3.1. Variations in the Taxonomical, Phylogenetical Diversity and Composition of Bacterial Communities
Significant variations in PD and taxonomical diversity of bacterial communities could be detected based on seagrass species within a coral reef ecosystem, but only significant variations in taxonomical diversity for the same seagrass species from different sampling locations (Table 2). Moreover, significant taxonomical and phylogenetic variations only existed among different seagrass species collected from the Xisha Islands. Therefore, the coral reef ecosystem’s seagrass species may be one important factor in shaping the rhizosphere bacterial communities.
We also found significant differences in the taxonomy composition of rhizosphere bacterial communities at the genus level based on different seagrass sampling locations. This was partly consistent with the investigation result of Cúcio et al. (2016) [
16], the result of which demonstrated that significant differences were detected for the same seagrass species from different sampling locations, but no significant differences existed between the rhizobiomes of different seagrass species from the same sampling location. The reason for this phenomenon may be that different seagrass species were included in each study. Three different seagrass species, namely
Z. marina,
Z. noltii, and
Cymodocea nodos, were studied for Cúcio et al. (2016) [
16], while four seagrass species (
C. nodos,
T. hemprichii,
H. ovalis, and
S. isoetifolium) were examined in our investigation. Another reason for this discrepancy may be the different growth habits. The seagrass habitats for their study was in the intertidal regions, while all the seagrasses in this study were collected from the coral reef ecosystem [
16]. Moreover, previous studies have highlighted the importance of temperature in constructing the rhizosphere bacterial community anwhich exhibited seasonal variations [
43,
44]. Therefore, there may also be seasonal variations in the seagrass rhizosphere bacterial community. More investigation on the temporal scale in the future needs to be performed.
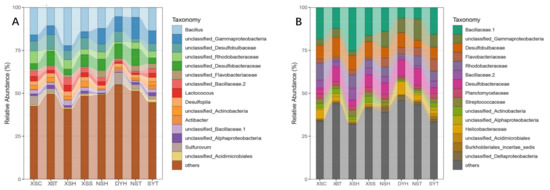
Proteobacteria (class alpha-, beta-, delta-, gamma-, and epsilon-proteobacteria) and the Firmicutes were the two most predominant phyla across the four coral reef ecosystems. Besides, class Deltaproteobacteria accounted for over 20% of all investigated bacterial communities. Cúcio et al. (2016) also reported that the phylum Proteobacteria was the most dominant in the rhizomes of seagrass
Z. marina,
Z. noltii, and
Cymodocea nodosa, with the proportion ranging from 65% to 68%. The existence of plants played a crucial role in shaping the microbial community in the rhizosphere of seagrasses as the seagrass rhizosphere bacterial community composition was quite different from that of the surrounding water and bulk sediment [
16]. Besides, seagrass (
Z. marina) colonization increased the abundance of the nitrogen fixation bacteria and other bacteria involved in benthic carbon and sulfur cycling [
45].
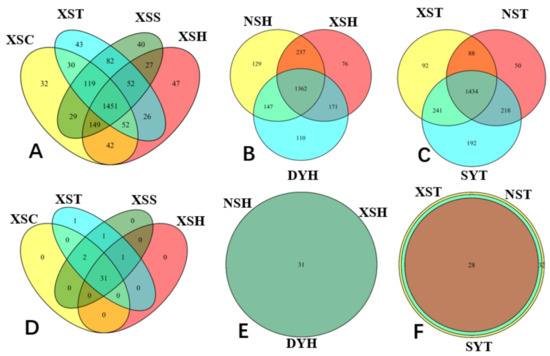
Moreover, some OTUs were peculiar to one coral reef ecosystem, and each coral reef had its own individual OTUs in our study. For instance, OTU1109 was affiliated to class Phycisphaerae SHA-43 belonging to the phylum Planctomycetes and could only be discovered at XS. It may play an important role in the nitrogen cycle by participating in the anammox process, which was assumed as a predominant source of N
2 production in anoxic marine environments [
46,
47,
48]. Moreover, bacteria from the family
Rhodothermaceae (phylum Bacteroidetes) were specially retrieved from Sanya Bay, and microorganisms from this family were usually isolated from the extreme environments and exhibited extreme thermophilic or halophilic characteristics [
49,
50].
