Bi-dimensional culture systems have represented the most used method to study cell biology outside the body for over a century. Although they convey useful information, such systems may lose tissue-specific architecture, biomechanical effectors, and biochemical cues deriving from the native extracellular matrix, with significant alterations in several cellular functions and processes. Notably, the introduction of three-dimensional (3D) platforms that are able to re-create in vitro the structures of the native tissue, have overcome some of these issues, since they better mimic the in vivo milieu and reduce the gap between the cell culture ambient and the tissue environment. 3D culture systems are currently used in a broad range of studies, from cancer and stem cell biology, to drug testing and discovery.
- tissue engineering
- 3D matrices
- biomechanical cues
- microenvironment remodeling
- hydrogel
- micro-bioreactor
- 3D printing and bioprinting
- nanofiber-based scaffolds
- decellularization
- nanomedicine
1. Introduction
In vitro two-dimensional (2D) culture systems have represented, till recently, the most widely used strategy to study the mechanisms underlying cell biology, as well as diseases and drug action outside the body [1]. The first 2D approach was developed in 1907 by Harrison and colleagues to maintain nerve fibers in culture [2]. Subsequently, this method was applied for the study of different cell types, ranging from pluripotent and/or multipotent stem cells to terminally differentiated adult cells. The major advantages of 2D systems are associated with the simple and low-cost maintenance of cell cultures, the fast-downstream processing, and the easy performance of functional tests [1]. During the years, several steps forwards have been made to ameliorate and turn this technique into a more flexible and quicker platform. An example is represented by the modification and functionalization of the plastic surface onto which cells are seeded, using different materials and/or proteins, in order to resemble certain microenvironments [3]. Nevertheless, these improvements were not sufficient to mimic the natural structures of the original tissue and cells continued to be cultured in a monolayer bidirectional manner, providing sub-optimal conditions for their growth and specific functions. This negatively influences the fundamental cellular features and, usually, compromises the viability and reliability of the experiments, as well as the correct understanding of the whole organ activity [1][4]. Indeed, cells isolated from tissues and transferred onto flat and hard plastic substrates have been shown to lose their specific phenotype, because of the absence of a tissue-specific architecture and of the naturally occurring biomechanical and biochemical cues that cannot be mimicked in 2D in vitro systems. In particular, one of the most compromising aspects is represented by the alterations and/or the complete loss of the cell-to-cell and cell-to-extracellular matrix interactions. This leads to significant changes in several cellular features and processes, including cell morphology, polarity, differentiation, proliferation, genetic pattern, responsiveness to stimuli and secretions, drug metabolism, and many other functions [5][6][7][8][9].
The recent biotechnological advances have partly overcome these hurdles, thanks to the use of three-dimensional (3D) culture systems, which more closely mimic the natural in vivo milieu. The 3D culture methods are currently used in a broad range of in vitro studies, including cancer and stem cell biology, and drug testing and discovery. The first pioneering attempt to produce a 3D culture model dates back to the 1970s, when Hamburg and Salmon used a solution of soft agar to embed and grow human single cells [10]. Since then, several cutting-edge techniques were developed, ranging from hydrogels to organoid models and synthetic or biological scaffolds. The common key aspect of these new approaches is represented by their ability to in vitro recreate the macro- and micro-architecture of tissues and organs, encouraging cells to re-organize in complex 3D structures that closely resemble the in vivo micro-topography. The resulting in vitro environment replicates the biochemical and biomechanical effectors, directly influencing both cell fate and behavior.
2. Biomechanical Sensors and Effectors of the Microenvironment
The cell microenvironment is regulated by several factors, which include not only temperature, pH, oxygen, metabolites, growth factors or peptides, and hormones, but also macro- and micro-architecture-related cues as well as stretching and contracting stimuli. Externally applied forces are directly controlled by the stiffness of the substrate that the cells adhere to [11] and result in biomechanical signals that have a broad impact on cell behavior [12][13]. Cells are indeed able to sense the surrounding microenvironment and to interact with it, regulating their shape, their intracellular organization, growth, and differentiation, as well as their functionality. Interestingly enough, this ability is not limited to somatic cells and has emerged as an active property of oocytes and early embryos, which may actively sense biomechanical stimuli and convert them into intracellular signals, tuning their own behavior. In particular, in the context of the female gamete, the two main mechanosensing signaling pathways, namely Hippo and RhoGTPase, have been demonstrated to be involved in many fundamental oogenesis processes and to influence oocyte quality. Indeed, while the RhoGTPase signaling pathways are important actors in the coordination of actin filaments and microtubule formation, as well as polar body extrusion and spindle rotation during meiosis, the main actors of the Hippo pathway, Yes-associated protein (YAP) and WW domain-containing transcription regulator protein 1 (WWTR1 or TAZ), have an essential role during oogenesis, are highly transcribed in mouse and human oocytes, are maternally accumulated, and have been reported as strong candidates for zygotic genome activation [14].
