at both cellular and systems levels. So far only a few of the identified PON2 interactions were deeply investigated to understand their relationships and significance and here we report last discoveries. Nagarajan et al. (2017) [56] demonstrated the role of PON2 protein/protein interaction in modulating an “old” cancer trait, namely the Warburg effect. In the pancreatic ductal adenocarcinoma cancer (PDAC), the release of p53 translational inhibition of PON2 results in an indirect effect of PON2 on GLUT1, a glucose transporter, mediated by the interaction with the protein stomatin (STOM) [57]. Furthermore, PON2/GLUT1 interaction prevents AMP-activated protein kinase (AMPK)-mediated anoikis via ATP inhibition of the pathway AMPK-FOXO3A-PUMA, which in turn favors metastasis [56]. In 2018 PON2 was demonstrated to interact with leucine-rich repeats and immunoglobulin-like domains 1 (LRIG1) [58]. PON2 was a determinant of the PDGFRA–down-regulating function of LRIG1. PON2 also bidirectionally interacted with LINGO1, another leucine-rich repeat and immunoglobulin-like domain-containing protein that has been reported to negatively regulate TRK (NTRK) receptors in a manner that appears to be highly similar to the mechanism described for LRIG1 [59].
This entry is adapted from the peer-reviewed paper 10.3390/antiox10020256
References
- Altenhöfer, S.; Witte, I.; Teiber, J.F.; Wilgenbus, P.; Pautz, A.; Li, H.; Daiber, A.; Witan, H.; Clement, A.M.; Förstermann, U.; et al. One Enzyme, Two Functions. Biol. Chem. 2010, 285, 24398–24403, doi:10.1074/jbc.m110.118604.
- Devarajan, A.; Bourquard, N.; Hama, S.; Navab, M.; Grijalva, V.R.; Morvardi, S.; Clarke, C.F.; Vergnes, L.; Reue, K.; Teiber, J.F.; et al. Paraoxonase 2 Deficiency Alters Mitochondrial Function and Exacerbates the Development of Atherosclerosis. Redox Signal. 2011, 14, 341–351, doi:10.1089/ars.2010.3430.
- Horke, S.; Witte, I.; Wilgenbus, P.; Krü; Strand, D.; FörstermannU. Paraoxonase-2 Reduces Oxidative Stress in Vascular Cells and Decreases Endoplasmic Reticulum Stress–Induced Caspase Activation. Circulation 2007, 115, 2055–2064, doi:10.1161/circulationaha.106.681700.
- Witte, I.; Altenhöfer, S.; Wilgenbus, P.; Amort, J.; Clement, A.M.; Pautz, A.; Li, H.; Förstermann, U.; Horke, S. Beyond reduction of atherosclerosis: PON2 provides apoptosis resistance and stabilizes tumor cells. Cell Death Dis. 2011, 2, e112, doi:10.1038/cddis.2010.91.
- Li, W.; Kennedy, D.; Shao, Z.; Wang, X.; Kamdar, A.K.; Weber, M.; Mislick, K.; Kiefer, K.; Morales, R.; Agatisa-Boyle, B.; et al. Paraoxonase 2 prevents the development of heart failure. Free Radic. Biol. Med. 2018, 121, 117–126, doi:10.1016/j.freeradbiomed.2018.04.583.
- Shakhparonov, M.I.M.I.; Antipova, N.V.; Shender, V.O.; Shnaider, P.V.; Arapidi, G.P.; Pestov, N.B.; Pavlyukov, M.S.; M.M. Shemyakin-Yu.A. Ovchinnikov Institute of Bioorganic Chemistry of the Russian Academy of Sciences Expression and Intracellular Localization of Paraoxonase 2 in Different Types of Malignancies. Acta Naturae 2018, 10, 92–99, doi:10.32607/20758251-2018-10-3-92-99.
- Taler-Verčič, A.; Goličnik, M.; Bavec, A. The Structure and Function of Paraoxonase-1 and Its Comparison to Paraoxonase-2 and -3. Molecules 2020, 25, 5980, doi:10.3390/molecules25245980.
- Primo-Parmo, S.L.; Sorenson, R.C.; Teiber, J.; Du, B.N. The Human Serum Paraoxonase/Arylesterase Gene (PON1) Is One Member of a Multigene Family. Genomics 1996, 33, 498–507, doi:10.1006/geno.1996.0225.