3.2. The Functional Structure of Microbial Communities in Seagrass Rhizosphere
Seagrass holobionts have been reported to play essential roles in the cycle of sulfur, nitrogen, and carbon, at both microbial structural and functional levels [
11,
13], and Ugarelli et al. (2019) [
51] reported that the seagrass plant and its microbiome were highly interlinked in the cycle of sulfur, nitrogen, and carbon. Likewise, FAPROTAX analysis revealed that many microbes in the seagrass rhizosphere of coral reef ecosystems participated in these processes (
Figure S3).
Previous studies showed that increased sulfide concentration in the sediment caused by the activity of sulfate-reducing prokaryotes was one of the main reasons for seagrass death all over the world [
11,
13]. However, seagrass could oxygenate their roots [
41] and lose radial oxygen in the rhizosphere of young roots to lower the concentration of sulfide to protect themselves [
52]. What is more, sulfur-oxidizing bacteria in this ecosystem may also alleviate the sulfide stress for seagrass by oxidation of sulfide [
53]. A higher abundance of genes was found to participate in the process of sulfur oxidation than sulfate reduction in the rhizosphere of the seagrass
Z. marina [
14]. We also found a high percentage of sulfate respiration (129 records) and respiration of sulfur compounds (131 records) in the FAPROTAX analysis result. This may indicate that microbes in the seagrass rhizosphere also play an important role in the sulfur-related cycle in the coral reef ecosystem.
Bioavailable nitrogen is crucial to all living organisms, but it is still a limiting nutrient globally [
54]. The nitrogen enters the ecosystem from the air in the form of ammonia by the microbial nitrogen fixation, which is an essential link of the nitrogen cycle due to nitrogen usually acting as the limiting factor for productivity in the oligotrophic seagrass meadow and coral reef ecosystems [
55]. Welsh et al. (2002) found that the microbes capable of sulfate-reducing are the significant component of the diazotrophs in many seagrass ecosystems [
56]. Besides, the microbes involved in the nitrogen cycle, as revealed by FAPROTAX analysis in this study, mainly involved in the process of nitrification, aerobic ammonia oxidation, nitrate reduction, nitrate respiration, and nitrogen respiration.
Furthermore, the microbes conducted of nitrification activity mainly came from the genus
Nitrosopumilus,
Nitrososphaera, and
Nitrospira. Nitrification is a process of oxidizing ammonia via nitrite to nitrate, which was assumed as a two-step process catalyzed by chemolithoautotrophic microorganisms before 2015 [
57,
58]. Daims et al. (2015) have reported that a completely nitrifying bacterium from the genus
Nitrospira, which was present in diverse environments, and those findings confirmed that completely nitrifying
Nitrospira played important roles in the nitrogen cycle-related microbial functional groups [
58]. Although the ammonia available concentrations in most ocean waters are low, this is suitable for the living of comammox organisms. However, no comammox gene has been found in ocean waters until now. To explore microbes capable of comammox, a future research hotspot for environmental microbiologists is underway [
54].
The diversity of carbon metabolism found in this study was very high, such as aerobic chemoheterotrophy, chemoheterotrophy, fermentation, aromatic compound degradation, photoautotrophy, methanogenesis, and methylotrophy (
Figure S3). Many microbes of the phylum Planctomycetes were involved in the process of aerobic chemoheterotrophy. Like the genus
Blastopirellula, a dominant chemoorganotrophic genus in the Black Sea sediments, are chemoheterotrophic [
59,
60], and their the major carbon and energy sources are carbohydrates [
59]. Eight OTUs were detected in the methylotrophy from the genus
Methanomassiliicoccus, unclassified Methylophilaceae, and
Methylophaga, which accounted for 0.87% of all detected functional groups. Moreover, the putative methylotrophic bacteria, such as
Methylotenera and
Methylophaga, were more abundant in healthy seagrasses and could be used as indicators of seagrass health root microbiomes [
61]. Besides, the microbes involved in sulfur-cycling, including sulfide-oxidizing (e.g., Candidatus Thiodiazotropha and
Candidatus Electrothrix) and sulfate-reducing (e.g., SEEP-SRB1,
Desulfomonile, and
Desulfonema), were more abundant in stressed seagrass [
61]. Hence, there is a need to investigate the relationship between the composition and functions of rhizosphere microbes and seagrass health.