Altogether, the properties described above are defined with the term “mechano-transduction”, which indicates the cellular mechanisms by which mechanical inputs, such as stretching, tension or fluid flow, are converted into intracellular signals [15]. They are regulated by several membrane proteins that modify their folding in response to extracellular biomechanical stimuli [16]. Integrins have been recognized as the main molecular link between cells and the extracellular matrix (ECM). This protein family is characterized by the ability to sense, respond to, and bidirectionally interact with the cellular environment [17]. More in detail, integrin’s key function is to trigger the cytoplasmatic recruitment and activation of adaptors and signaling proteins, such as Protein Tyrosine Kinase 2 (PTK2B, also known as FAK), SRC Proto-Oncogene Non-Receptor Tyrosine Kinase (SRC), and Mitogen-Activated Protein Kinase 1 (MAPK1, known as ERK), which assemble into the focal adhesion complex [18]. Force-mediated formation and/or reorganization of focal adhesion complexes, in turn, modify the Rho family small GTPase activities, leading to enhanced actin polymerization and the formation of stress fibers. This latter event can produce two different effects, described as short- or long-range transmission of force. The first directly affects the nuclear localization and function of the mechanosensitive transcription regulators, such as Myocardin Related Transcription Factor A (MRTFA, also known as MKL1) and YAP/TAZ [19]. The long-range effect transmits the force from the surrounding environment into the cell nucleus, thanks to a physical actin-mediated connection between the ECM and the linker of the nucleoskeleton and cytoskeleton (LINC) complex (Figure 1) [20]. This results in several changes, including chromatin remodeling, exposure of specific sites to transcription factors, gene expression changes, and the modifications of nuclear pore conformation and size, finally promoting YAP/TAZ nuclear translocation.
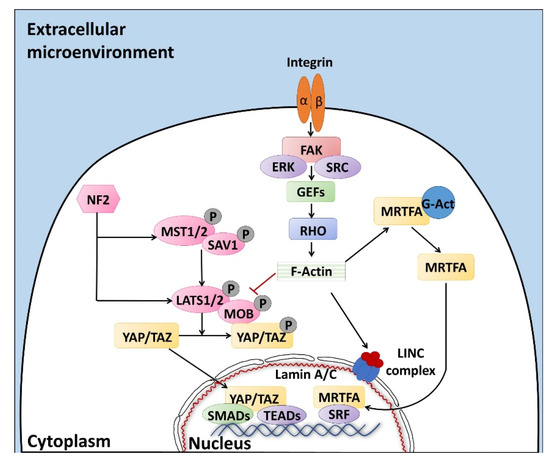
Figure 1. The sensors (integrins) and effectors (Rho and Hippo pathways) driving mechano-transduction processes, regulating YAP/TAZ localization and the subsequent cell behavior. Signaling pathways are described in detail in the text. Black arrow: activation, red T bar: inhibition of phosphorylation and molecule activation.
YAP/TAZ represent the main downstream effectors of the Hippo pathway cascade [21], which is involved in the cellular response to extracellular stimuli. Their compartmentalization is strictly correlated to mechanical signals, cellular stress, polarity, and adhesion cues [20], and influences gene expression, thus controlling cell fate. Recent studies also demonstrate the presence of upstream molecules, such as mammalian STE20-like protein kinase 1/2 (MST1/2), Salvador family WW domain containing protein 1 (SAV1), MOB kinase activator 1A/B (MOB1A/B), and large tumor suppressor 1/2 (LATS1/2), which regulate the Hippo pathway activity [22][23][24]. In particular, it has been shown that phosphorylation events are involved in the activation of the cascade, inducing cytoplasmatic retention of YAP/TAZ (Figure 1). In contrast, when the cascade is inactive, these two proteins translocate into the nucleus, interacting with different transcription factors (TFs). It is interesting to note that the YAP/TAZ subcellular localization is tightly controlled by substrate rigidity and topography [25][26][27], as well as by actin remodeling [28][29][30] and cell stretching [28]. This clearly demonstrates a specific role of these molecules as mechanotransducers and mechanosensors. In agreement with this, cells grown on a 3D soft matrix have been demonstrated to display YAP/TAZ localized to the cytoplasm, whereas cells plated on standard plastic supports (2D system) showed YAP/TAZ localized to the nucleus [26]. In addition, substrate stiffness is able to modulate the expression of the two molecules, which are upregulated in cells grown on hard hydrogels compared to soft substrates [31].