- Chen, Y.; Bharill, S.; Altun, Z.; O’Hagan, R.; Coblitz, B.; Isacoff, E.Y.; Chalfie, M. Caenorhabditis elegans paraoxonase-like proteins control the functional expression of DEG/ENaC mechanosensory proteins. Biol. Cell 2016, 27, 1272–1285, doi:10.1091/mbc.e15-08-0561.
- Mochizuki, H.; Scherer, S.W.; Xi, T.; Nickle, D.C.; Majer, M.; Huizenga, J.J.; Tsui, L.-C.; Prochazka, M. Human PON2 gene at 7q21.3: Cloning, multiple mRNA forms, and missense polymorphisms in the coding sequence. Gene 1998, 213, 149–157, doi:10.1016/s0378-1119(98)00193-0.
- Diepgen, T.L.; Mallinckrodt, M.G.-V. Interethnic Differences in the Detoxification of Organophosphates: The Human Serum Paraoxonase Polymorphism. Toxicol. 1986, 9, 154–158, doi:10.1007/978-3-642-71248-7_18.
- Jaouad, L.; De Guise, C.; Berrougui, H.; Cloutier, M.; Isabelle, M.; Fülöp, T.; Payette, H.; Khalil, A. Age-related decrease in high-density lipoproteins antioxidant activity is due to an alteration in the PON1’s free sulfhydyl groups. Atherosclerosis 2006, 185, 191–200, doi:10.1016/j.atherosclerosis.2005.06.012.
- Leviev, I.; Negro, F.; James, R.W. Two Alleles of the Human Paraoxonase Gene Produce Different Amounts of mRNA. Thromb. Vasc. Biol. 1997, 17, 2935–2939, doi:10.1161/01.atv.17.11.2935.
- Sierksma, A.; Van Der Gaag, M.S.; Van Tol, A.; James, R.W.; Hendriks, H.F.J. Kinetics of HDL cholesterol and paraoxonase activity in moderate alcohol consumers. Clin. Exp. Res. 2002, 26, 1430–1435.
- Ng, C.J.; Wadleigh, D.J.; Gangopadhyay, A.; Hama, S.; Grijalva, V.R.; Navab, M.; Fogelman, A.M.; Reddy, S.T. Paraoxonase-2 Is a Ubiquitously Expressed Protein with Antioxidant Properties and Is Capable of Preventing Cell-mediated Oxidative Modification of Low Density Lipoprotein. Biol. Chem. 2001, 276, 44444–44449, doi:10.1074/jbc.m105660200.
- Ng, C.J.; Shih, D.M.; Hama, S.Y.; Villa, N.; Navab, M.; Reddy, S.T. The paraoxonase gene family and atherosclerosis. Free Radic. Biol. Med. 2005, 38, 153–163, doi:10.1016/j.freeradbiomed.2004.09.035.
- Harel, M.; Aharoni, A.; Gaidukov, L.; Brumshtein, B.; Khersonsky, O.; Meged, R.; Dvir, H.; Ravelli, R.B.G.; McCarthy, A.; Toker, L.; et al. Structure and evolution of the serum paraoxonase family of detoxifying and anti-atherosclerotic enzymes. Struct. Mol. Biol. 2004, 11, 412–419, doi:10.1038/nsmb767.
- Kuo, C.L.; La Du, B.N. Calcium binding by human and rabbit serum paraoxonases. Structural stability and enzymatic activity. Drug Metab. Dispos. 1998, 26, 653–660.
- Stoltz, D.A.; Ozer, E.A.; Recker, T.J.; Estin, M.; Yang, X.; Shih, D.M.; Lusis, A.J.; Zabner, J. A Common Mutation in Paraoxonase-2 Results in Impaired Lactonase Activity. Biol. Chem. 2009, 284, 35564–35571, doi:10.1074/jbc.m109.051706.
- Chen, R.; Jiang, X.; Sun, D.; Han, G.; Wang, F.; Ye, M.; Wang, L.; Zou, H. Glycoproteomics Analysis of Human Liver Tissue by Combination of Multiple Enzyme Digestion and Hydrazide Chemistry. Proteome Res. 2009, 8, 651–661, doi:10.1021/pr8008012.
- Hagmann, H.; Kuczkowski, A.; Ruehl, M.; Lamkemeyer, T.; Brodesser, S.; Horke, S.; Dryer, S.; Schermer, B.; Benzing, T.; Brinkkoetter, P.T. Breaking the chain at the membrane: Paraoxonase 2 counteracts lipid peroxidation at the plasma membrane. FASEB J. 2014, 28, 1769–1779, doi:10.1096/fj.13-240309.
