This a review that comprehensively covers the modalities that act as disruptors of the YAP-TEAD interaction. The transcriptional co-activator YAP (Yes-associated protein) by pairing with the transcription factor TEAD (TEA domain) orchestrates the expression of several oncogenic transcriptional programs. These programs are seen in a proportion of all solid tumors. Therefore, the disruption of YAP-TEAD interaction is proposed as an attractive option to target cancers.
- YAP-TEAD Interaction
1. Introduction
The TEADs are a family of four conserved proteins (TEAD 1-4 in humans) that have similar domain architecture (Figure 1A) [14,15]. All TEADs have an N-terminal TEA domain that binds to DNA and adopts a homeodomain fold. At the C-terminus, all TEADs have a transactivation domain that binds to YAP/TAZ. TEADs differ largely at the linker that connects the TEA domain with the transactivation domain. It was known early on that an N-terminal motif of YAP interacts with the TEAD transactivation domain (Figures 1A and 1B). However, the determination of the crystal structures of TEADs in complex with YAP was a breakthrough as it gave a clear, detailed picture of the interaction [11,16,17]. The TEAD transactivation domain adopts an immunoglobulin-like β-sandwich fold; additionally, it has two helix-turn-helix motifs. The N-terminal motif of YAP wraps around TEAD extensively and forms three interfaces (Figure 1B).
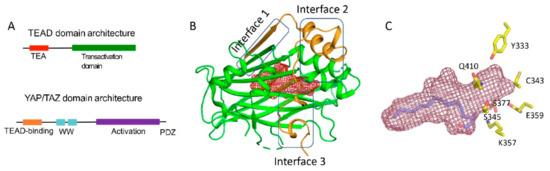
Figure 1. Domain architecture and structures of YAP/TAZ and TEAD (A) Domain architecture of YAP and TEAD (B) Ribbon diagram of the core complex structure of the YAP-TEAD complex (PDB ID: 3KYS). N-terminus of YAP interacts with the transactivation domain of TEAD by forming three interfaces. The central pocket is shown as red mesh. (C) Central pocket (red) and the orientation of the palmitate in the pocket is shown. The conserved polar/charged TEAD2 residues lining the central pocket is shown as yellow sticks.
2. Structural Characterization of YAP-TEAD Interaction
Interface 1 is an anti-parallel β-sheet where both YAP and TEAD contribute to a single β-strand. This interface contributes little to the affinity between YAP and TEAD. Although there are drugs that target higher-order oligomers of β-sheets such as the amyloid plaques formed in Alzheimer disease [18], a simple β-sheet as in interface 1 has poor ligand-binding ability. Interface 2 is formed when a helix of YAP/TAZ slots into a pocket formed by one of the helix-turn-helix motifs of TEAD (Figure 1B). Such a helix-pocket interaction commonly occurs between proteins and is usually mediated by leucine residues (the LxxLL motif) [19,20]. These three hydrophobic leucine residues that are oriented towards the pocket are the key contact residues. YAP and TAZ have a variant of this motif, they have L65xxL68F69 motif instead of LxxLL (the numbers correspond to human YAP residues). Peptides and small molecules have been shown to occupy the interface 2 pocket (Table 1). The YAP/TAZ helix on its own binds poorly to TEAD and interface 3 acts to increase its binding affinity. Interface 3 is the most crucial of the three interfaces [11,16,17]. At this interface, YAP adopts a three-dimensional structure resembling the Greek letter Ω (Figure 1B), and this conformation is generally referred to as the Ω-loop. Crucial for interface 3 interaction are three hydrophobic core residues -- in human YAP, these are M86, L91, and F95 -- that fit into a pocket on the TEAD surface. A cation-π interaction formed between R87 and F96 YAP residues is also crucial and appears to stabilize the Ω-loop conformation. Appreciable progress has been made on the peptidomimetics front, linear and cyclic peptides have been designed to act as PPIDs by occupying the interface 3 pocket on TEAD (Table 1). Interface 3 TEAD residues that interact with YAP are highly conserved, therefore it is difficult to design modalities at this interface that display selective TEAD inhibition. That said, it is not clear whether there is a biological need for selective TEAD inhibitors. In the field, researchers tend to work more on YAP than TAZ, but as YAP and TAZ have similar primary sequences and structural features, PPIDs that disrupt the YAP-TEAD interaction should also have the ability to disrupt the TAZ-TEAD interaction.
Table 1. Compounds that target TEAD surface pocket interfaces 2 and 3.
