Multiple myeloma is the second most common hematologic malignancy in the United States. Eventually, all myeloma patients will relapse and develop resistance to currently available agents. There is an unmet medical need to identify novel therapeutic targets. PIM kinases play an important role in myeloma pathogenesis and disease relapse.
- PIM kinase
- inhibitor
- myeloma
- resistance
- PI3K/Akt/mTOR
[1][2][3][4][5][6][7][8][9][10][11][12][13][14][15][16][17][18][19][20][21][22][23][24][25][26][27][28][29][30][31][32][33][34][35][36][37][38][39][40][41][42][43][44]
1. Introduction
Multiple myeloma (MM) is a hematologic malignancy characterized by the proliferation of malignant plasma cells. Precision medicine has heralded an era of change and challenge in the treatment of patients with MM. Although targeted therapies and immunotherapies have made significant advances in the personalized treatment of MM, clinicians still face the persistence of disease recurrence and drug resistance. The acquisition of anti-cancer drug resistance is a major issue with therapies in MM. Cancer cells utilize multiple intercellular and intracellular signaling cascades mediated by oncogenes such as PIM kinases to maintain cell growth and survival. In normal cells, the activity of these kinases is tightly controlled, whereas their sustained activation promotes apoptotic resistance and uncontrolled proliferation in cancer cells . The complexity of the kinase molecular signaling network along with its crosstalk with alternative oncogenic signaling pathways provides ample opportunities for MM to develop productive adaptive mechanisms. PIM kinase activation has been shown to play a significant role in this bypass signaling mechanism. A better understanding of PIM kinase synergism, in addition to other signaling pathways, is important to the development of PIM inhibitors and to provide the rationale of combination therapy to improve the treatment efficacy for patients with MM.
2. Background—Expression and Regulation of PIM Kinases
3. PIM Kinase and Cancers
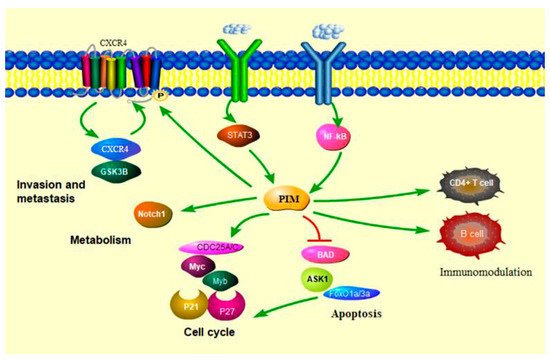
3.1. PIM Kinases in Cancer Cell Cycle
3.2. PIM Kinases in Cancer Cell Survival
3.3. PIM Kinases in Cancer Cell Metabolism
3.4. PIM Kinases and Immune Modulation
This entry is adapted from the peer-reviewed paper 10.3390/cancers13174304
References
- [1] N.A. Warfel, A.S. Kraft, PIM kinase (and Akt) biology and signaling in tumors, Pharmacol Ther, 151 (2015) 41-49.
- [2] N. An, Y.W. Lin, S. Mahajan, J.N. Kellner, Y. Wang, Z. Li, A.S. Kraft, Y. Kang, Pim1 serine/threonine kinase regulates the number and functions of murine hematopoietic stem cells, Stem Cells, 31 (2013) 1202-1212.
- [3] N. An, A.S. Kraft, Y. Kang, Abnormal hematopoietic phenotypes in Pim kinase triple knockout mice, J Hematol Oncol, 6 (2013) 12.
- [4] X. Le, R. Antony, P. Razavi, D.J. Treacy, F. Luo, M. Ghandi, P. Castel, M. Scaltriti, J. Baselga, L.A. Garraway, Systematic Functional Characterization of Resistance to PI3K Inhibition in Breast Cancer, Cancer Discov, 6 (2016) 1134-1147.
- [5] R.S. Iyer, L. Chatham, R. Sleigh, D.W. Meek, A functional SUMO-motif in the active site of PIM1 promotes its degradation via RNF4, and stimulates protein kinase activity, Sci Rep, 7 (2017) 3598.
- [6] K. Adam, M. Lambert, E. Lestang, G. Champenois, I. Dusanter-Fourt, J. Tamburini, D. Bouscary, C. Lacombe, Y. Zermati, P. Mayeux, Control of Pim2 kinase stability and expression in transformed human haematopoietic cells, Biosci Rep, 35 (2015).
