In vitro lung models are used to faithfully model basic human pathology and the complexity and diversity of human respiratory tissues and to study emerging viral respiratory pathogens and diseases. These models include conventional cell lines, primary human airway epithelial cell (hAEC) cultures, lung organoids, lung-on-a-chip technology, ex vivo lung perfusion (EVLP) models and human lung tissue explants.
- SARS-CoV-2
- in vitro lung model
- cell culture
- human airway epithelial cell culture
- human lung organoids
- lung-on-a-chip
- ex vivo lung
- air–liquid interface
1. Introduction
1.1. Anatomy and Cellular Characteristics of the Human Respiratory System
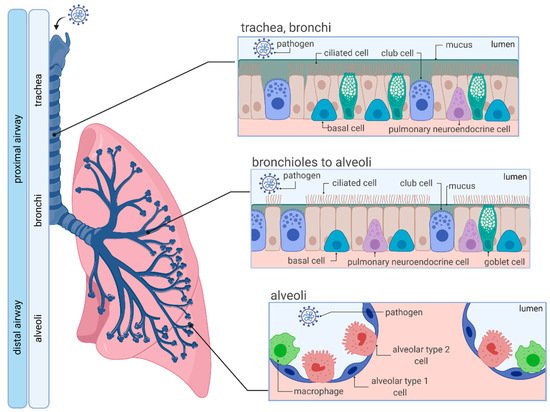
1.2. SARS-CoV-2
2. In Vitro Cell Culture Models for SARS-CoV-2 Infections
2.1. Conventional Cell Lines—Invaluable Tools for Life Sciences
|
Cell Lines |
Origin |
Characteristics |
ACE2 Expression |
Reference |
|---|---|---|---|---|
|
Vero E6 |
Kidney epithelial cells extracted from African green monkey (Chlorocebus sp.) |
Interferon-deficient (do not secrete IFNα or IFNβ when infected by viruses), non-tumorigenic, pseudodiploid karyotypes |
+++ |
|
|
Calu-3 |
Human lung adenocarcinoma |
Epithelial cells |
+ |
[36] |
|
A549 |
Human lung adenocarcinoma |
Epithelium-like, hypotriproid, synthesizes comparably large amounts of lecithin |
− |
|
|
Caco-2 |
Human colorectal adenocarcinoma |
Epithelium-like, upon reaching confluence, the cells express characteristics of enterocytic differentiation; express heat stable enterotoxin and epidermal growth factor |
++ |
[36] |
|
HEK-293T |
Human embryonic kidney |
Epithelial cells, highly transfectable, contains the SV40 T-antigen, widely used for retroviral production, gene expression and protein production |
