In eukaryotes, DNA methylation refers to the addition of a methyl group to the fifth atom in the six-atom ring of cytosine residues (Cs). As a chemical modification, DNA methylation was discovered in 1948 by Rollin Hotchkiss, almost simultaneous to the identification of DNA as genetic material by Avery, MacLeold and McCarty. Almost thirty years later, in 1975, Holliday and Pugh proposed that DNA methylation is an important epigenetic mark [
33]. However, the mechanism by which DNA methylation was induced remained elusive for many years. In plants, DNA methylation was widely considered to be mediated by DNA–DNA interactions. However, in 1994, a study using viroid-infected tobacco plants showed that de novo DNA methylation is mediated by RNA molecules, thus aptly termed ‘RNA-directed DNA methylation’ (RdDM) [
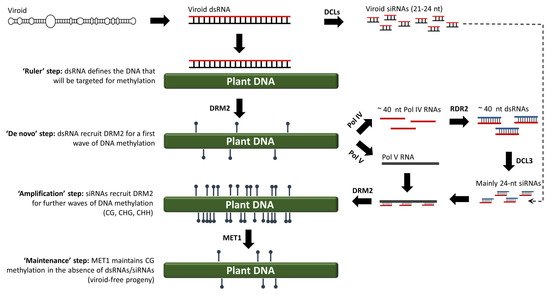
34]. Today, almost 30 years after the RdDM discovery, the widely accepted model suggests that 24-nucleotide siRNAs (canonical RdDM) or, in exceptional cases, 21/22-nucleotide siRNAs (non-canonical RdDM), are loaded onto AGO4 and guide domains-rearranged methyltransferases (DRMs) to methylate cognate DNA, most likely through a process wherein siRNAs directly interact with DNA or interact with the nascent transcripts produced by RNA polymerase V (POL V) [
35,
36,
37]. However, a growing body of evidence challenges the validity of this model. To begin with, it has been suggested that POL V is recruited to an already methylated DNA template, thus it can hardly be involved in the very first step of de novo methylation of a completely unmethylated DNA [
38]. More importantly, RdDM is not eliminated in an
Arabidopsis thaliana quadruple
dcl1 dcl2 dcl3 dcl4 mutant, suggesting that DCL-produced siRNAs are dispensable for RdDM [
39]. In addition, AGO4, which is thought to be involved in both canonical and non-canonical RdDM, is not always required for RdDM. AGO4 contains the DDH motif and has a slicer activity [
40]. During sense RdDM (S-RdDM), a single-stranded RNA is cleaved by the siRNAs loaded onto AGO4 and the cleaved transcripts are copied by RDRs into dsRNAs [
40]. However, during inverted repeat RdDM (IR-RdDM), the generation of a dsRNA would not rely on AGO4 activity, but on the mere transcription of hairpin RNA-producing DNA. Indeed, AGO4 is not required in IR-RdDM, at least not in
Arabidopsis thaliana [
41]. Collectively, these data suggest that RdDM is triggered not by siRNAs, but by long dsRNAs (>50 bp) [
42,
43,
44,
45]. According to our model, long dsRNAs appear to define the DNA region that will be methylated in a ruler-like fashion (Step 1, ). The dsRNA–DNA interaction may involve the formation of triple helices, as has been shown to take place in mammals between long non-coding RNAs (lncRNAs) and DNA [
46]. Such aberrant structures could attract DRMs to impose a first (perhaps incomplete) wave of de novo methylation on fully unmethylated DNA (Step 2, ). The resulting (and incompletely) methylated DNA seems to recruit POL IV and POL V. POL IV generates short transcripts (40–50 nt) that are copied by RDR2 into 40–50 bp dsRNAs that are cleaved by DCL3 into 24-nucleotide siRNAs [
47]. The AGO4-loaded 24-nucleotide siRNAs will now hybridize the newly produced POL V transcript, and this re-initiates the recruitment of DRMs to impose a second wave of methylation marks (Step 3, ) [
48,
49]. As siRNAs are amplified by this self-reinforcing loop mechanism, methylation will sequentially increase until nearly all of the Cs are methylated to CG, CHG and CHH sequence contexts in both of the DNA strands [
50]. Importantly, upon cell division and in the absence of dsRNAs/siRNAs, CG methylation can be maintained through the action of methyltransferase 1 (MET1) (Step 4, ) [
51]. To a lesser extent, CHG methylation will be maintained by chromomethylase 3 (CMT3) [
52]. However, CHH methylation cannot be maintained, as it always needs to be de novo established and requires the presence of dsRNAs/siRNAs [
45].
Over the last 30 years, the role of viroid-based systems in the elucidation of the RdDM mechanism and in refining corresponding models has been undisputable. Based on our current knowledge of RdDM and the above model, in the following, we will retrospectively navigate through the various stations of viroid-based RdDM, from 1994 to today.