4. The Double Role of Autophagy in Cancer
Autophagy is a highly conserved catabolic mechanism that involves the formation of double-membrane vesicles, the autophagosomes, by which cellular materials are delivered to lysosomes for degradation and recycled into metabolic and biosynthetic pathways. Cytoplasmic materials are delivered to the lysosome through various type of autophagy: macroautophagy, microautophagy and chaperone-mediated autophagy (CMA). Among these various types of autophagy, macroautophagy (autophagy hereafter) is the most extensively analyzed and the major catabolic mechanism used by eukaryotic cells to maintain nutrient homeostasis and organellar quality control. On the contrary, CMA is not mediated by the autophagosome and the cytoplasmic substrates are recognized by chaperone proteins, such as Hsc70, and directed toward translocation into the lysosome for degradation [
59].
Autophagy is mediated by a set of conserved genes. From yeast genetic studies to those on mammalian, the breakthrough in elucidation of the molecular machinery in autophagy came from the discovery of 35 autophagy-related (
ATGs) genes. All
ATG genes are required for the different steps of the autophagy: among them,
ATG 1–10,
12–14,
16–18,
29 and
31 are essential for the efficient formation of autophagosome. The lysosomal degradation pathway is usually described as involving a set of about 16–20 core conserved genes. The ATG proteins encoded by these genes are traditionally classified into distinct biochemical and functional groups that act at specific stages of the autophagic flux [
59,
60]. The formation and turnover of the autophagosome is divided into five distinct stages: (i) initiation due to starvation conditions or other stress factors, during which a decrease of glucose transport results in the release of mTOR inhibition of the ULK1 complex; (ii) nucleation of the autophagosome by ULK1 and class III PI3K complexes; (iii) expansion and elongation of the autophagosome membrane mediated by two ubiquitin-like conjugation systems, the first system being represented by LC3I/PE and LC3II complex, while the second one involving ATG5-ATG12 conjugate mediated by ATG7 and ATG10 genes; (iv) closure and autophagosome fusion with lysosome to form an autophagolysosome by the SNARE protein syntaxin 17 (STX17); and (v) degradation of intravesicular products due to the low pH of the lysosome [
61].
Autophagy has opposite and context-dependent roles in cancer: under certain circumstances, autophagy may be detrimental either via its prosurvival effects or via possible cell-death promoting effects. Thus, the role of autophagy in cancer is complex and controversial. Autophagy was originally thought to represent only a tumor suppression mechanism since Aita et al. and Liang et al. found an allelic loss of an autophagic gene,
BECN1 (
ATG6), whose position was in close proximity to the tumor suppressor breast cancer 1 gene (
BRCA1) [
8,
62].
In the early stages of neoplastic transformation, autophagy can act as a mechanism to counteract tumorigenesis by preventing the accumulation of damaged proteins and organelles and the excessive production of ROS that can promote DNA mutations and thus the development of neoplastic cells. In this way, autophagy limits oncogenic signaling and suppresses the onset of cancer. This may suggest a role for the stimulation of the autophagic process in the prevention of cancer occurrence [
63]. Over the past 10 years, significant progress has been made in understanding the molecular mechanisms of autophagy and one conceptual advance is that autophagy can act also as a tumor promotion mechanism. The ability of autophagy to support cell survival under unfavorable environmental conditions, such as lack of nutrients or oxygen, which are extremely frequent in a growing tumor could help the survival of cancer cells. The tumors therefore exploit autophagy to their own advantage to promote their survival through the self-production of metabolic substrates necessary for the sustenance and spread of the tumor. Although it has been recognized that autophagy has an impact on the regulation of cell growth, the precise role of autophagy is highly contextual. Indeed, the aberrant stimulation of autophagy can determine an excessive auto-degradative event and self-eating mechanism supporting cell death.
Knocking down the expression of essential autophagy genes or deleting them can reduce tumorigenesis, confirming the functional importance of autophagy in tumor promotion. Autophagy is also upregulated in hypoxic tumor regions where it is required for tumor cell survival [
64]. Thus, both the activation of cancer pathways within tumor cells and stress in the tumor microenvironment can increase the requirement for autophagy to promote tumor growth and survival [
63].