- Mandrich, L.; Cerreta, M.; Manco, G. An Engineered Version of Human PON2 Opens the Way to Understand the Role of Its Post-Translational Modifications in Modulating Catalytic Activity. PLoS ONE 2015, 10, doi:10.1371/journal.pone.0144579.
- Carusone, T.M.; Cardiero, G.; Cerreta, M.; Mandrich, L.; Moran, O.; Porzio, E.; Catara, G.; Lacerra, G.; Manco, G. WTAP and BIRC3 are involved in the posttranscriptional mechanisms that impact on the expression and activity of the human lactonase PON2. Cell Death Dis. 2020, 11, 324–327, doi:10.1038/s41419-020-2504-2.
- Draganov, D.I.; Teiber, J.F.; Speelman, A.; Osawa, Y.; Sunahara, R.; La Du, B.N. Human paraoxonases (PON1, PON2, and PON3) are lactonases with overlapping and distinct substrate specificities. Lipid Res. 2005, 46, 1239–1247, doi:10.1194/jlr.m400511-jlr200.
- Teiber, J.F.; Horke, S.; Haines, D.C.; Chowdhary, P.K.; Xiao, J.; Kramer, G.L.; Haley, R.W.; Draganov, D.I. Dominant Role of Paraoxonases in Inactivation of the Pseudomonas aeruginosa Quorum-Sensing Signal N-(3-Oxododecanoyl)-l-Homoserine Lactone. Immun. 2008, 76, 2512–2519, doi:10.1128/iai.01606-07.
- Sameshima, E.; Tabata, Y.; Hayashi, A.; Lida, K.; Mitsuyama, M.; Kanai, S.; Saito, T. Paraoxonase mRNA, nirs splice variant1. Submitted (FEB-2003) to the EMBL/GenBank/DDBJ databases.
- Ota, T.; Suzuki, Y.; Nishikawa, T.; Otsuki, T.; Sugiyama, T.; Irie, R.; Wakamatsu, A.; Hayashi, K.; Sato, H.; Nagai, K.; et al. Complete sequencing and characterization of 21,243 full-length human cDNAs. Genet. 2003, 36, 40–45, doi:10.1038/ng1285.
- Sorenson, R.C.; Primo-Parmo, S.L.; Camper, S.A.; La Du, B.N. The Genetic Mapping and Gene Structure of Mouse Paraoxonase/Arylesterase. Genomics 1995, 30, 431–438, doi:10.1006/geno.1995.1261.
- Aviram, M.; Billecke, S.; Sorenson, R.; Bisgaier, C.; Newton, R.; Rosenblat, M.; Erogul, J.; Hsu, C.; Dunlop, C.; La Du, B. Paraoxonase Active Site Required for Protection Against LDL Oxidation Involves Its Free Sulfhydryl Group and Is Different From That Required for Its Arylesterase/Paraoxonase Activities. Thromb. Vasc. Biol. 1998, 18, 1617–1624, doi:10.1161/01.atv.18.10.1617.
- Rozenberg, O.; Aviram, M. S-Glutathionylation regulates HDL-associated paraoxonase 1 (PON1) activity. Biophys. Res. Commun. 2006, 351, 492–498, doi:10.1016/j.bbrc.2006.10.059.
- Wang, X.; Fan, Z.; Huang, J.; Su, S.; Yu, Q.; Zhao, J.; Hui, R.; Yao, Z.; Shen, Y.; Qiang, B.; et al. Extensive Association Analysis Between Polymorphisms of PON Gene Cluster with Coronary Heart Disease in Chinese Han Population. Thromb. Vasc. Biol. 2003, 23, 328–334, doi:10.1161/01.atv.0000051702.38086.c1.
- Martinelli, N.; Girelli, D.; Olivieri, O.; Stranieri, C.; Trabetti, E.; Pizzolo, F.; Friso, S.; Tenuti, I.; Cheng, S.; Grow, M.A.; et al. Interaction between smoking and PON2 Ser311Cys polymorphism as a determinant of the risk of myocardial infarction. J. Clin. Investig. 2004, 34, 14–20, doi:10.1111/j.1365-2362.2004.01292.x.