− |
[39] |
|
Huh-7 |
Human hepatocellular carcinoma |
Epithelial cells, tumorigenic |
+ |
[37] |
2.2. Primary Human Airway Epithelial Cell (hAEC) Air–Liquid Interface (ALI) Cultures—The Gold Standard
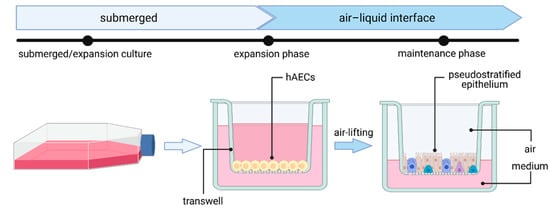
2.3. Lung Organoids—Innovative Technology
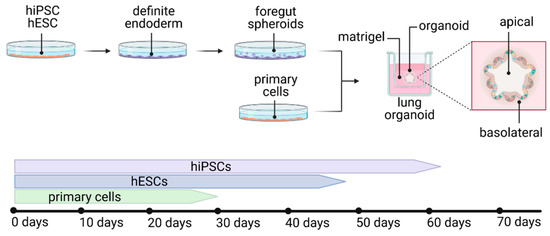
2.4. Lung-on-a-Chip—The Future?
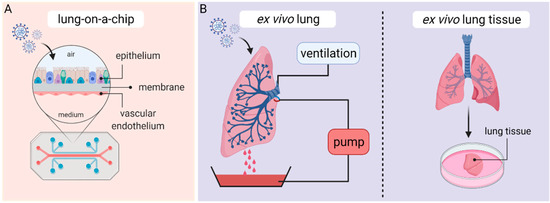
3. Ex Vivo Lung Perfusion (EVLP) Models and Human Lung Tissue Explants
|
Continuous Cell Lines |
hAEC-ALI Cultures |
Lung Organoids |
Lung-on-Chip |
EVLP/Lung Tissue Explants |
|
|---|---|---|---|---|---|
|
Availability |
+++ |
++ |
++ |
+ |
+ |
|
Affordability |
+++ |
++ |
+ |
+ |
− |
|
Authenticity/Physiological relevance |
+ |
+++ |
++ |
+++ |
+++ |
|
Handling |
+++ |
++ |
++ |
+ |
+ |
|
Reproducibility |
+++ |
++ |
+ |
+ |
− |
|
Genetic manipulation |
+++ |
+ |
++ |
+ |
− |
|
Application |
Virus propagation, diagnostics, high-throughput drug screenings, host–pathogen interactions |
Cell tropism, disease pathology immune response, transcriptomics, cell signaling, virus–host interactions, drug testing |
Cell tropism, disease pathology, transcriptomics, cell signaling, virus–host interactions, drug testing |
Immune and inflammatory responses, disease pathology, virus–host interactions, cross-talk, drug testing |
Drug testing, host–pathogen interactions, cellular tropism |
The symbols indicate high (+++), moderate (++), mild (+) and low (−) applicability.
This entry is adapted from the peer-reviewed paper 10.3390/v13050792
References
- Hsia, C.C.W.; Hyde, D.M.; Weibel, E.R. Lung Structure and the Intrinsic Challenges of Gas Exchange. Compr. Physiol. 2016, 6, 827–895.
- Crystal, R.G.; Randell, S.H.; Engelhardt, J.F.; Voynow, J.; Sunday, M.E. Airway epithelial cells: Current concepts and challenges. Proc. Am. Thorac. Soc. 2008, 5, 772–777.
- Hewitt, R.J.; Lloyd, C.M. Regulation of immune responses by the airway epithelial cell landscape. Nat. Rev. Immunol. 2021.
- Carcaterra, M.; Caruso, C. Alveolar epithelial cell type II as main target of SARS-CoV-2 virus and COVID-19 development via NF-Kb pathway deregulation: A physio-pathological theory. Med. Hypotheses 2021, 146, 110412.
- Ravindra, N.G.; Alfajaro, M.M.; Gasque, V.; Huston, N.C.; Wan, H.; Szigeti-Buck, K.; Yasumoto, Y.; Greaney, A.M.; Habet, V.; Chow, R.D.; et al. Single-cell longitudinal analysis of SARS-CoV-2 infection in human airway epithelium identifies target cells, alterations in gene expression, and cell state changes. PLoS Biol. 2021, 19, e3001143.
- Dijkman, R.; Jebbink, M.F.; Koekkoek, S.M.; Deijs, M.; Jónsdóttir, H.R.; Molenkamp, R.; Ieven, M.; Goossens, H.; Thiel, V.; van der Hoek, L. Isolation and characterization of current human coronavirus strains in primary human epithelial cell cultures reveal differences in target cell tropism. J. Virol. 2013, 87, 6081–6090.
- Vareille, M.; Kieninger, E.; Edwards, M.R.; Regamey, N. The airway epithelium: Soldier in the fight against respiratory viruses. Clin. Microbiol. Rev. 2011, 24, 210–229.
- The lungs at the frontlines of immunity. Nat. Immunol. 2015, 16, 17.
- Martin, T.R.; Frevert, C.W. Innate immunity in the lungs. Proc. Am. Thorac. Soc. 2005, 2, 403–411.
- Tyrrell, D.; Bynoe, M. Cultivation of viruses from a high proportion of patients with colds. Lancet 1966, 287, 76–77.
- Chan, J.F.-W.; To, K.K.-W.; Tse, H.; Jin, D.-Y.; Yuen, K.-Y. Interspecies transmission and emergence of novel viruses: Lessons from bats and birds. Trends Microbiol. 2013, 21, 544–555.
- Fehr, A.R.; Perlman, S. Coronaviruses: An overview of their replication and pathogenesis. Methods Mol. Biol. 2015, 1282, 1–23.
- Chan-Yeung, M.; Xu, R.-H. SARS: Epidemiology. Respirology 2003, 8, S9–S14.
- Zaki, A.M.; van Boheemen, S.; Bestebroer, T.M.; Osterhaus, A.D.M.E.; Fouchier, R.A.M. Isolation of a novel coronavirus from a man with pneumonia in Saudi Arabia. N. Engl. J. Med. 2012, 367, 1814–1820.
- Wu, F.; Zhao, S.; Yu, B.; Chen, Y.-M.; Wang, W.; Song, Z.-G.; Hu, Y.; Tao, Z.-W.; Tian, J.-H.; Pei, Y.-Y.; et al. A new coronavirus associated with human respiratory disease in China. Nature 2020, 579, 265–269.