Furthermore, we recently demonstrated that YAP/TAZ localization is mirrored by a parallel compartmentalization of SMAD Family Member 2/3 (SMAD2/3). In particular, we observed that cells with cytoplasmic retention of YAP/TAZ exhibited a SMAD2/3 cytoplasmic distribution, while cells displaying the two molecules localized in the nucleus showed concomitant SMAD2/3 nuclear accumulation [23]. This latter event led to the formation of the YAP/TAZ–SMAD2/3 complex, which is known to bind to TEA domain transcription factors (TEAD), regulating cell fate and differentiation processes [32], both in bi-parental and mono-parental parthenogenetic cell lines [33][34][35].
It is indeed important to note that YAP/TAZ are transcriptional coactivators, unable to directly interact with DNA, but rather binding other TFs in order to elicit their functions [36]. Accumulating evidences point, in particular, to the TEAD protein family as the main actor, mediating YAP and TAZ nuclear activity and modulating the expression of different target genes involved in cell growth, proliferation, and organ development [23][37].
3. D Cell Culture Systems
The cellular microenvironment is a highly complex 3D natural structure, regulating in vivo cell functional activities through specific stimuli [38][39][40][41]. As a consequence, the organization and composition of the ECM, as well as the contribution of cell-to-cell and cell-to-ECM interactions, play a fundamental role in the control of cell behaviors and are strictly tied to the functions of tissues and whole organs.
The major aim of the recently introduced 3D cell culture systems is to model and in vitro re-create the tissue and organ in vivo milieu, mimicking the underlying biochemical and biomechanical signals [42][43]. A well-selected and -designed microenvironment can be therefore used in tissue and cell engineering to faithfully re-produce in vitro the conditions that promote all cell functions, including proliferation, migration, matrix production, and stem cell differentiation.
4. Tissue and Organ Regeneration Approaches
The first reports describing in vitro skin reconstruction date back to over 30 years ago [44][45][46][47], while the first bio-artificial bone tissue was developed in the 1990s by Ishaug et al. [48] and Baksh et al. [49]. Following these pioneering studies, extraordinary progress has been made in tissue engineering, not only for the optimization of scaffold production processes, but also for the regeneration and recellularization methods [50][51]. Indeed, cell repopulation plays a key role in getting fully functional bio-artificial organs. Needless to say, this procedure needs to be improved and optimized, and above all, adapted to the specific biophysical and biomechanical proprieties of the scaffold used as well as to the complexity of the tissue or organ of interest. Indeed, each organ varies in its unique structural components, namely, its cell types, matrix, architecture, as well as in its biophysical and biochemical environment, such as pressure, flow, oxygen tension, cytokines, and growth factors. As a consequence, the repopulation step requires the use of appropriate specific cell types, while commanding an optimized seeding method and a physiologically relevant culture approach (Figure 2). Skin cell sheet or blood vessel in vitro re-creation, for example, requires a single cell type, whereas whole organ reconstruction necessitates the seeding of multiple cell types to reestablish all the functional structures, including parenchyma, vasculature, and supporting components.
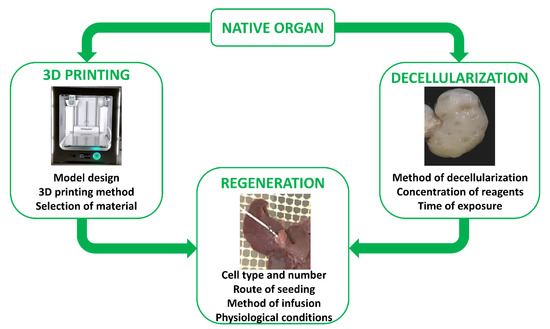
Figure 2. The two main ways and essential key aspects to successfully recreate in vitro a fully functional bio-artificial organ.
To date, distinct cell sources, such as embryonic stem cells, mesenchymal stem cells, fetal and adult tissue cells, and induced pluripotent cells, have been used to repopulate both artificial scaffolds as well as biological ones. The first tissue-engineered airway was transplanted into a patient in 2008. In this work, the authors repopulated a trachea scaffold with epithelial cells and mesenchymal stem-cell-derived chondrocytes. The results obtained demonstrated that it was possible to in vitro re-create a tissue-engineered trachea with mechanical properties that allow normal functioning, providing successful treatment for patients [52]. Following this, many studies using a single type of tissue-specific stem cells [53][54][55][56][57][58][59] or induced pluripotent cells [59][60][61][62] were carried out, and others are currently ongoing, in order to identify the best 3D supports able to boost cell differentiation and successfully produce in vitro functional tissues and organs for future possible transplantation. In parallel, both synthetic and biological scaffolds were repopulated with terminally differentiated cells, including fibroblasts [63], mature hepatocytes [64][65][66], and pancreatic endocrine [67][68][69], cardiac [70], and ovarian cells [71].