Our group demonstrated that the inhibition of expression of autophagic genes by mutant p53 increases the proliferation of pancreatic cancer cells [
14]. These results support the hypothesis of a new mechanism by which oncogenic mutant p53 protein promotes tumor proliferation with the concomitant inhibition of autophagy. The discovery of this double role of autophagy in human cancers has already led to the development of promising new cancer drugs so far. In another recent study, Ranieri et al. identified a new biological mechanism, which acts on the blocking of autophagic protective process and consequently inducing cell death and the reduction and elimination of the tumor [
65]. In this study, they also identified a new pharmacologically active molecule, which can modulate this process resulting in a specific antitumor activity in melanoma cells.
5. GAPDH-Mediated Autophagy
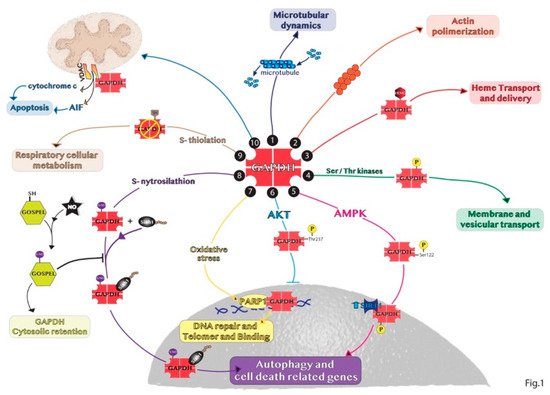
The moonlight GAPDH protein is one of the regulators of autophagy. Besides playing the glycolytic role in the cytosol, GAPDH participates in several non-glycolytic functions including autophagy [
66]. Various conditions exhibit a correlation between autophagy and the translocation of GAPDH in different subcellular compartments, especially to nucleus. For instance, GAPDH negatively regulates autophagy through the interaction with the key autophagy component ATG3 in plants [
67,
68]. Indeed, ROS affects the interaction between cytoplasmic GAPDH and ATG3, making free ATG3 proteins available for use in autophagy. In cardiomyocytes, oxidative stress conditions induce mitochondrial association with GAPDH promoting mitophagy, the selective degradation of mitochondria by autophagy. The formation of multiorganellar lysosomal-like (LL) structures for elimination of damaged mitochondria, also by GAPDH, helps cardiomyocytes to survive during reperfusion or reoxygenation-induced injury [
53]. In brain, autophagic cytotoxic effect of cocaine is mediated by the nitric oxide-GAPDH signaling pathway [
69]. Several molecular mechanisms may trigger or prevent GAPDH nuclear translocation to regulate autophagic events particularly in cancer. For instance, we discovered that the oncogenic mutant p53 protein prevents GAPDH-mediated autophagy in pancreatic cancer cells blocking the nuclear translocation of GAPDH through the stimulation of AKT and inhibition of AMPK signaling pathways [
39,
70], which are reported to directly phosphorylate GAPDH resulting in the inhibition or in the stimulation of GAPDH nuclear translocation, respectively [
47,
71]. Furthermore, in the same cancer type the production of ROS by inhibition of the antioxidant mitochondrial uncoupling protein 2 (UCP2) stimulates the nuclear translocation of GAPDH which promotes autophagy [
38].
The molecular mechanisms involved in GAPDH-mediated autophagy have not yet been clarified. However, GAPDH is supposed to regulate autophagy through the direct interaction of regulatory proteins in the nucleus. In chaperone-mediated autophagy (CMA), oxidized GAPDH may be a specific substrate of this proteolytic pathway. This occurs in various steps, which include binding of the enzyme to the lysosomal membrane through the interaction with monomeric LAMP-2A (lysosome-associated membrane protein type 2A), uptake into lysosomal matrix, and degradation inside this cellular compartment [
72,
73]. Both heat shock cognate proteins, hsc70 and hsp90, play critical roles in the dynamics of these steps.