- [7] L. Mologni, V. Magistroni, F. Casuscelli, M. Montemartini, C. Gambacorti-Passerini, The Novel PIM1 Inhibitor NMS-P645 Reverses PIM1-Dependent Effects on TMPRSS2/ERG Positive Prostate Cancer Cells And Shows Anti-Proliferative Activity in Combination with PI3K Inhibition, J Cancer, 8 (2017) 140-145.
- [8] D. Hildebrand, K. Heeg, K.F. Kubatzky, Pasteurella multocida Toxin Manipulates T Cell Differentiation, Front Microbiol, 6 (2015) 1273.
- [9] S. Darici, H. Alkhaldi, G. Horne, H.G. Jorgensen, S. Marmiroli, X. Huang, Targeting PI3K/Akt/mTOR in AML: Rationale and Clinical Evidence, J Clin Med, 9 (2020).
- [10] D.S. Hoover, D.G. Wingett, J. Zhang, R. Reeves, N.S. Magnuson, Pim-1 protein expression is regulated by its 5'-untranslated region and translation initiation factor elF-4E, Cell Growth Differ, 8 (1997) 1371-1380.
- [11] N.A. Keane, M. Reidy, A. Natoni, M.S. Raab, M. O'Dwyer, Targeting the Pim kinases in multiple myeloma, Blood Cancer J, 5 (2015) e325.
- [12] C.O. Leung, C.C. Wong, D.N. Fan, A.K. Kai, E.K. Tung, I.M. Xu, I.O. Ng, R.C. Lo, PIM1 regulates glycolysis and promotes tumor progression in hepatocellular carcinoma, Oncotarget, 6 (2015) 10880-10892.
- [13] O. Kim, T. Jiang, Y. Xie, Z. Guo, H. Chen, Y. Qiu, Synergism of cytoplasmic kinases in IL6-induced ligand-independent activation of androgen receptor in prostate cancer cells, Oncogene, 23 (2004) 1838-1844.
- [14] H. Mikkers, M. Nawijn, J. Allen, C. Brouwers, E. Verhoeven, J. Jonkers, A. Berns, Mice deficient for all PIM kinases display reduced body size and impaired responses to hematopoietic growth factors, Mol Cell Biol, 24 (2004) 6104-6115.
- [15] J. Wang, P.D. Anderson, W. Luo, D. Gius, M. Roh, S.A. Abdulkadir, Pim1 kinase is required to maintain tumorigenicity in MYC-expressing prostate cancer cells, Oncogene, 31 (2012) 1794-1803.
- [16] F. Braso-Maristany, S. Filosto, S. Catchpole, R. Marlow, J. Quist, E. Francesch-Domenech, D.A. Plumb, L. Zakka, P. Gazinska, G. Liccardi, P. Meier, A. Gris-Oliver, M.C. Cheang, A. Perdrix-Rosell, M. Shafat, E. Noel, N. Patel, K. McEachern, M. Scaltriti, P. Castel, F. Noor, R. Buus, S. Mathew, J. Watkins, V. Serra, P. Marra, A. Grigoriadis, A.N. Tutt, PIM1 kinase regulates cell death, tumor growth and chemotherapy response in triple-negative breast cancer, Nat Med, 22 (2016) 1303-1313.
- [17] B. Zhao, L. Liu, J. Mao, Z. Zhang, Q. Wang, Q. Li, PIM1 mediates epithelial-mesenchymal transition by targeting Smads and c-Myc in the nucleus and potentiates clear-cell renal-cell carcinoma oncogenesis, Cell Death Dis, 9 (2018) 307.
- [18] X.H. Zhang, H.L. Yu, F.J. Wang, Y.L. Han, W.L. Yang, Pim-2 Modulates Aerobic Glycolysis and Energy Production during the Development of Colorectal Tumors, Int J Med Sci, 12 (2015) 487-493.
- [19] X. Zhang, M. Song, J.K. Kundu, M.H. Lee, Z.Z. Liu, PIM Kinase as an Executional Target in Cancer, J Cancer Prev, 23 (2018) 109-116.