- Sanghera, D.K.; Aston, C.E.; Saha, N.; Kamboh, M.I. DNA Polymorphisms in Two Paraoxonase Genes (PON1 and PON2) Are Associated with the Risk of Coronary Heart Disease. J. Hum. Genet. 1998, 62, 36–44, doi:10.1086/301669.
- Janka, Z.; Juhász, A.; Rimanóczy, Á.; Boda, K.; Márki-Zay, J.; Kálmán, J. Codon 311 (Cys → Ser) polymorphism of paraoxonase-2 gene is associated with apolipoprotein E4 allele in both Alzheimer’s and vascular dementias. Psychiatry 2002, 7, 110–112, doi:10.1038/sj.mp.4000916.
- Shi, J.; Zhang, S.; Tang, M.; Liu, X.; Li, T.; Han, H.; Wang, Y.; Guo, Y.; Zhao, J.; Li, H.; et al. Possible association between Cys311Ser polymorphism of paraoxonase 2 gene and late-onset Alzheimer’s disease in Chinese. Brain Res. 2004, 120, 201–204, doi:10.1016/j.molbrainres.2003.10.018.
- Yamada, Y.; Ando, F.; Niino, N.; Miki, T.; Shimokata, H. Association of polymorphisms of paraoxonase 1 and 2 genes, alone or in combination, with bone mineral density in community-dwelling Japanese. Hum. Genet. 2003, 48, 469–475, doi:10.1007/s10038-003-0063-x.
- Hegele, R.A. Paraoxonase genes and disease. Med. 1999, 31, 217–224, doi:10.3109/07853899909115981.
- Shin, B.-S.; Oh, S.-Y.; Kim, Y.-S.; Kim, K.-W. The paraoxonase gene polymorphism in stroke patients and lipid profile. Acta Neurol. Scand. 2008, 117, 237–243, doi:10.1111/j.1600-0404.2007.00929.x.
- Chen, Q.; Reis, S.E.; Kammerer, C.M.; McNamara, D.M.; Holubkov, R.; Sharaf, B.L.; Sopko, G.; Pauly, D.F.; Merz, C.N.B.; Kamboh, M.I. Association between the Severity of Angiographic Coronary Artery Disease and Paraoxonase Gene Polymorphisms in the National Heart, Lung, and Blood Institute–Sponsored Women’s Ischemia Syndrome Evaluation (WISE) Study. J. Hum. Genet. 2003, 72, 13–22, doi:10.1086/345312.
- Wheeler, J.G.; Keavney, B.D.; Watkins, H.; Collins, R.; Danesh, J. Four paraoxonase gene polymorphisms in 11 212 cases of coronary heart disease and 12 786 controls: Meta-analysis of 43 studies. Lancet 2004, 363, 689–695, doi:10.1016/s0140-6736(04)15642-0.
- Chen, M.-L.; Zhao, H.; Liao, N.; Xie, Z.-F. Association Between Paraoxonase 2 Ser311Cys Polymorphism and Coronary Heart Disease Risk: A Meta-Analysis. Med Sci. Monit. 2016, 22, 3196–3201, doi:10.12659/msm.896601.
- Marchegiani, F.; Spazzafumo, L.; Provinciali, M.; Cardelli, M.; Olivieri, F.; Franceschi, C.; Lattanzio, F.; Antonicelli, R. Paraoxonase2 C311S polymorphism and low levels of HDL contribute to a higher mortality risk after acute myocardial infarction in elderly patients. Genet. Metab. 2009, 98, 314–318, doi:10.1016/j.ymgme.2009.05.008.
- Słowik, A.; Wloch, D.; Szermer, P.; Wolkow, P.P.; Malecki, M.; Pera, J.; Turaj, W.; Dziedzic, T.; Klimkowicz-Mrowiec, A.; Kopec, G.; et al. Paraoxonase 2 Gene C311S Polymorphism Is Associated with a Risk of Large Vessel Disease Stroke in a Polish Population. Dis. 2007, 23, 395–400, doi:10.1159/000101462.
- Porntadavity, S.; Permpongpaiboon, T.; Sukketsiri, W. Human paraoxonase 2. EXCLI J. 2010, 9, 159–172.
- Leus, F.R.; Zwart, M.; Kastelein, J.J.; Voorbij, H.A. PON2 gene variants are associated with clinical manifestations of cardiovascular disease in familial hypercholesterolemia patients. Atherosclerosis 2001, 154, 641–649, doi:10.1016/s0021-9150(00)00440-8.