Nevertheless, it is still difficult to identify and standardize common protocols indicating routes of delivery, methods of infusion, and the correct cell number to be applied for the generation of functional organs. The main limitations are currently related to the complexity and the specialization of tissues, and, in some instances, to the elevated experimental costs. Indeed, although, different studies have already described distinct approaches obtaining encouraging results, none of them have proved their absolute potential to replace a damaged organ or tissue and several issues need to be solved.
In the kidney, a variety of successful strategies were developed [72]. However, the current techniques still show important limitations that impede the obtainment of a complete and functional whole-organ. In particular, while a piecemeal of renal components were reconstructed in vitro, the nephron structure has not yet been recreated [73].
In contrast, the liver represents a simpler, more feasible model, thanks to its regular structure and clear cellular organization. The current technical approaches in liver tissue engineering are based on the use of different methods, such as cell encapsulation, 3D printing, 3D bioprinting, and decellularized organs, which display the appropriate topography and biomechanical properties that facilitate hepatocyte colonization, migration, differentiation, proliferation, and cell polarity [74][75][76]. It is, however, important to highlight that, to date, 3D liver tissue organization and function has been mostly demonstrated in animal models and further studies and evidence are necessary in order to confirm the generation of a functional liver for human transplantation.
Many strategies were also developed for 3D heart recreation. Nevertheless, they led to the production of functionally immature contractile cardiac constructs [77][78][79]. Furthermore, although cardiomyocytes represent the key cell type of this tissue, the engrafting of stromal and endothelial populations, for the recreation of vascular system, remains the most immediate need for obtaining functional organs [73].
Lastly, the lung represents an extraordinary challenge to engineer almost from scratch. The studies carried out up to now demonstrated the recreation of some microscopic aspects of alveolar organization, without any gas exchange [80][81][82]. This organ, in fact, displays a unique and complicated architecture and is composed of more than 40 different highly specialized cell types. Furthermore, the peculiar structure of each capillary vessel, surrounded on both sides by a one cell-thick epithelium, which is needed to ensure carbon dioxide–oxygen exchange, represents the greatest obstacle for successful lung tissue engineering [80].
This entry is adapted from the peer-reviewed paper 10.3390/ijms22020830
References
- Kapałczyńska, M.; Kolenda, T.; Przybyła, W.; Zajączkowska, M.; Teresiak, A.; Filas, V.; Ibbs, M.; Bliźniak, R.; Łuczewski, Ł.; Lamperska, K. 2D and 3D cell cultures—A comparison of different types of cancer cell cultures. Arch. Med. Sci. 2018, 14, 910–919.
- Harrison, R.G.; Greenman, M.J.; Mall, F.P.; Jackson, C.M. Observations of the living developing nerve fiber. Anat. Rec. 1907, 1, 116–128.
- Langhans, S.A. Three-dimensional in vitro cell culture models in drug discovery and drug repositioning. Front. Pharmacol. 2018, 9, 6.
- Duval, K.; Grover, H.; Han, L.H.; Mou, Y.; Pegoraro, A.F.; Fredberg, J.; Chen, Z. Modeling physiological events in 2D vs. 3D cell culture. Physiology 2017, 32, 266–277.
- Baker, B.M.; Chen, C.S. Deconstructing the third dimension—How 3D culture microenvironments alter cellular cues. J. Cell Sci. 2012, 125, 3015–3024.
- Bissell, M.J.; Rizki, A.; Mian, I.S. Tissue architecture: The ultimate regulator of breast epithelial function. Curr. Opin. Cell Biol. 2003, 15, 753–762.
- Hickman, J.A.; Graeser, R.; de Hoogt, R.; Vidic, S.; Brito, C.; Gutekunst, M.; van der Kuip, H. IMI PREDECT consortium Three-dimensional models of cancer for pharmacology and cancer cell biology: Capturing tumor complexity in vitro/ex vivo. Biotechnol. J. 2014, 9, 1115–1128.
- Kleinman, H.K.; Philp, D.; Hoffman, M.P. Role of the extracellular matrix in morphogenesis. Curr. Opin. Biotechnol. 2003, 14, 526–532.
- Pampaloni, F.; Reynaud, E.G.; Stelzer, E.H.K. The third dimension bridges the gap between cell culture and live tissue. Nat. Rev. Mol. Cell Biol. 2007, 8, 839–845.