- [20] A.K. Yadav, V. Kumar, D.B. Bailey, B.C. Jang, AZD1208, a Pan-Pim Kinase Inhibitor, Has Anti-Growth Effect on 93T449 Human Liposarcoma Cells via Control of the Expression and Phosphorylation of Pim-3, mTOR, 4EBP-1, S6, STAT-3 and AMPK, Int J Mol Sci, 20 (2019).
- [21] F. Cervantes-Gomez, L.S. Chen, R.Z. Orlowski, V. Gandhi, Biological effects of the Pim kinase inhibitor, SGI-1776, in multiple myeloma, Clin Lymphoma Myeloma Leuk, 13 Suppl 2 (2013) S317-329.
- [22] H. Koblish, Y.L. Li, N. Shin, L. Hall, Q. Wang, K. Wang, M. Covington, C. Marando, K. Bowman, J. Boer, K. Burke, R. Wynn, A. Margulis, G.W. Reuther, Q.T. Lambert, V. Dostalik Roman, K. Zhang, H. Feng, C.B. Xue, S. Diamond, G. Hollis, S. Yeleswaram, W. Yao, R. Huber, K. Vaddi, P. Scherle, Preclinical characterization of INCB053914, a novel pan-PIM kinase inhibitor, alone and in combination with anticancer agents, in models of hematologic malignancies, PLoS One, 13 (2018) e0199108.
- [23] E. Motylewska, M. Braun, H. Stepien, High Expression of NEK2 and PIM1, but Not PIM3, Is Linked to an Aggressive Phenotype of Bronchopulmonary Neuroendocrine Neoplasms, Endocr Pathol, 31 (2020) 264-273.
- [24] K. Saurabh, M.T. Scherzer, P.P. Shah, A.S. Mims, W.W. Lockwood, A.S. Kraft, L.J. Beverly, The PIM family of oncoproteins: small kinases with huge implications in myeloid leukemogenesis and as therapeutic targets, Oncotarget, 5 (2014) 8503-8514.
- [25] C.T. Cottage, B. Bailey, K.M. Fischer, D. Avitabile, B. Collins, S. Tuck, P. Quijada, N. Gude, R. Alvarez, J. Muraski, M.A. Sussman, Cardiac progenitor cell cycling stimulated by pim-1 kinase, Circ Res, 106 (2010) 891-901.
- [26] P. Mondello, S. Cuzzocrea, M. Mian, Pim kinases in hematological malignancies: where are we now and where are we going?, J Hematol Oncol, 7 (2014) 95.
- [27] L.S. Chen, K. Balakrishnan, V. Gandhi, Inflammation and survival pathways: chronic lymphocytic leukemia as a model system, Biochem Pharmacol, 80 (2010) 1936-1945.
- [28] T.L. Aho, J. Sandholm, K.J. Peltola, H.P. Mankonen, M. Lilly, P.J. Koskinen, Pim-1 kinase promotes inactivation of the pro-apoptotic Bad protein by phosphorylating it on the Ser112 gatekeeper site, FEBS Lett, 571 (2004) 43-49.
- [29] B. Yan, M. Zemskova, S. Holder, V. Chin, A. Kraft, P.J. Koskinen, M. Lilly, The PIM-2 kinase phosphorylates BAD on serine 112 and reverses BAD-induced cell death, J Biol Chem, 278 (2003) 45358-45367.
- [30] J.J. Gu, Z. Wang, R. Reeves, N.S. Magnuson, PIM1 phosphorylates and negatively regulates ASK1-mediated apoptosis, Oncogene, 28 (2009) 4261-4271.
- [31] C. Hogan, C. Hutchison, L. Marcar, D. Milne, M. Saville, J. Goodlad, N. Kernohan, D. Meek, Elevated levels of oncogenic protein kinase Pim-1 induce the p53 pathway in cultured cells and correlate with increased Mdm2 in mantle cell lymphoma, J Biol Chem, 283 (2008) 18012-18023.
- [32] N.M. Santio, S.K. Landor, L. Vahtera, J. Yla-Pelto, E. Paloniemi, S.Y. Imanishi, G. Corthals, M. Varjosalo, G.B. Manoharan, A. Uri, U. Lendahl, C. Sahlgren, P.J. Koskinen, Phosphorylation of Notch1 by Pim kinases promotes oncogenic signaling in breast and prostate cancer cells, Oncotarget, 7 (2016) 43220-43238.