In low-glucose conditions, GAPDH stimulates autophagy by different pathways. One is the inhibition of mTOR signaling by the interaction between GAPDH and the Ras superfamily of GTPases Rheb, preventing Rheb binding to mTOR [
74] and regulating the cross talk between glycolysis pathway and the mTORC1 pathway. In this way, GAPDH may stimulate autophagy, as mTOR inhibition causes autophagic induction [
75]. In addition, under glucose deficiency GAPDH is phosphorylated by AMPK and it translocates into the nucleus, binds and activates SIRT1 deacetylase, disassociating its inhibitor DBC1 [
29]. Consequently, activated SIRT1 is known to stimulate autophagy, i.e., through the deacetylation of the key autophagy component LC3 in the nucleus, which is essential for its redistribution to the cytoplasm and association with autophagic membranes [
76,
77]. Thus, GAPDH-mediated SIRT1 activation favors the initiation of autophagy. Interestingly, since autophagy might have a role in the promotion of cancer cell survival [
4], GAPDH may act as a prosurvival factor in cancer through the induction of autophagy to support the energy consumption by rapid cell proliferation. Colell et al. showed that nuclear GAPDH protects cells from caspase-independent cell death (CICD), inducing autophagy [
78]. Specifically, the nuclear function of GAPDH in protecting cells from CICD is mediated by the transcriptional up-regulation of Atg12. Since nuclear GAPDH has been involved in transcriptional regulation [
28], the authors suggested that GAPDH may transcriptionally regulate ATG12 directly or indirectly. Thus, GAPDH coordinates two metabolic pathways producing ATP by glycolysis and removing damaged mitochondria by autophagy to shift the cells away from CICD [
78,
79]. Furthermore, Bertin et al. showed that in colon carcinoma cells the increase of GAPDH expression is sufficient to induce autophagy in vitro and in vivo [
80]. Even in esophageal cancer tissues, there is a strong correlation between overexpression of GAPDH and upregulation of autophagy-related genes, like ATG12 and PIK3C3 (phosphatidylinositol 3-kinase catalytic subunit type 3) [
81]. Thus, the knock-down of GAPDH was found to be sufficient to reduce autophagy and ATP levels in tumor cells [
82]. Indeed, the usage of koningic acid (KA), a specific GAPDH inhibitor, inhibited autophagic flux in neuroblastoma cells [
83].
In summary, the manipulation of autophagy through the regulation of the nuclear translocation of GAPDH may be an important therapeutic application in oncology. For instance, we discovered that the combined treatment with target drugs, i.e., genipin and everolimus, which are inhibitors of UCP2 and mTOR, respectively, promotes GAPDH nuclear translocation stimulating the formation of autophagic vesicles and pancreas cancer cell death [
38]. On the contrary, when tumors promote autophagy as a survival mechanism, the modulation of nuclear GAPDH may be a useful target to counteract autophagy. Indeed, Guan et al. showed that GAPDH-siRNA encapsulated in nano-targeted liposomes reduces the autophagic flux in cancer cells and intriguingly favors the outcome of cancer cell drug resistance [
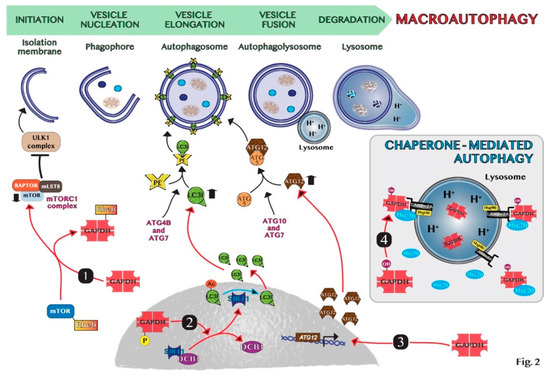
82]. Thus, GAPDH may constitute a valuable target to modulate autophagy in cancer therapy and the main mechanisms at the basis of GAPDH-mediated autophagy regulation are schematically shown in .