- Zhou, P.; Yang, X.-L.; Wang, X.-G.; Hu, B.; Zhang, L.; Zhang, W.; Si, H.-R.; Zhu, Y.; Li, B.; Huang, C.-L.; et al. A pneumonia outbreak associated with a new coronavirus of probable bat origin. Nature 2020, 579, 270–273.
- WHO. WHO Coronavirus (COVID-19) Dashboard. Available online: (accessed on 12 March 2021).
- Wang, D.; Hu, B.; Hu, C.; Zhu, F.; Liu, X.; Zhang, J.; Wang, B.; Xiang, H.; Cheng, Z.; Xiong, Y.; et al. Clinical Characteristics of 138 Hospitalized Patients With 2019 Novel Coronavirus-Infected Pneumonia in Wuhan, China. JAMA 2020, 323, 1061–1069.
- Huang, C.; Wang, Y.; Li, X.; Ren, L.; Zhao, J.; Hu, Y.; Zhang, L.; Fan, G.; Xu, J.; Gu, X.; et al. Clinical features of patients infected with 2019 novel coronavirus in Wuhan, China. Lancet 2020, 395, 497–506.
- Song, P.; Li, W.; Xie, J.; Hou, Y.; You, C. Cytokine storm induced by SARS-CoV-2. Clin. Chim. Acta 2020, 509, 280–287.
- Wang, M.-Y.; Zhao, R.; Gao, L.-J.; Gao, X.-F.; Wang, D.-P.; Cao, J.-M. SARS-CoV-2: Structure, Biology, and Structure-Based Therapeutics Development. Front. Cell. Infect. Microbiol. 2020, 10, 587269.
- Guo, G.; Ye, L.; Pan, K.; Chen, Y.; Xing, D.; Yan, K.; Chen, Z.; Ding, N.; Li, W.; Huang, H.; et al. New Insights of Emerging SARS-CoV-2: Epidemiology, Etiology, Clinical Features, Clinical Treatment, and Prevention. Front. Cell Dev. Biol. 2020, 8, 410.
- Scialo, F.; Daniele, A.; Amato, F.; Pastore, L.; Matera, M.G.; Cazzola, M.; Castaldo, G.; Bianco, A. ACE2: The Major Cell Entry Receptor for SARS-CoV-2. Lung 2020, 198, 867–877.
- Hoffmann, M.; Kleine-Weber, H.; Schroeder, S.; Krüger, N.; Herrler, T.; Erichsen, S.; Schiergens, T.S.; Herrler, G.; Wu, N.-H.; Nitsche, A.; et al. SARS-CoV-2 Cell Entry Depends on ACE2 and TMPRSS2 and Is Blocked by a Clinically Proven Protease Inhibitor. Cell 2020, 181, 271–280.e8.
- WHO. Draft Landscape and Tracker of COVID-19 Candidate Vaccines. Available online: (accessed on 12 March 2021).
- Polack, F.P.; Thomas, S.J.; Kitchin, N.; Absalon, J.; Gurtman, A.; Lockhart, S.; Perez, J.L.; Pérez Marc, G.; Moreira, E.D.; Zerbini, C.; et al. Safety and Efficacy of the BNT162b2 mRNA Covid-19 Vaccine. N. Engl. J. Med. 2020, 383, 2603–2615.
- Voysey, M.; Costa Clemens, S.A.; Madhi, S.A.; Weckx, L.Y.; Folegatti, P.M.; Aley, P.K.; Angus, B.; Baillie, V.L.; Barnabas, S.L.; Bhorat, Q.E.; et al. Single-dose administration and the influence of the timing of the booster dose on immunogenicity and efficacy of ChAdOx1 nCoV-19 (AZD1222) vaccine: A pooled analysis of four randomised trials. Lancet 2021, 397, 881–891.
- Baden, L.R.; El Sahly, H.M.; Essink, B.; Kotloff, K.; Frey, S.; Novak, R.; Diemert, D.; Spector, S.A.; Rouphael, N.; Creech, C.B.; et al. Efficacy and Safety of the mRNA-1273 SARS-CoV-2 Vaccine. N. Engl. J. Med. 2021, 384, 403–416.
- WHO. Therapeutics and COVID-19. Available online: (accessed on 12 March 2021).
- Bao, L.; Deng, W.; Gao, H.; Xiao, C.; Liu, J.; Xue, J.; Lv, Q.; Liu, J.; Yu, P.; Xu, Y.; et al. Lack of Reinfection in Rhesus Macaques Infected with SARS-CoV-2. bioRxiv 2020.
- Bao, L.; Deng, W.; Huang, B.; Gao, H.; Liu, J.; Ren, L.; Wei, Q.; Yu, P.; Xu, Y.; Qi, F.; et al. The pathogenicity of SARS-CoV-2 in hACE2 transgenic mice. Nature 2020, 583, 830–833.
