Pseudomonas biofilms have been studied intensively for several decades and research outcomes have been successfully implemented in various medical and agricultural applications. Research on biofilm synthesis and composition has also overlapped with the objectives of environmental sciences, since biofilm components show exceptional physicochemical properties applicable to remediation techniques. Especially, exopolysaccharides (ExPs) have been at the center of scientific interest, indicating their potential in solving the environmental issues of heavy metal land and water contamination via sorptive interactions and flocculation.
- exopolysaccharides
- Pseudomonas
- biosorption
- bioremediation
1. Introduction
2. Biosynthesis of Extracellular Polysaccharides in Pseudomonas
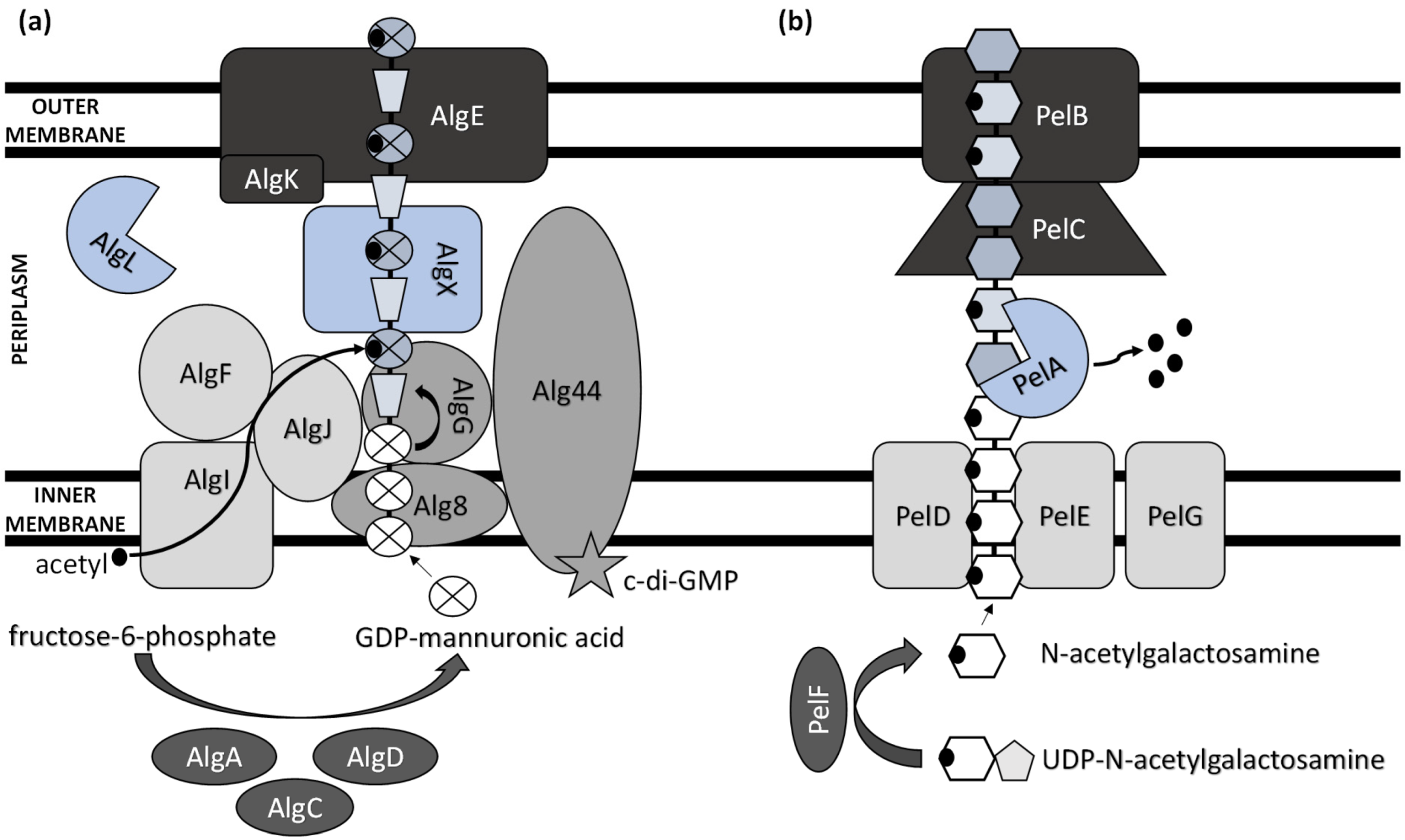
This entry is adapted from the peer-reviewed paper 10.3390/polym14204253
References
- Mohd Nadzir, M.; Nurhayati, R.W.; Idris, F.N. Biomedical applications of bacterial exopolysaccharides: A review. Polymers 2021, 13, 530.