- Hamburger, A.; Salmon, S. Primary bioassay of human tumor stem cells. Science 1977, 197, 461–463.
- Discher, D.E.; Janmey, P.; Wang, Y.L. Tissue cells feel and respond to the stiffness of their substrate. Science 2005, 310, 1139–1143.
- Chicurel, M.E.; Singer, R.H.; Meyer, C.J.; Ingber, D.E. Integrin binding and mechanical tension induce movement of mRNA and ribosomes to focal adhesions. Nature 1998, 392, 730–733.
- Wozniak, M.A.; Modzelewska, K.; Kwong, L.; Keely, P.J. Focal adhesion regulation of cell behavior. Biochim. Biophys. Acta Mol. Cell Res. 2004, 1692, 103–119.
- Pennarossa, G.; Gandolfi, F.; Brevini, T.A.L. Biomechanical signaling in oocytes and parthenogenetic cells. Front. Cell Dev. Biol. Under revision.
- Pruitt, B.L.; Dunn, A.R.; Weis, W.I.; Nelson, W.J. Mechano-Transduction: From Molecules to Tissues. PLoS Biol. 2014, 12, e1001996.
- Shen, Z.; Niethammer, P. A cellular sense of space and pressure. Science 2020, 370, 295–296.
- Kechagia, J.Z.; Ivaska, J.; Roca-Cusachs, P. Integrins as biomechanical sensors of the microenvironment. Nat. Rev. Mol. Cell Biol. 2019, 20, 457–473.
- Huveneers, S.; Danen, E.H.J. Adhesion signaling—Crosstalk between integrins, Src and Rho. J. Cell Sci. 2009, 122, 1059–1069.
- Trembley, M.A.; Quijada, P.; Agullo-Pascual, E.; Tylock, K.M.; Colpan, M.; Dirkx, R.A.; Myers, J.R.; Mickelsen, D.M.; De Mesy Bentley, K.; Rothenberg, E.; et al. Mechanosensitive gene regulation by myocardin-related transcription factors is required for cardiomyocyte integrity in load-induced ventricular hypertrophy. Circulation 2018, 138, 1864–1878.
- Martino, F.; Perestrelo, A.R.; Vinarský, V.; Pagliari, S.; Forte, G. Cellular Mechanotransduction: From Tension to Function. Front. Physiol. 2018, 9, 824.
- Varelas, X. The Hippo pathway effectors TAZ and YAP in development, homeostasis and disease. Development 2014, 141, 1614–1626.
- Hansen, C.G.; Moroishi, T.; Guan, K.-L. YAP and TAZ: A nexus for Hippo signaling and beyond. Trends Cell Biol. 2015, 25, 499–513.
- Pennarossa, G.; Paffoni, A.; Ragni, G.; Gandolfi, F.; Brevini, T.A.L. Rho Signaling-Directed YAP/TAZ Regulation Encourages 3D Spheroid Colony Formation and Boosts Plasticity of Parthenogenetic Stem Cells. Adv. Exp. Med. Biol. 2020, 1237, 49–60.
- Brevini, T.A.L.; Pennarossa, G.; Gandolfi, F. A 3D approach to reproduction. Theriogenology 2020.
- Halder, G.; Dupont, S.; Piccolo, S. Transduction of mechanical and cytoskeletal cues by YAP and TAZ. Nat. Rev. Mol. Cell Biol. 2012, 13, 591–600.
- Dupont, S.; Morsut, L.; Aragona, M.; Enzo, E.; Giulitti, S.; Cordenonsi, M.; Zanconato, F.; Le Digabel, J.; Forcato, M.; Bicciato, S.; et al. Role of YAP/TAZ in mechanotransduction. Nature 2011, 474, 179–183.
- Schroeder, M.C.; Halder, G. Regulation of the Hippo pathway by cell architecture and mechanical signals. Semin. Cell Dev. Biol. 2012, 23, 803–811.
- Aragona, M.; Panciera, T.; Manfrin, A.; Giulitti, S.; Michielin, F.; Elvassore, N.; Dupont, S.; Piccolo, S. A mechanical checkpoint controls multicellular growth through YAP/TAZ regulation by actin-processing factors. Cell 2013, 154, 1047–1059.
- Fernández, B.G.; Gaspar, P.; Brás-Pereira, C.; Jezowska, B.; Rebelo, S.R.; Janody, F. Actin-capping Protein and the Hippo pathway regulate F-actin and tissue growth in Drosophila. Development 2011, 138, 2337–2346.