Figure 2. Schematic representation of molecular mechanisms of GAPDH-mediated autophagy: (1) GAPDH binds to Rheb, preventing Rheb binding to mTOR. (2) Phosphorylated GAPDH by AMPK translocates into the nucleus and disruptions the link between SIRT1 and DBC1. SIRT1 deacetylates LC3, which sustains autophagosome formation. (3) GAPDH enters the nucleus, favoring the induction of ATG12 gene. (4) In chaperone-mediated autophagy (CMA), GAPDH binds to LAMP-2A receptor on the lysosomal membrane. The chaperon proteins hsc70 and hsp90 are essential for CMA activity. Cytosolic Hsc70 after binding to CMA-substrates, such as GAPDH, transfers the complex to LAMP-2A. Hsp90 stabilizes LAMP-2A at the luminal side of the lysosomal membrane for the translocation of substrates.
6. Aggregation Mechanisms of GAPDH and Impact on Diseases
Protein aggregation is another process of the multiple functions involving GAPDH [
84]. The high-resolution structure of the enzyme shows a highly stable tetrameric protein with identical subunits, each constituting an active site [
40]. Tetrameric dissociation of the enzyme produces dimers and monomers that aggregate or bind to other biomolecules such as proteins and nucleic acids [
16,
85]. The apoform of the enzyme without its cofactors NAD
+ or NADH, is susceptible to denaturation and consequently aggregation. These events are dependent on the content of cofactors and other ligands and on the presence of different signal molecules such as ROS that may influence the amyloidogenic processes through intermolecular disulfide bonds [
25,
84]. Indeed, GAPDH is an intracellular sensor of oxidative stress as discussed above. Some cysteine residues of the active site of the enzyme are strongly susceptible to various types of oxidation, decreasing the enzyme affinity to the cofactor and promoting its dissociation. The active site Cys152 of GAPDH is crucial for its aggregation induced by nitric oxide [
86]. Furthermore, the site-directed mutagenesis experiments revealed that Cys149 is one of the most aggregate-prone cysteine residue of GAPDH induced by oxidative stress [
25]. Several pieces of evidence show the involvement of GAPDH in the development of neurodegenerative disorders such as Alzheimer’s, Huntington’s, and Parkinson’s diseases, which are characterized by the accumulation of protein aggregates [
87,
88,
89]. Indeed, denatured GAPDH forms may bind soluble Aβ species yielding insoluble aggregates [
90,
91]. The deposition of β-amyloid proteins is one of the most distinct features in Alzheimer’s disease and results from proteolytic processing of β-amyloid precursor protein (β-APP) [
91]. GAPDH binds β-APP altering the normal processing of β-APP to produce β-amyloid protein and this interaction was found in amyloid plaques from the brains of patients with AD [
42,
92,
93]. Furthermore, GAPDH aggregates promote the formation of Lewy bodies in the brains of individuals with PD [
88]. GAPDH–protein interactions also occur with spinocerebellar ataxia type-1 and spinobulbar muscular atrophy gene products [
94,
95,
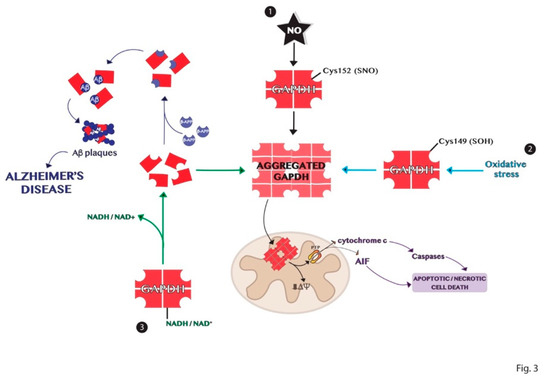
96]. In conclusion, the comprehension of the mechanisms involved in the formation of GAPDH aggregates, represented in , may help in the understanding of the biological alterations observed in neurodegenerative diseases.
Figure 3. Schematic representation of the mechanisms of GAPDH aggregation. (1) Nitrosylation of Cys152 in the active site of GAPDH is crucial for its aggregation induced by nitric oxide. (2) Oxidation of the redox-sensitive Cys149 of GAPDH induced by oxidative stress is involved in its aggregation. (3) GAPDH without its cofactor NADH or NAD+ is susceptible to denaturation and consequent aggregation. Dimer or monomers of GAPDH can aggregate or bind other biomolecules in neurodegenerative diseases, as Alzheimer’s disease.