- Nwodo, U.U.; Green, E.; Okoh, A.I. Bacterial exopolysaccharides: Functionality and prospects. Int. J. Mol. Sci. 2012, 13, 14002–14015.
- Kanmani, P.; Yuvapriya, S. Exopolysaccharide from Bacillus sp. YP03: Its properties and application as a flocculating agent in wastewater treatment. Int. J. Environ. Sci. Technol. 2018, 15, 2551–2560.
- Mulligan, C.N.; Yong, R.N.; Gibbs, B.F. Remediation technologies for metal-contaminated soils and groundwater: An evaluation. Eng. Geol. 2001, 60, 193–207.
- Hagarová, I.; Kudrík, I. Optimization of extraction procedure with nonionic surfactant for determination of trace lead in waters. Chem. Listy 2016, 110, 504–510.
- Hagarová, I. Utilization of Supramolecular Solvents in the Extraction of Metals. Chem. Listy 2014, 108, 949–955.
- Zhang, T.; Liu, J.-M.; Huang, X.-F.; Xia, B.; Su, C.-Y.; Luo, G.-F.; Xu, Y.-W.; Wu, Y.-X.; Mao, Z.-W.; Qiu, R.-L. Chelant extraction of heavy metals from contaminated soils using new selective EDTA derivatives. J. Hazard. Mater. 2013, 262, 464–471.
- Hiller, E.; Jurkovič, Ľ.; Faragó, T.; Vítková, M.; Tóth, R.; Komárek, M. Contaminated soils of different natural pH and industrial origin: The role of (nano) iron- and manganese-based amendments in As, Sb, Pb, and Zn leachability. Environ. Pollut. 2021, 285, 117268.
- Chmielewska, E.; Tylus, W.; Bujdoš, M. Study of Mono- and Bimetallic Fe and Mn Oxide-Supported Clinoptilolite for Improved Pb(II) Removal. Molecules 2021, 26, 4143.
- Dudová, J.; Bujdoš, M. Study of selenium sorption on iron oxide hydroxides. Chem. Listy 2015, 109, 770–774.
- Hagarová, I.; Nemček, L. Application of Metallic Nanoparticles and Their Hybrids as Innovative Sorbents for Separation and Pre-concentration of Trace Elements by Dispersive Micro-Solid Phase Extraction: A Minireview. Front. Chem. 2021, 9, 672755.
- Pamukcu, S. In Situ Soil and Sediment Remediation. In Handbook of Environmental Engineering; John Wiley & Sons: Hoboken, NJ, USA, 2018; pp. 209–248.
- Song, T.-S.; Zhang, J.; Hou, S.; Wang, H.; Zhang, D.; Li, S.; Xie, J. In situ electrokinetic remediation of toxic metal-contaminated soil driven by solid phase microbial fuel cells with a wheat straw addition. J. Appl. Chem. Biotechnol. 2018, 93, 2860–2867.
- You, Y.; Dou, J.; Xue, Y.; Jin, N.; Yang, K. Chelating Agents in Assisting Phytoremediation of Uranium-Contaminated Soils: A Review. Sustainability 2022, 14, 6379.
- Zhou, W.; Shen, B.; Meng, F.; Liu, S.; Zhang, Y. Coagulation enhancement of exopolysaccharide secreted by an Antarctic sea-ice bacterium on dye wastewater. Sep. Purif. Technol. 2010, 76, 215–221.
- Kumar, A.S.; Mody, K.; Jha, B. Bacterial exopolysaccharides—A perception. J. Basic Microbiol. 2007, 47, 103–117.
- Li, C.; Yu, Y.; Fang, A.; Feng, D.; Du, M.; Tang, A.; Chen, S.; Li, A. Insight into biosorption of heavy metals by extracellular polymer substances and the improvement of the efficacy: A review. Lett. Appl. Microbiol. 2021.
- Hagarová, I. Utilization of biosurfactants in remediation of environmental media contaminated with heavy metals. Chem. Listy 2015, 109, 431–436.
- Villela, H.D.M.; Peixoto, R.S.; Soriano, A.U.; Carmo, F.L. Microbial bioremediation of oil contaminated seawater: A survey of patent deposits and the characterization of the top genera applied. Sci. Total Environ. 2019, 666, 743–758.