- Calvo, F.; Ege, N.; Grande-Garcia, A.; Hooper, S.; Jenkins, R.P.; Chaudhry, S.I.; Harrington, K.; Williamson, P.; Moeendarbary, E.; Charras, G.; et al. Mechanotransduction and YAP-dependent matrix remodelling is required for the generation and maintenance of cancer-associated fibroblasts. Nat. Cell Biol. 2013, 15, 637–646.
- Thomasy, S.M.; Morgan, J.T.; Wood, J.A.; Murphy, C.J.; Russell, P. Substratum stiffness and latrunculin B modulate the gene expression of the mechanotransducers YAP and TAZ in human trabecular meshwork cells. Exp. Eye Res. 2013, 113, 66–73.
- Pennarossa, G.; Manzoni, E.F.M.E.F.M.; Ledda, S.; de Eguileor, M.; Gandolfi, F.; Brevini, T.A. Use of a PTFE Micro-Bioreactor to Promote 3D Cell Rearrangement and Maintain High Plasticity in Epigenetically Erased Fibroblasts. Stem Cell Rev. Rep. 2019, 15, 82–92.
- Brevini, T.A.L.; Pennarossa, G.; Maffei, S.; Gandolfi, F. Pluripotency network in porcine embryos and derived cell lines. Reprod. Domest. Anim. 2012, 47.
- Brevini, T.A.L.; Pennarossa, G.; Antonini, S.; Gandolfi, F. Parthenogenesis as an approach to pluripotency: Advantages and limitations involved. Stem Cell Rev. 2008, 4.
- Brevini, T.A.L.; Pennarossa, G.; Gandolfi, F. No shortcuts to pig embryonic stem cells. Theriogenology 2010, 74.
- Pan, D. The Hippo Signaling Pathway in Development and Cancer. Dev. Cell 2010, 19, 491–505.
- Holden, J.K.; Cunningham, C.N. Targeting the Hippo Pathway and Cancer through the TEAD Family of Transcription Factors. Cancers 2018, 10, 81.
- Burdick, J.A.; Vunjak-Novakovic, G. Engineered microenvironments for controlled stem cell differentiation. Tissue Eng. Part A 2009, 15, 205–219.
- Scadden, D.T. The stem-cell niche as an entity of action. Nature 2006, 441, 1075–1079.
- Underhill, G.H.; Bhatia, S.N. High-throughput analysis of signals regulating stem cell fate and function. Curr. Opin. Chem. Biol. 2007, 11, 357–366.
- Vining, K.H.; Mooney, D.J. Mechanical forces direct stem cell behaviour in development and regeneration. Nat. Rev. Mol. Cell Biol. 2017, 18, 728.
- Huh, D.; Hamilton, G.A.; Ingber, D.E. From 3D cell culture to organs-on-chips. Trends Cell Biol. 2011, 21, 745–754.
- Gu, L.; Mooney, D.J. Biomaterials and emerging anticancer therapeutics: Engineering the microenvironment. Nat. Rev. Cancer 2016, 16, 56–66.
- Burke, J.F.; Yannas, O.V.; Quinby, W.C.; Bondoc, C.C.; Jung, W.K. Successful use of a physiologically acceptable artificial skin in the treatment of extensive burn injury. Ann. Surg. 1981, 194, 413–427.
- Spira, M.; Fissette, J.; Hall, C.W.; Hardy, S.B.; Gerow, F.J. Evaluation of synthetic fabrics as artificial skin grafts to experimental burn wounds. J. Biomed. Mater. Res. 1969, 3, 213–234.
- Hall, C.W.; Liotta, D.; De Bakey, M.E. Artificial skin. Trans. Am. Soc. Artif. Intern. Organs 1966, 12, 340–343.
- O’Connor, N.E.; Mulliken, J.B.; Banks-Schlegel, S.; Kehinde, O.; Green, H. Grafting of Burns with Cultured Epithelium Prepared from Autologous Epidermal Cells. Lancet 1981, 317, 75–78.
- Ishaug, S.L.; Crane, G.M.; Miller, M.J.; Yasko, A.W.; Yaszemski, M.J.; Mikos, A.G. Bone formation by three-dimensional stromal osteoblast culture in biodegradable polymer scaffolds. J. Biomed. Mater. Res. 1997, 36, 17–28.
- Baksh, D.; Davies, J.E.; Kim, S. Three-dimensional matrices of calcium polyphosphates support bone growth in vitro and in vivo. J. Mater. Sci. Mater. Med. 1998, 9, 743–748.
- Khan, A.A.; Vishwakarma, S.K.; Bardia, A.; Venkateshwarulu, J. Repopulation of decellularized whole organ scaffold using stem cells: An emerging technology for the development of neo-organ. J. Artif. Organs 2014, 17, 291–300.