- Chan, J.F.-W.; Zhang, A.J.; Yuan, S.; Poon, V.K.-M.; Chan, C.C.-S.; Lee, A.C.-Y.; Chan, W.-M.; Fan, Z.; Tsoi, H.-W.; Wen, L.; et al. Simulation of the Clinical and Pathological Manifestations of Coronavirus Disease 2019 (COVID-19) in a Golden Syrian Hamster Model: Implications for Disease Pathogenesis and Transmissibility. Clin. Infect. Dis. 2020, 71, 2428–2446.
- Johansen, M.D.; Irving, A.; Montagutelli, X.; Tate, M.D.; Rudloff, I.; Nold, M.F.; Hansbro, N.G.; Kim, R.Y.; Donovan, C.; Liu, G.; et al. Animal and translational models of SARS-CoV-2 infection and COVID-19. Mucosal Immunol. 2020, 13, 877–891.
- Leland, D.S.; Ginocchio, C.C. Role of cell culture for virus detection in the age of technology. Clin. Microbiol. Rev. 2007, 20, 49–78.
- Dolskiy, A.A.; Grishchenko, I.V.; Yudkin, D.V. Cell Cultures for Virology: Usability, Advantages, and Prospects. Int. J. Mol. Sci. 2020, 21, 7978.
- Ren, X.; Glende, J.; Al-Falah, M.; de Vries, V.; Schwegmann-Wessels, C.; Qu, X.; Tan, L.; Tschernig, T.; Deng, H.; Naim, H.Y.; et al. Analysis of ACE2 in polarized epithelial cells: Surface expression and function as receptor for severe acute respiratory syndrome-associated coronavirus. J. Gen. Virol. 2006, 87, 1691–1695.
- Nie, Y.; Wang, P.; Shi, X.; Wang, G.; Chen, J.; Zheng, A.; Wang, W.; Wang, Z.; Qu, X.; Luo, M.; et al. Highly infectious SARS-CoV pseudotyped virus reveals the cell tropism and its correlation with receptor expression. Biochem. Biophys. Res. Commun. 2004, 321, 994–1000.
- Giard, D.J.; Aaronson, S.A.; Todaro, G.J.; Arnstein, P.; Kersey, J.H.; Dosik, H.; Parks, W.P. In vitro cultivation of human tumors: Establishment of cell lines derived from a series of solid tumors. J. Natl. Cancer Inst. 1973, 51, 1417–1423.
- Lambert, D.W.; Yarski, M.; Warner, F.J.; Thornhill, P.; Parkin, E.T.; Smith, A.I.; Hooper, N.M.; Turner, A.J. Tumor necrosis factor-alpha convertase (ADAM17) mediates regulated ectodomain shedding of the severe-acute respiratory syndrome-coronavirus (SARS-CoV) receptor, angiotensin-converting enzyme-2 (ACE2). J. Biol. Chem. 2005, 280, 30113–30119.
- Chu, H.; Chan, J.F.-W.; Yuen, T.T.-T.; Shuai, H.; Yuan, S.; Wang, Y.; Hu, B.; Yip, C.C.-Y.; Tsang, J.O.-L.; Huang, X.; et al. Comparative tropism, replication kinetics, and cell damage profiling of SARS-CoV-2 and SARS-CoV with implications for clinical manifestations, transmissibility, and laboratory studies of COVID-19: An observational study. Lancet Microbe 2020, 1, e14–e23.
- Harcourt, J.; Tamin, A.; Lu, X.; Kamili, S.; Sakthivel, S.K.; Murray, J.; Queen, K.; Tao, Y.; Paden, C.R.; Zhang, J.; et al. Severe Acute Respiratory Syndrome Coronavirus 2 from Patient with Coronavirus Disease, United States. Emerg. Infect. Dis. 2020, 26, 1266–1273.
- Ogando, N.S.; Dalebout, T.J.; Zevenhoven-Dobbe, J.C.; Limpens, R.W.A.L.; van der Meer, Y.; Caly, L.; Druce, J.; de Vries, J.J.C.; Kikkert, M.; Bárcena, M.; et al. SARS-coronavirus-2 replication in Vero E6 cells: Replication kinetics, rapid adaptation and cytopathology. J. Gen. Virol. 2020, 101, 925–940.
- Blanco-Melo, D.; Nilsson-Payant, B.E.; Liu, W.-C.; Uhl, S.; Hoagland, D.; Møller, R.; Jordan, T.X.; Oishi, K.; Panis, M.; Sachs, D.; et al. Imbalanced Host Response to SARS-CoV-2 Drives Development of COVID-19. Cell 2020, 181, 1036–1045.e9.