- Al Disi, Z.; Al-Ghouti, M.A.; Zouari, N. Investigating the simultaneous removal of hydrocarbons and heavy metals by highly adapted Bacillus and Pseudomonas strains. Environ. Technol. Innov. 2022, 27, 102513.
- Braud, A.; Jézéquel, K.; Bazot, S.; Lebeau, T. Enhanced phytoextraction of an agricultural Cr- and Pb-contaminated soil by bioaugmentation with siderophore-producing bacteria. Chemosphere 2009, 74, 280–286.
- Chlebek, D.; Płociniczak, T.; Gobetti, S.; Kumor, A.; Hupert-Kocurek, K.; Pacwa-Płociniczak, M. Analysis of the genome of the heavy metal resistant and hydrocarbon-degrading rhizospheric Pseudomonas qingdaonensis zcr6 strain and assessment of its plant-growth-promoting traits. Int. J. Mol. Sci. 2022, 23, 214.
- Moore, E.R.B.; Tindall, B.J.; Martins Dos Santos, V.A.P.; Pieper, D.H.; Ramos, J.-L.; Palleroni, N.J. Nonmedical: Pseudomonas. In The Prokaryotes: A Handbook on the Biology of Bacteria Volume 6: Proteobacteria: Gamma Subclass; Dworkin, M., Falkow, S., Rosenberg, E., Schleifer, K.-H., Stackebrandt, E., Eds.; Springer: New York, NY, USA, 2006; pp. 646–703.
- Abdelbary, S.; Elgamal, M.S.; Farrag, A. Trends in Heavy Metals Tolerance and Uptake by Pseudomonas aeruginosa. In Pseudomonas aeruginosa—An Armory Within; Sriramulu, D., Ed.; IntechOpen: London, UK, 2018.
- Nouha, K.; Kumar, R.S.; Balasubramanian, S.; Tyagi, R.D. Critical review of EPS production, synthesis and composition for sludge flocculation. J. Environ. Sci. 2018, 66, 225–245.
- Hay, I.D.; Rehman, Z.U.; Moradali, M.F.; Wang, Y.; Rehm, B.H.A. Microbial alginate production, modification and its applications. Microb. Biotechnol. 2013, 6, 637–650.
- Wu, Q.; Tun, H.M.; Leung, F.C.-C.; Shah, N.P. Genomic insights into high exopolysaccharide-producing dairy starter bacterium Streptococcus thermophilus ASCC 1275. Sci. Rep. 2014, 4, 4974.
- Osemwegie, O.O.; Adetunji, C.O.; Ayeni, E.A.; Adejobi, O.I.; Arise, R.O.; Nwonuma, C.O.; Oghenekaro, A.O. Exopolysaccharides from bacteria and fungi: Current status and perspectives in Africa. Heliyon 2020, 6, e04205.
- Rehm, B.H.A. Bacterial polymers: Biosynthesis, modifications and applications. Nat. Rev. Microbiol. 2010, 8, 578–592.
- Ohman, D.E. Molecular genetics of exopolysaccharide production by mucoid Pseudomonas aeruginosa. Eur. J. Clin. Microbiol. 1986, 5, 6–10.
- Kasak, P.; Sasová, J.; Shoheeduzzaman, R.; Baig, M.T.; Alyafei, A.A.H.A.; Tkac, J. Influence of direct electric field on PMCG-alginate-based microcapsule. Emergent Mater. 2021, 4, 769–779.
- Kasak, P.; Danko, M.; Zavahir, S.; Mrlik, M.; Xiong, Y.; Yousaf, A.B.; Lai, W.-F.; Krupa, I.; Tkac, J.; Rogach, A.L. Identification of molecular fluorophore as a component of carbon dots able to induce gelation in a fluorescent multivalent-metal-ion-free alginate hydrogel. Sci. Rep. 2019, 9, 15080.
- Franklin, M.; Nivens, D.; Weadge, J.; Howell, P. Biosynthesis of the Pseudomonas aeruginosa extracellular polysaccharides, alginate, Pel, and Psl. Front. Microbiol. 2011, 2, 167.
- May, T.B.; Shinabarger, D.; Boyd, A.; Chakrabarty, A.M. Identification of amino acid residues involved in the activity of phosphomannose isomerase-guanosine 5′-diphospho-D-mannose pyrophosphorylase. A bifunctional enzyme in the alginate biosynthetic pathway of Pseudomonas aeruginosa. J. Biol. Chem. 1994, 269, 4872–4877.