- Scarritt, M.E.; Pashos, N.C.; Bunnell, B.A.; Bussolati, B. A review of cellularization strategies for tissue engineering of whole organs. Front. Bioeng. Biotechnol. 2015, 3, 43.
- Macchiarini, P.; Jungebluth, P.; Go, T.; Asnaghi, M.A.; Rees, L.E.; Cogan, T.A.; Dodson, A.; Martorell, J.; Bellini, S.; Parnigotto, P.P.; et al. Clinical transplantation of a tissue-engineered airway. Lancet 2008, 372, 2023–2030.
- Pennarossa, G.; Ghiringhelli, M.; Gandolfi, F.; Brevini, T.A.L. Creation of a bioengineered ovary: Isolation of female germline stem cells for the repopulation of a decellularized ovarian bio-scaffold. Methods Mol. Biol. in press.
- Ghiringhelli, M.; Zenobi, A.; Brizzola, S.; Gandolfi, F.; Bontempo, V.; Rossi, S.; Brevini, T.A.L.; Acocella, F. Simple and quick method to obtain a decellularized, functional liver bioscaffold. Methods Mol. Biol. 2018, 1577, 283–292.
- Shackleton, M.; Vaillant, F.; Simpson, K.J.; Stingl, J.; Smyth, G.K.; Asselin-Labat, M.L.; Wu, L.; Lindeman, G.J.; Visvader, J.E. Generation of a functional mammary gland from a single stem cell. Nature 2006, 439, 84–88.
- Stingl, J.; Eirew, P.; Ricketson, I.; Shackleton, M.; Vaillant, F.; Choi, D.; Li, H.I.; Eaves, C.J. Purification and unique properties of mammary epithelial stem cells. Nature 2006, 439, 993–997.
- Schlieve, C.R.; Grikscheit, T.C. A Purpose in Liquidity: Perfusing 3D Open Scaffolds Improves “Mini-gut” Morphogenesis and Longevity. Cell Stem Cell 2020, 27, 699–701.
- Han, Y.; Baltriukienė, D.; Kozlova, E.N. Effect of scaffold properties on adhesion and maintenance of boundary cap neural crest stem cells in vitro. J. Biomed. Mater. Res. Part A 2020, 108, 1274–1280.
- Croce, S.; Peloso, A.; Zoro, T.; Avanzini, M.A.; Cobianchi, L. A hepatic scaffold from decellularized liver tissue: Food for thought. Biomolecules 2019, 9, 813.
- Santarella, F.; Sridharan, R.; Marinkovic, M.; Do Amaral, R.J.F.C.; Cavanagh, B.; Smith, A.; Kashpur, O.; Gerami-Naini, B.; Garlick, J.A.; O’Brien, F.J.; et al. Scaffolds Functionalized with Matrix from Induced Pluripotent Stem Cell Fibroblasts for Diabetic Wound Healing. Adv. Healthc. Mater. 2020, 9, 2000307.
- Murphy, A.R.; Haynes, J.M.; Laslett, A.L.; Cameron, N.R.; O’Brien, C.M. Three-dimensional differentiation of human pluripotent stem cell-derived neural precursor cells using tailored porous polymer scaffolds. Acta Biomater. 2020, 101, 102–116.
- Ding, M.; Andersson, H.; Martinsson, S.; Sabirsh, A.; Jonebring, A.; Wang, Q.D.; Plowright, A.T.; Drowley, L. Aligned nanofiber scaffolds improve functionality of cardiomyocytes differentiated from human induced pluripotent stem cell-derived cardiac progenitor cells. Sci. Rep. 2020, 10.
- Pennarossa, G.; Ghiringhelli, M.; Gandolfi, F.; Brevini, T.A.L. Whole-ovary decellularization generates an effective 3D bioscaffold for ovarian bioengineering. J. Assist. Reprod. Genet. 2020, 1–11.
- Nyberg, S.L.; Remmel, R.P.; Mann, H.J.; Peshwa, M.V.; Hu, W.S.; Cerra, F.B. Primary hepatocytes outperform Hep G2 cells as the source of biotransformation functions in a bioartificial liver. Ann. Surg. 1994, 220, 59–67.
- Nagaki, M.; Miki, K.; Kim, Y.I.; Ishiyama, H.; Hirahara, I.; Takahashi, H.; Sugiyama, A.; Muto, Y.; Moriwaki, H. Development and characterization of a hybrid bioartificial liver using primary hepatocytes entrapped in a basement membrane matrix. Dig. Dis. Sci. 2001, 46, 1046–1056.