- Riva, L.; Yuan, S.; Yin, X.; Martin-Sancho, L.; Matsunaga, N.; Pache, L.; Burgstaller-Muehlbacher, S.; de Jesus, P.D.; Teriete, P.; Hull, M.V.; et al. Discovery of SARS-CoV-2 antiviral drugs through large-scale compound repurposing. Nature 2020, 586, 113–119.
- Touret, F.; Gilles, M.; Barral, K.; Nougairède, A.; van Helden, J.; Decroly, E.; de Lamballerie, X.; Coutard, B. In vitro screening of a FDA approved chemical library reveals potential inhibitors of SARS-CoV-2 replication. Sci. Rep. 2020, 10, 13093.
- Wang, M.; Cao, R.; Zhang, L.; Yang, X.; Liu, J.; Xu, M.; Shi, Z.; Hu, Z.; Zhong, W.; Xiao, G. Remdesivir and chloroquine effectively inhibit the recently emerged novel coronavirus (2019-nCoV) in vitro. Cell Res. 2020, 30, 269–271.
- Liu, J.; Cao, R.; Xu, M.; Wang, X.; Zhang, H.; Hu, H.; Li, Y.; Hu, Z.; Zhong, W.; Wang, M. Hydroxychloroquine, a less toxic derivative of chloroquine, is effective in inhibiting SARS-CoV-2 infection in vitro. Cell Discov. 2020, 6, 16.
- Pruijssers, A.J.; George, A.S.; Schäfer, A.; Leist, S.R.; Gralinksi, L.E.; Dinnon, K.H.; Yount, B.L.; Agostini, M.L.; Stevens, L.J.; Chappell, J.D.; et al. Remdesivir Inhibits SARS-CoV-2 in Human Lung Cells and Chimeric SARS-CoV Expressing the SARS-CoV-2 RNA Polymerase in Mice. Cell Rep. 2020, 32, 107940.
- Zettl, F.; Meister, T.L.; Vollmer, T.; Fischer, B.; Steinmann, J.; Krawczyk, A.; V’kovski, P.; Todt, D.; Steinmann, E.; Pfaender, S.; et al. Rapid Quantification of SARS-CoV-2-Neutralizing Antibodies Using Propagation-Defective Vesicular Stomatitis Virus Pseudotypes. Vaccines 2020, 8, 386.
- Luchsinger, L.L.; Ransegnola, B.P.; Jin, D.K.; Muecksch, F.; Weisblum, Y.; Bao, W.; George, P.J.; Rodriguez, M.; Tricoche, N.; Schmidt, F.; et al. Serological Assays Estimate Highly Variable SARS-CoV-2 Neutralizing Antibody Activity in Recovered COVID-19 Patients. J. Clin. Microbiol. 2020, 58.
- Puchelle, E.; Zahm, J.-M.; Tournier, J.-M.; Coraux, C. Airway epithelial repair, regeneration, and remodeling after injury in chronic obstructive pulmonary disease. Proc. Am. Thorac. Soc. 2006, 3, 726–733.
- Rawlins, E.L.; Ostrowski, L.E.; Randell, S.H.; Hogan, B.L.M. Lung development and repair: Contribution of the ciliated lineage. Proc. Natl. Acad. Sci. USA 2007, 104, 410–417.
- Comer, D.M.; Kidney, J.C.; Ennis, M.; Elborn, J.S. Airway epithelial cell apoptosis and inflammation in COPD, smokers and nonsmokers. Eur. Respir. J. 2013, 41, 1058–1067.
- Awatade, N.T.; Wong, S.L.; Hewson, C.K.; Fawcett, L.K.; Kicic, A.; Jaffe, A.; Waters, S.A. Human Primary Epithelial Cell Models: Promising Tools in the Era of Cystic Fibrosis Personalized Medicine. Front. Pharmacol. 2018, 9, 1429.
- Mulay, A.; Konda, B.; Garcia, G.; Yao, C.; Beil, S.; Sen, C.; Purkayastha, A.; Kolls, J.K.; Pociask, D.A.; Pessina, P.; et al. SARS-CoV-2 infection of primary human lung epithelium for COVID-19 modeling and drug discovery. bioRxiv 2020.
- Jonsdottir, H.R.; Dijkman, R. Coronaviruses and the human airway: A universal system for virus-host interaction studies. Virol. J. 2016, 13, 24.