- Zielinski, N.A.; Chakrabarty, A.M.; Berry, A. Characterization and regulation of the Pseudomonas aeruginosa algC gene encoding phosphomannomutase. J. Biol. Chem. 1991, 266, 9754–9763.
- Tavares, I.M.; Leitão, J.H.; Fialho, A.M.; Sá-Correia, I. Pattern of changes in the activity of enzymes of GDP-D-mannuronic acid synthesis and in the level of transcription of algA, algC and algD genes accompanying the loss and emergence of mucoidy in Pseudomonas aeruginosa. Res. Microbiol. 1999, 150, 105–116.
- Oglesby, L.L.; Jain, S.; Ohman, D.E. Membrane topology and roles of Pseudomonas aeruginosa Alg8 and Alg44 in alginate polymerization. Microbiology 2008, 154, 1605–1615.
- Remminghorst, U.; Rehm, B.H.A. Alg44, a unique protein required for alginate biosynthesis in Pseudomonas aeruginosa. FEBS Lett. 2006, 580, 3883–3888.
- Maharaj, R.; May, T.B.; Shang-Kwei, W.; Chakrabarty, A.M. Sequence of the alg8 and alg44 genes involved in the synthesis of alginate by Pseudomonas aeruginosa. Gene 1993, 136, 267–269.
- Hay, I.D.; Remminghorst, U.; Rehm, B.H.A. MucR, a novel membrane-associated regulator of alginate biosynthesis in Pseudomonas aeruginosa. Appl. Environ. Microbiol. 2009, 75, 1110–1120.
- Nivens, D.E.; Ohman, D.E.; Williams, J.; Franklin, M.J. Role of alginate and its O acetylation in formation of Pseudomonas aeruginosa microcolonies and biofilms. J. Bacteriol. 2001, 183, 1047–1057.
- Franklin, M.J.; Chitnis, C.E.; Gacesa, P.; Sonesson, A.; White, D.C.; Ohman, D.E. Pseudomonas aeruginosa AlgG is a polymer level alginate C5-mannuronan epimerase. J. Bacteriol. 1994, 176, 1821–1830.
- Robles-Price, A.; Wong Thiang, Y.; Sletta, H.; Valla, S.; Schiller Neal, L. AlgX is a periplasmic protein required for alginate biosynthesis in Pseudomonas aeruginosa. J. Bacteriol. 2004, 186, 7369–7377.
- Schiller, N.L.; Monday, S.R.; Boyd, C.M.; Keen, N.T.; Ohman, D.E. Characterization of the Pseudomonas aeruginosa alginate lyase gene (algL): Cloning, sequencing, and expression in Escherichia coli. J. Bacteriol. 1993, 175, 4780–4789.
- Whitney, J.C.; Hay, I.D.; Li, C.; Eckford, P.D.W.; Robinson, H.; Amaya, M.F.; Wood, L.F.; Ohman, D.E.; Bear, C.E.; Rehm, B.H.; et al. Structural basis for alginate secretion across the bacterial outer membrane. Proc. Natl. Acad. Sci. USA 2011, 108, 13083–13088.
- Keiski, C.-L.; Harwich, M.; Jain, S.; Neculai, A.M.; Yip, P.; Robinson, H.; Whitney, J.C.; Riley, L.; Burrows, L.L.; Ohman, D.E.; et al. AlgK is a TPR-containing protein and the periplasmic component of a novel exopolysaccharide secretin. Structure 2010, 18, 265–273.
- Myszka, K.; Czaczyk, K. Characterization of adhesive exopolysaccharide (EPS) produced by Pseudomonas aeruginosa under starvation conditions. Curr. Microbiol. 2009, 58, 541–546.
- Grobe, S.; Wingender, J.; Flemming, H.-C. Capability of mucoid Pseudomonas aeruginosa to survive in chlorinated water. Int. J. Hyg. Environ. Health 2001, 204, 139–142.
- Grobe, S.; Wingender, J.; Trüper, H.G. Characterization of mucoid Pseudomonas aeruginosa strains isolated from technical water systems. J. Appl. Bacteriol. 1995, 79, 94–102.
- Wozniak, D.J.; Wyckoff, T.J.O.; Starkey, M.; Keyser, R.; Azadi, P.; O’Toole, G.A.; Parsek, M.R. Alginate is not a significant component of the extracellular polysaccharide matrix of PA14 and PAO1 Pseudomonas aeruginosa biofilms. Proc. Natl. Acad. Sci. USA 2003, 100, 7907–7912.