- Hosseini, V.; Maroufi, N.F.; Saghati, S.; Asadi, N.; Darabi, M.; Ahmad, S.N.S.; Hosseinkhani, H.; Rahbarghazi, R. Current progress in hepatic tissue regeneration by tissue engineering. J. Transl. Med. 2019, 17, 383.
- De Carlo, E.; Baiguera, S.; Conconi, M.T.; Vigolo, S.; Grandi, C.; Lora, S.; Martini, C.; Maffei, P.; Tamagno, G.; Vettor, R.; et al. Pancreatic acellular matrix supports islet survival and function in a synthetic tubular device: In vitro and in vivo studies. Int. J. Mol. Med. 2010, 25, 195–202.
- Conrad, C.; Schuetz, C.; Clippinger, B.; Vacanti, J.P.; Markmann, J.F.; Ott, H.C. Bio-engineered endocrine pancreas based on decellularized pancreatic matrix and mesenchymal stem cell/islet cell coculture. J. Am. Coll. Surg. 2010, 211, S62.
- Peloso, A.; Citro, A.; Oldani, G.; Brambilla, S.; Piemonti, L.; Cobianchi, L. Bioengineering the Pancreas: Cell-on-Scaffold Technology. In Scaffolds in Tissue Engineering—Materials, Technologies and Clinical Applications; IntechOpen: London, UK, 2017.
- Silva, A.C.; Rodrigues, S.C.; Caldeira, J.; Nunes, A.M.; Sampaio-Pinto, V.; Resende, T.P.; Oliveira, M.J.; Barbosa, M.A.; Thorsteinsdóttir, S.; Nascimento, D.S.; et al. Three-dimensional scaffolds of fetal decellularized hearts exhibit enhanced potential to support cardiac cells in comparison to the adult. Biomaterials 2016, 104, 52–64.
- Laronda, M.M. Engineering a bioprosthetic ovary for fertility and hormone restoration. Theriogenology 2020.
- Wragg, N.M.; Burke, L.; Wilson, S.L. A critical review of current progress in 3D kidney biomanufacturing: Advances, challenges, and recommendations. Ren. Replace. Ther. 2019, 5, 1–16.
- Mandrycky, C.; Phong, K.; Zheng, Y. Tissue engineering toward organ-specific regeneration and disease modeling. MRS Commun. 2017, 7, 332–347.
- Mirdamadi, E.S.; Kalhori, D.; Zakeri, N.; Azarpira, N.; Solati-Hashjin, M. Liver tissue engineering as an emerging alternative for liver disease treatment. Tissue Eng. Part B Rev. 2020, 26, 145–163.
- Vo, T.N.; Chen, A.X.; Smith, Q.B.; Chhabra, A.; Bhatia, S.N. Integrated Technologies for Liver Tissue Engineering. In The Liver; Wiley: Hoboken, NJ, USA, 2020; pp. 1028–1035.
- Heydari, Z.; Najimi, M.; Mirzaei, H.; Shpichka, A.; Ruoss, M.; Farzaneh, Z.; Montazeri, L.; Piryaei, A.; Timashev, P.; Gramignoli, R.; et al. Tissue Engineering in Liver Regenerative Medicine: Insights into Novel Translational Technologies. Cells 2020, 9, 304.
- West, J.L. Biomaterials for Cardiovascular Tissue Engineering. In Biomaterials Science; Elsevier: Amsterdam, The Netherlands, 2020; pp. 1389–1397.
- Jang, Y.; Park, Y.; Kim, J. Engineering Biomaterials to Guide Heart Cells for Matured Cardiac Tissue. Coatings 2020, 10, 925.
- Bolonduro, O.A.; Duffy, B.M.; Rao, A.A.; Black, L.D.; Timko, B.P. From biomimicry to bioelectronics: Smart materials for cardiac tissue engineering. Nano Res. 2020, 13, 1253–1267.
- Tebyanian, H.; Karami, A.; Nourani, M.R.; Motavallian, E.; Barkhordari, A.; Yazdanian, M.; Seifalian, A. Lung tissue engineering: An update. J. Cell. Physiol. 2019, 234, 19256–19270.
- Calle, E.A.; Ghaedi, M.; Sundaram, S.; Sivarapatna, A.; Tseng, M.K.; Niklason, L.E. Strategies for whole lung tissue engineering. IEEE Trans. Biomed. Eng. 2014, 61, 1482–1496.
- Petersen, T.H.; Calle, E.A.; Zhao, L.; Lee, E.J.; Gui, L.; Raredon, M.S.B.; Gavrilov, K.; Yi, T.; Zhuang, Z.W.; Breuer, C.; et al. Tissue-engineered lungs for in vivo implantation. Science 2010, 329, 538–541.