- Hao, S.; Ning, K.; Kuz, C.A.; Vorhies, K.; Yan, Z.; Qiu, J. Long-Term Modeling of SARS-CoV-2 Infection of In Vitro Cultured Polarized Human Airway Epithelium. mBio 2020, 11.
- Sims, A.C.; Baric, R.S.; Yount, B.; Burkett, S.E.; Collins, P.L.; Pickles, R.J. Severe acute respiratory syndrome coronavirus infection of human ciliated airway epithelia: Role of ciliated cells in viral spread in the conducting airways of the lungs. J. Virol. 2005, 79, 15511–15524.
- Sims, A.C.; Burkett, S.E.; Yount, B.; Pickles, R.J. SARS-CoV replication and pathogenesis in an in vitro model of the human conducting airway epithelium. Virus Res. 2008, 133, 33–44.
- Banach, B.S.; Orenstein, J.M.; Fox, L.M.; Randell, S.H.; Rowley, A.H.; Baker, S.C. Human airway epithelial cell culture to identify new respiratory viruses: Coronavirus NL63 as a model. J. Virol. Methods 2009, 156, 19–26.
- Wang, G.; Deering, C.; Macke, M.; Shao, J.; Burns, R.; Blau, D.M.; Holmes, K.V.; Davidson, B.L.; Perlman, S.; McCray, P.B. Human coronavirus 229E infects polarized airway epithelia from the apical surface. J. Virol. 2000, 74, 9234–9239.
- Zhang, L.; Peeples, M.E.; Boucher, R.C.; Collins, P.L.; Pickles, R.J. Respiratory syncytial virus infection of human airway epithelial cells is polarized, specific to ciliated cells, and without obvious cytopathology. J. Virol. 2002, 76, 5654–5666.
- Wu, N.-H.; Yang, W.; Beineke, A.; Dijkman, R.; Matrosovich, M.; Baumgärtner, W.; Thiel, V.; Valentin-Weigand, P.; Meng, F.; Herrler, G. The differentiated airway epithelium infected by influenza viruses maintains the barrier function despite a dramatic loss of ciliated cells. Sci. Rep. 2016, 6, 39668.
- Gultom, M.; Laloli, L.; Dijkman, R. Well-Differentiated Primary Mammalian Airway Epithelial Cell Cultures. Methods Mol. Biol. 2020, 2203, 119–134.
- Kindler, E.; Jónsdóttir, H.R.; Muth, D.; Hamming, O.J.; Hartmann, R.; Rodriguez, R.; Geffers, R.; Fouchier, R.A.M.; Drosten, C.; Müller, M.A.; et al. Efficient replication of the novel human betacoronavirus EMC on primary human epithelium highlights its zoonotic potential. mBio 2013, 4, e00611-12.
- Sungnak, W.; Huang, N.; Bécavin, C.; Berg, M.; Queen, R.; Litvinukova, M.; Talavera-López, C.; Maatz, H.; Reichart, D.; Sampaziotis, F.; et al. SARS-CoV-2 entry factors are highly expressed in nasal epithelial cells together with innate immune genes. Nat. Med. 2020, 26, 681–687.
- Zhu, N.; Wang, W.; Liu, Z.; Liang, C.; Wang, W.; Ye, F.; Huang, B.; Zhao, L.; Wang, H.; Zhou, W.; et al. Morphogenesis and cytopathic effect of SARS-CoV-2 infection in human airway epithelial cells. Nat. Commun. 2020, 11, 3910.
- Pizzorno, A.; Padey, B.; Julien, T.; Trouillet-Assant, S.; Traversier, A.; Errazuriz-Cerda, E.; Fouret, J.; Dubois, J.; Gaymard, A.; Lescure, F.-X.; et al. Characterization and Treatment of SARS-CoV-2 in Nasal and Bronchial Human Airway Epithelia. Cell Rep. Med. 2020, 1, 100059.
- Sheahan, T.P.; Sims, A.C.; Zhou, S.; Graham, R.L.; Pruijssers, A.J.; Agostini, M.L.; Leist, S.R.; Schäfer, A.; Dinnon, K.H.; Stevens, L.J.; et al. An orally bioavailable broad-spectrum antiviral inhibits SARS-CoV-2 in human airway epithelial cell cultures and multiple coronaviruses in mice. Sci. Transl. Med. 2020, 12.
- Fiege, J.K.; Thiede, J.M.; Nanda, H.A.; Matchett, W.E.; Moore, P.J.; Montanari, N.R.; Thielen, B.K.; Daniel, J.; Stanley, E.; Hunter, R.C.; et al. Single cell resolution of SARS-CoV-2 tropism, antiviral responses, and susceptibility to therapies in primary human airway epithelium. PLoS Pathog. 2021, 17, e1009292.