- Colvin, K.M.; Irie, Y.; Tart, C.S.; Urbano, R.; Whitney, J.C.; Ryder, C.; Howell, P.L.; Wozniak, D.J.; Parsek, M.R. The Pel and Psl polysaccharides provide Pseudomonas aeruginosa structural redundancy within the biofilm matrix. Environ. Microbiol. 2012, 14, 1913–1928.
- Jennings, L.K.; Storek, K.M.; Ledvina, H.E.; Coulon, C.; Marmont, L.S.; Sadovskaya, I.; Secor, P.R.; Tseng, B.S.; Scian, M.; Filloux, A.; et al. Pel is a cationic exopolysaccharide that cross-links extracellular DNA in the Pseudomonas aeruginosa biofilm matrix. Proc. Natl. Acad. Sci. USA 2015, 112, 11353–11358.
- Le Mauff, F.; Razvi, E.; Reichhardt, C.; Sivarajah, P.; Parsek, M.R.; Howell, P.L.; Sheppard, D.C. The Pel polysaccharide is predominantly composed of a dimeric repeat of α-1,4 linked galactosamine and N-acetylgalactosamine. Commun. Biol. 2022, 5, 502.
- Friedman, L.; Kolter, R. Genes involved in matrix formation in Pseudomonas aeruginosa PA14 biofilms. Mol. Microbiol. 2004, 51, 675–690.
- Ghafoor, A.; Jordens, Z.; Rehma, H.A.B. Role of pelf in pel polysaccharide biosynthesis in Pseudomonas aeruginosa. Appl. Environ. Microbiol. 2013, 79, 2968–2978.
- Whitfield Gregory, B.; Marmont Lindsey, S.; Ostaszewski, A.; Rich Jacquelyn, D.; Whitney John, C.; Parsek Matthew, R.; Harrison Joe, J.; Howell, P.L. Pel Polysaccharide Biosynthesis Requires an Inner Membrane Complex Comprised of PelD, PelE, PelF, and PelG. J. Bacteriol. 2020, 202, e00684-19.
- Low, K.E.; Howell, P.L. Gram-negative synthase-dependent exopolysaccharide biosynthetic machines. Curr. Opin. Struct. Biol. 2018, 53, 32–44.
- Colvin Kelly, M.; Alnabelseya, N.; Baker, P.; Whitney John, C.; Howell, P.L.; Parsek Matthew, R. PelA deacetylase activity is required for Pel polysaccharide synthesis in Pseudomonas aeruginosa. J. Bacteriol. 2013, 195, 2329–2339.
- Marmont, L.S.; Rich, J.D.; Whitney, J.C.; Whitfield, G.B.; Almblad, H.; Robinson, H.; Parsek, M.R.; Harrison, J.J.; Howell, P.L. Oligomeric lipoprotein PelC guides Pel polysaccharide export across the outer membrane of Pseudomonas aeruginosa. Proc. Natl. Acad. Sci. USA 2017, 114, 2892–2897.
- Byrd, M.S.; Sadovskaya, I.; Vinogradov, E.; Lu, H.; Sprinkle, A.B.; Richardson, S.H.; Ma, L.; Ralston, B.; Parsek, M.R.; Anderson, E.M.; et al. Genetic and biochemical analyses of the Pseudomonas aeruginosa Psl exopolysaccharide reveal overlapping roles for polysaccharide synthesis enzymes in Psl and LPS production. Mol. Microbiol. 2009, 73, 622–638.
- Lee, H.-J.; Chang, H.-Y.; Venkatesan, N.; Peng, H.-L. Identification of amino acid residues important for the phosphomannose isomerase activity of PslB in Pseudomonas aeruginosa PAO1. FEBS Lett. 2008, 582, 3479–3483.
- Baker, P.; Whitfield, G.B.; Hill, P.J.; Little, D.J.; Pestrak, M.J.; Robinson, H.; Wozniak, D.J.; Howell, P.L. Characterization of the Pseudomonas aeruginosa glycoside hydrolase PslG reveals that Its levels are critical for Psl polysaccharide biosynthesis and biofilm formation. J. Biol. Chem. 2015, 290, 28374–28387.
- Wu, H.; Wang, D.; Tang, M.; Ma, L.Z. The advance of assembly of exopolysaccharide Psl biosynthesis machinery in Pseudomonas aeruginosa. MicrobiologyOpen 2019, 8, e857.