- [33] V. Carafa, L. Altucci, A. Nebbioso, Dual Tumor Suppressor and Tumor Promoter Action of Sirtuins in Determining Malignant Phenotype, Front Pharmacol, 10 (2019) 38.
- [34] S. Guo, X. Mao, J. Chen, B. Huang, C. Jin, Z. Xu, S. Qiu, Overexpression of Pim-1 in bladder cancer, J Exp Clin Cancer Res, 29 (2010) 161.
- [35] C. Xue, Y. He, Q. Hu, Y. Yu, X. Chen, J. Chen, F. Ren, J. Li, Z. Ren, G. Cui, R. Sun, Downregulation of PIM1 regulates glycolysis and suppresses tumor progression in gallbladder cancer, Cancer Manag Res, 10 (2018) 5101-5112.
- [36] S. Chatterjee, P. Chakraborty, A. Daenthanasanmak, S. Iamsawat, G. Andrejeva, L.A. Luevano, M. Wolf, U. Baliga, C. Krieg, C.C. Beeson, M. Mehrotra, E.G. Hill, J.C. Rathmell, X.Z. Yu, A.S. Kraft, S. Mehrotra, Targeting PIM Kinase with PD1 Inhibition Improves Immunotherapeutic Antitumor T-cell Response, Clin Cancer Res, 25 (2019) 1036-1049.
- [37] L.J. Jackson, J.A. Pheneger, T.J. Pheneger, G. Davis, A.D. Wright, J.E. Robinson, S. Allen, M.C. Munson, L.L. Carter, The role of PIM kinases in human and mouse CD4+ T cell activation and inflammatory bowel disease, Cell Immunol, 272 (2012) 200-213.
- [38] T.L. Aho, R.J. Lund, E.K. Ylikoski, S. Matikainen, R. Lahesmaa, P.J. Koskinen, Expression of human pim family genes is selectively up-regulated by cytokines promoting T helper type 1, but not T helper type 2, cell differentiation, Immunology, 116 (2005) 82-88.
- [39] W. Du, T. Chen, Y. Ni, X. Hou, Y. Yu, Q. Zhou, F. Wu, W. Tang, G. Shi, Role of PIM2 in allergic asthma, Mol Med Rep, 16 (2017) 7504-7512.
- [40] M. Szydlowski, S. Debek, M. Prochorec-Sobieszek, M. Szolkowska, A.M. Tomirotti, P. Juszczynski, A. Szumera-Cieckiewicz, PIM Kinases Promote Survival and Immune Escape in Primary Mediastinal Large B-Cell Lymphoma through Modulation of JAK-STAT and NF-kappaB Activity, Am J Pathol, 191 (2021) 567-574.
- [41] J. Asano, A. Nakano, A. Oda, H. Amou, M. Hiasa, K. Takeuchi, H. Miki, S. Nakamura, T. Harada, S. Fujii, K. Kagawa, I. Endo, K. Yata, A. Sakai, S. Ozaki, T. Matsumoto, M. Abe, The serine/threonine kinase Pim-2 is a novel anti-apoptotic mediator in myeloma cells, Leukemia, 25 (2011) 1182-1188.
- [42] M. Hiasa, J. Teramachi, A. Oda, R. Amachi, T. Harada, S. Nakamura, H. Miki, S. Fujii, K. Kagawa, K. Watanabe, I. Endo, Y. Kuroda, T. Yoneda, D. Tsuji, M. Nakao, E. Tanaka, K. Hamada, S. Sano, K. Itoh, T. Matsumoto, M. Abe, Pim-2 kinase is an important target of treatment for tumor progression and bone loss in myeloma, Leukemia, 29 (2015) 207-217.
- [43] T.R. Ullah, The role of CXCR4 in multiple myeloma: Cells' journey from bone marrow to beyond, J Bone Oncol, 17 (2019) 100253.
- [44] I.M. Ghobrial, Y. Zhang, Y. Liu, H. Ngo, F. Azab, A. Sacco, A. Azab, P. Maiso, B. Morgan, P. Quang, G.C. Issa, X. Leleu, A.M. Roccaro, Targeting the bone marrow in Waldenstrom macroglobulinemia, Clin Lymphoma Myeloma Leuk, 11 Suppl 1 (2011) S65-69.