- Salgado-Benvindo, C.; Thaler, M.; Tas, A.; Ogando, N.S.; Bredenbeek, P.J.; Ninaber, D.K.; Wang, Y.; Hiemstra, P.S.; Snijder, E.J.; van Hemert, M.J. Suramin Inhibits SARS-CoV-2 Infection in Cell Culture by Interfering with Early Steps of the Replication Cycle. Antimicrob. Agents Chemother. 2020, 64.
- Bojkova, D.; Bechtel, M.; McLaughlin, K.-M.; McGreig, J.E.; Klann, K.; Bellinghausen, C.; Rohde, G.; Jonigk, D.; Braubach, P.; Ciesek, S.; et al. Aprotinin Inhibits SARS-CoV-2 Replication. Cells 2020, 9, 2377.
- Leibel, S.L.; McVicar, R.N.; Winquist, A.M.; Niles, W.D.; Snyder, E.Y. Generation of Complete Multi-Cell Type Lung Organoids From Human Embryonic and Patient-Specific Induced Pluripotent Stem Cells for Infectious Disease Modeling and Therapeutics Validation. Curr. Protoc. Stem Cell Biol. 2020, 54, e118.
- Miller, A.J.; Dye, B.R.; Ferrer-Torres, D.; Hill, D.R.; Overeem, A.W.; Shea, L.D.; Spence, J.R. Generation of lung organoids from human pluripotent stem cells in vitro. Nat. Protoc. 2019, 14, 518–540.
- Sachs, N.; Papaspyropoulos, A.; Zomer-van Ommen, D.D.; Heo, I.; Böttinger, L.; Klay, D.; Weeber, F.; Huelsz-Prince, G.; Iakobachvili, N.; Amatngalim, G.D.; et al. Long-term expanding human airway organoids for disease modeling. EMBO J. 2019, 38.
- Suzuki, T.; Itoh, Y.; Sakai, Y.; Saito, A.; Okuzaki, D.; Motooka, D.; Minami, S.; Kobayashi, T.; Yamamoto, T.; Okamoto, T.; et al. Generation of human bronchial organoids for SARS-CoV-2 research. bioRxiv 2020.
- Pei, R.; Feng, J.; Zhang, Y.; Sun, H.; Li, L.; Yang, X.; He, J.; Xiao, S.; Xiong, J.; Lin, Y.; et al. Host metabolism dysregulation and cell tropism identification in human airway and alveolar organoids upon SARS-CoV-2 infection. Protein Cell 2020.
- Han, Y.; Duan, X.; Yang, L.; Nilsson-Payant, B.E.; Wang, P.; Duan, F.; Tang, X.; Yaron, T.M.; Zhang, T.; Uhl, S.; et al. Identification of SARS-CoV-2 inhibitors using lung and colonic organoids. Nature 2020.
- Katsura, H.; Sontake, V.; Tata, A.; Kobayashi, Y.; Edwards, C.E.; Heaton, B.E.; Konkimalla, A.; Asakura, T.; Mikami, Y.; Fritch, E.J.; et al. Human Lung Stem Cell-Based Alveolospheres Provide Insights into SARS-CoV-2-Mediated Interferon Responses and Pneumocyte Dysfunction. Cell Stem Cell 2020, 27, 890–904.e8.
- Youk, J.; Kim, T.; Evans, K.V.; Jeong, Y.-I.; Hur, Y.; Hong, S.P.; Kim, J.H.; Yi, K.; Kim, S.Y.; Na, K.J.; et al. Three-Dimensional Human Alveolar Stem Cell Culture Models Reveal Infection Response to SARS-CoV-2. Cell Stem Cell 2020, 27, 905–919.e10.
- Salahudeen, A.A.; Choi, S.S.; Rustagi, A.; Zhu, J.; van Unen, V.; Sean, M.d.l.O.; Flynn, R.A.; Margalef-Català, M.; Santos, A.J.M.; Ju, J.; et al. Progenitor identification and SARS-CoV-2 infection in human distal lung organoids. Nature 2020.
- Chen, Y.-W.; Huang, S.X.; de Carvalho, A.L.R.T.; Ho, S.-H.; Islam, M.N.; Volpi, S.; Notarangelo, L.D.; Ciancanelli, M.; Casanova, J.-L.; Bhattacharya, J.; et al. A three-dimensional model of human lung development and disease from pluripotent stem cells. Nat. Cell Biol. 2017, 19, 542–549.
- Dye, B.R.; Dedhia, P.H.; Miller, A.J.; Nagy, M.S.; White, E.S.; Shea, L.D.; Spence, J.R. A bioengineered niche promotes in vivo engraftment and maturation of pluripotent stem cell derived human lung organoids. eLife 2016, 5.