| No. | Molecule | Structure | Surface Pocket Validation | Molecule Type | Validation Method | Reference |
|---|---|---|---|---|---|---|
| Modalities binding to Interface 2 | ||||||
| 1. | TB2G1 | Cystine-dense peptide | Rosetta modeling | PPID | Co-IP assay | [21] |
| 2. | Fragment 1 | 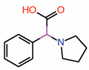 |
Crystal structure | PPID unlikely |
GST pull-down | [22] |
| 3. | Tri-substitutedpyrazoles | 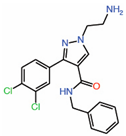 |
Crystal structure | Potential PPID |
||
| Modalities binding to Interface 3 | ||||||
| 4. | Peptide 17 | YAP cyclic peptide | Molecular modeling | PPID | GST pull-down | [23] |
| 5. | Peptide 10 | YAP cyclic peptide | Crystal structure |
PPID | GST pull-down | [24] |
| 6. | Peptides 9, 10 | YAP linear peptide | Crystal structure |
PPID | TR-FRET assay | [25] |
| 7. | Flufenamic acid | 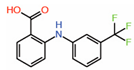 |
Crystal structure |
PPID unlikely |
[13] | |
| 8. | TEAD-binding fragment |
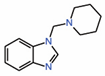 |
NMR | PPID unlikely |
[26] | |
| 9. | Dioxo-benzoisothiazole Example 22 |
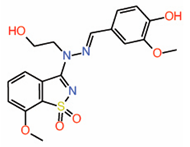 |
NMR | PPID | AlphaLISA assay |
[27] |
| 10. | Triazole carbohydrazides Hit 2 |
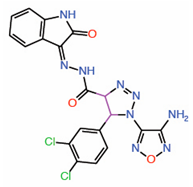 |
Molecular modeling |
Potential PPID |
[28] | |
| 11. | Compound 3.1 | 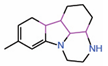 |
Molecular docking |
PPID | Co-IP assay | [29] |
| Peptide binding to both Interfaces 2 & 3 | ||||||
| 12. | Super-TDU | YAP-VGLL4 fusion peptide | Molecular modeling |
PPID | Co-IP assay | [30] |
Interestingly, Pobbati and colleagues identified a pocket with an appropriate geometry and hydrophobicity in the center of the transactivating domain of TEADs that is accessible to small molecules [13] (Figure 1B). This pocket is distinct from the TEAD surface pockets. One end of the pocket is solvent-exposed. However, it is blocked by the interface 1 β-strand of YAP that acts as a gate to ligand access. The other end of the central pocket extends into the domain interior. Palmitate is the natural ligand that binds to the central pocket [31,32]. In addition to the hydrophobic residues that line the TEAD central pocket, there are seven conserved charged/polar residues (Figure 1C). Remarkably, all seven residues are located at the solvent-exposed end of the pocket. Notably, several central pocket binders have been identified that, not only inhibit TEAD palmitoylation, but also disrupt YAP-TEAD interaction, thus, acting as allosteric PPIDs. Several other molecules bind the pocket but only act as palmitoylation inhibitors and not as allosteric PPIDs. Despite this, they inhibit TEAD activity and function (Table 2).
Table 2. Compounds that target the TEAD central pocket.
| No. | Molecule | Structure | Binding Validation | Molecule Type | Validation Method | Reference |
|---|---|---|---|---|---|---|
| Flufenamates | ||||||
| 1. | Flufenamic acid | 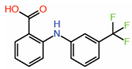 |
Crystal structure | Palmitoylation inhibitor |
Mass Spec analysis | [13] |
| 2. | TED-347 | 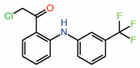 |
Crystal structure | Allosteric PPID |
FP assay with YAP peptide | [33] |
| 3. | MYF-01-037 | 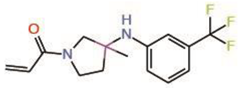 |
Molecular docking |
Allosteric PPID |
Split luciferase assay | [34] |
| 4. | Non-fused tricyclic Compound 42 | 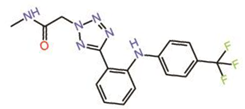 |
unknown | PPID? | [35] | |
| 5. | FA-Palmitate fusion | 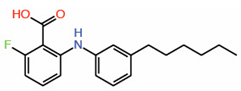 |
Crystal structure | PPID? | ||
| Non-covalent allosteric PPIDs | ||||||
| 6. | MGH-CP1 | 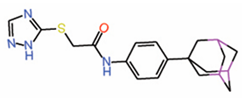 |
Crystal structure | Allosteric PPID |
Co-IP assay | [36] |
| 7. | Compound 9 Indole-focused | 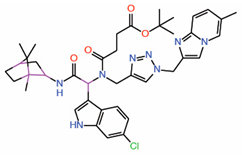 |
Palmitate-based FP assay | Allosteric PPID |
FP assay with YAP peptide | [37] |
| 8. | Dihydropyrazolo pyrimidines Compound 7 |
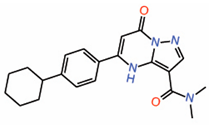 |
unknown | AllostericPPID? | TR-FRET assay | [38] |
| Covalent allosteric PPIDs | ||||||
| 9. | K-975 | 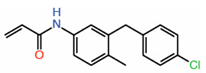 |
Crystal structure | Allosteric PPID |
Co-IP assay | [39] |
| 10. | Kojic acid analogs Compound 19 | 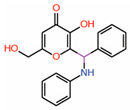 |
Thiol conjugation assay | Allosteric PPID |
FP assay with YAP peptide | [40] |
| Other central pocket binders | ||||||
| 11. | Compound 2 | 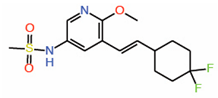 |
Crystal structure | Palmitoylation inhibitor | FP assay with palmitate | [41] |
| 12. | Vinylsulfonamide DC-TEADin02 | 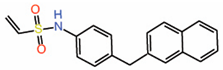 |
Molecular docking | Palmitoylation inhibitor | Competitive NMR |
[42] |
| 13. | Quinolinol Q2 | 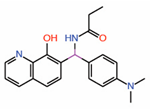 |
Molecular dynamics simulations |
TEAD activator | RNA-Seq | [43] |
This entry is adapted from the peer-reviewed paper 10.3390/molecules25246001