- Erlich, P.M.; Lunetta, K.L.; Cupples, L.A.; Huyck, M.; Green, R.C.; Baldwin, C.T.; Farrer, L.A.; Auerbach, S.; Akomolafe, A.; Griffith, P.; et al. Polymorphisms in the PON gene cluster are associated with Alzheimer disease. Mol. Genet. 2006, 15, 77–85, doi:10.1093/hmg/ddi428.
- Xie, C.; Jin, R.; Zhao, Y.; Lin, L.; Li, L.; Chen, J.; Zhang, Y. Paraoxonase 2 gene polymorphisms and prenatal phthalates’ exposure in Chinese newborns. Res. 2015, 140, 354–359, doi:10.1016/j.envres.2015.03.028.
- Baig, A.; Rehman, A.-U.; Zarina, S. Association of PON2 and PON3 polymorphism with risk of developing cataract. Saudi J. Ophthalmol. 2019, 33, 153–158, doi:10.1016/j.sjopt.2019.05.001.
- Pinizzotto, M.; Castillo, E.; Fiaux, M.; Temler, E.; Gaillard, R.C.; Ruiz, J. Paraoxonase2 polymorphisms are associated with nephropathy in Type II diabetes. Diabetology 2001, 44, 104–107, doi:10.1007/s001250051586.
- Qujeq, D.; Mahrooz, A.; Alizadeh, A.; Boorank, R. Paraoxonase-2 variants potentially influence insulin resistance, beta-cell function, and their interrelationships with alanine aminotransferase in type 2 diabetes. Res. Med. Sci. 2018, 23, 107.
- Mackness, B.; McElduff, P.; Mackness, M.I. The paraoxonase-2-310 polymorphism is associated with the presence of microvascular complications in diabetes mellitus. Intern. Med. 2005, 258, 363–368, doi:10.1111/j.1365-2796.2005.01554.x.
- Jäger, S.; Cimermancic, P.; Gulbahce, N.; Johnson, J.R.; McGovern, K.E.; Clarke, S.C.; Shales, M.; Mercenne, G.; Pache, L.; Li, K.; et al. Global landscape of HIV–human protein complexes. Nature 2011, 481, 365–370, doi:10.1038/nature10719.
- Naji, S.; Ambrus, G.; Cimermančič, P.; Reyes, J.R.; Johnson, J.R.; Filbrandt, R.; Huber, M.D.; Vesely, P.; Krogan, N.J.; Yates, J.R.; et al. Host Cell Interactome of HIV-1 Rev Includes RNA Helicases Involved in Multiple Facets of Virus Production. Cell. Proteom. 2012, 11, doi:10.1074/mcp.m111.015313.
- Available online: https://www.biorxiv.org/content/10.1101/2020.09.03.282103v1 (accessed on 20 December 2020 ).
- Available online: https://www.biorxiv.org/content/10.1101/2020.08.28.272955v1 (accessed on20 December 2020 )
- Nagarajan, A.; Dogra, S.K.; Sun, L.; Gandotra, N.; Ho, T.; Cai, G.; Cline, G.; Kumar, P.; Cowles, R.A.; Wajapeyee, N. Paraoxonase 2 Facilitates Pancreatic Cancer Growth and Metastasis by Stimulating GLUT1-Mediated Glucose Transport. Cell 2017, 67, 685–701.e6, doi:10.1016/j.molcel.2017.07.014.
- Havugimana, P.C.; Hart, G.T.; Nepusz, T.; Yang, H.; Turinsky, A.L.; Li, Z.; Wang, P.I.; Boutz, D.R.; Fong, V.; Phanse, S.; et al. A Census of Human Soluble Protein Complexes. Cell 2012, 150, 1068–1081, doi:10.1016/j.cell.2012.08.011.
- Faraz, M.; Herdenberg, C.; Holmlund, C.; Henriksson, R.; Hedman, H. A protein interaction network centered on leucine-rich repeats and immunoglobulin-like domains 1 (LRIG1) regulates growth factor receptors. Biol. Chem. 2018, 293, 3421–3435, doi:10.1074/jbc.m117.807487.
- Meabon, J.S.; De Laat, R.; Ieguchi, K.; Serbzhinsky, D.; Hudson, M.P.; Huber, B.R.; Wiley, J.C.; Bothwell, M. Intracellular LINGO-1 negatively regulates Trk neurotrophin receptor signaling. Cell. Neurosci. 2016, 70, 1–10, doi:10.1016/j.mcn.2015.11.002.