- Duffy, D.C.; McDonald, J.C.; Schueller, O.J.; Whitesides, G.M. Rapid Prototyping of Microfluidic Systems in Poly(dimethylsiloxane). Anal. Chem. 1998, 70, 4974–4984.
- Bhatia, S.N.; Ingber, D.E. Microfluidic organs-on-chips. Nat. Biotechnol. 2014, 32, 760–772.
- Tang, H.; Abouleila, Y.; Si, L.; Ortega-Prieto, A.M.; Mummery, C.L.; Ingber, D.E.; Mashaghi, A. Human Organs-on-Chips for Virology. Trends Microbiol. 2020, 28, 934–946.
- Si, L.; Prantil-Baun, R.; Benam, K.H.; Bai, H.; Rodas, M.; Burt, M.; Ingber, D.E. Discovery of influenza drug resistance mutations and host therapeutic targets using a human airway chip. bioRxiv 2019.
- Si, L.; Bai, H.; Rodas, M.; Cao, W.; Oh, C.Y.; Jiang, A.; Moller, R.; Hoagland, D.; Oishi, K.; Horiuchi, S.; et al. Human organ chip-enabled pipeline to rapidly repurpose therapeutics during viral pandemics. bioRxiv 2020.
- Fisher, A.; Andreasson, A.; Chrysos, A.; Lally, J.; Mamasoula, C.; Exley, C.; Wilkinson, J.; Qian, J.; Watson, G.; Lewington, O.; et al. An observational study of Donor Ex Vivo Lung Perfusion in UK lung transplantation: DEVELOP-UK. Health Technol. Assess. 2016, 20, 1–276.
- Steen, S.; Liao, Q.; Wierup, P.N.; Bolys, R.; Pierre, L.; Sjöberg, T. Transplantation of lungs from non–heart-beating donors after functional assessment ex vivo. Ann. Thorac. Surg. 2003, 76, 244–252.
- Ross, J.T.; Nesseler, N.; Lee, J.-W.; Ware, L.B.; Matthay, M.A. The ex vivo human lung: Research value for translational science. JCI Insight 2019, 4.
- Frank, J.A.; Briot, R.; Lee, J.W.; Ishizaka, A.; Uchida, T.; Matthay, M.A. Physiological and biochemical markers of alveolar epithelial barrier dysfunction in perfused human lungs. Am. J. Physiol. Lung Cell. Mol. Physiol. 2007, 293, L52–L59.
- Lee, J.W.; Krasnodembskaya, A.; McKenna, D.H.; Song, Y.; Abbott, J.; Matthay, M.A. Therapeutic Effects of Human Mesenchymal Stem Cells in Ex Vivo Human Lungs Injured with Live Bacteria. Am. J. Respir. Crit. Care Med. 2013, 187, 751–760.
- Park, J.; Kim, S.; Lim, H.; Liu, A.; Hu, S.; Lee, J.; Zhuo, H.; Hao, Q.; Matthay, M.A.; Lee, J.W. Therapeutic Effects of Human Mesenchymal Stem Cell Microvesicles in an Ex Vivo Perfused Human Lung Injured with Severe E.coli Pneumonia. Thorax 2018, 74, 43–50.
- Beck-Broichsitter, M.; Gauss, J.; Packhaeuser, C.B.; Lahnstein, K.; Schmehl, T.; Seeger, W.; Kissel, T.; Gessler, T. Pulmonary drug delivery with aerosolizable nanoparticles in an ex vivo lung model. Int. J. Pharm. 2009, 367, 169–178.
- Galasso, M.; Feld, J.J.; Watanabe, Y.; Pipkin, M.; Summers, C.; Ali, A.; Qaqish, R.; Chen, M.; Ribeiro, R.V.P.; Ramadan, K.; et al. Inactivating hepatitis C virus in donor lungs using light therapies during normothermic ex vivo lung perfusion. Nat. Commun. 2019, 10.
- Weathington, N.M.; Álvarez, D.; Sembrat, J.; Radder, J.; Cárdenes, N.; Noda, K.; Gong, Q.; Wong, H.; Kolls, J.; D’Cunha, J.; et al. Ex vivo lung perfusion as a human platform for preclinical small molecule testing. JCI Insight 2018, 3.
- Chen, M.; Rosenberg, J.; Cai, X.; Lee, A.C.H.; Shi, J.; Nguyen, M.; Wignakumar, T.; Mirle, V.; Edobor, A.J.; Fung, J.; et al. Nanotraps for the containment and clearance of SARS-CoV-2. bioRxiv 2021.
