The Golgi apparatus is an important organelle found in most eukaryotic cells. It plays a vital role in the processing and sorting of proteins, lipids and other cellular components for delivery to their appropriate destinations within the cell or for secretion outside of the cell. The Golgi complex also plays a role in the regulation of protein trafficking, secretion and post-translational modifications, which are significant in the development and progression of cancer. Abnormalities in this organelle have been observed in various types of cancer, although research into chemotherapies that target the Golgi apparatus is still in its early stages.
1. Golgi’s Major Roles
Camillo Golgi made an important observation while studying Purkinje cells in the cerebellum in 1898 using silver nitrate staining. He identified what would later become known as the Golgi apparatus using this technique and, shortly after, discovered its presence among neural cells of spinal ganglia. Soon thereafter, it was identified in other structures; initially known as an “internal reticular apparatus”, its function remained unclear at that time, and, for an extended period, it was thought to be just a staining artifact
[1].
The Golgi apparatus, also known as the Golgi complex or Golgi body, is a cellular organelle present in eukaryotic cells where it fulfills its function in the modification and transportation of proteins and lipids. It is made up of a series of flattened, membrane-bound sacs called cisternae, which are arranged in stacks called Golgi stacks. There are typically between three and eight Golgi stacks in a cell, and each stack can contain five to seven cisternae
[2].
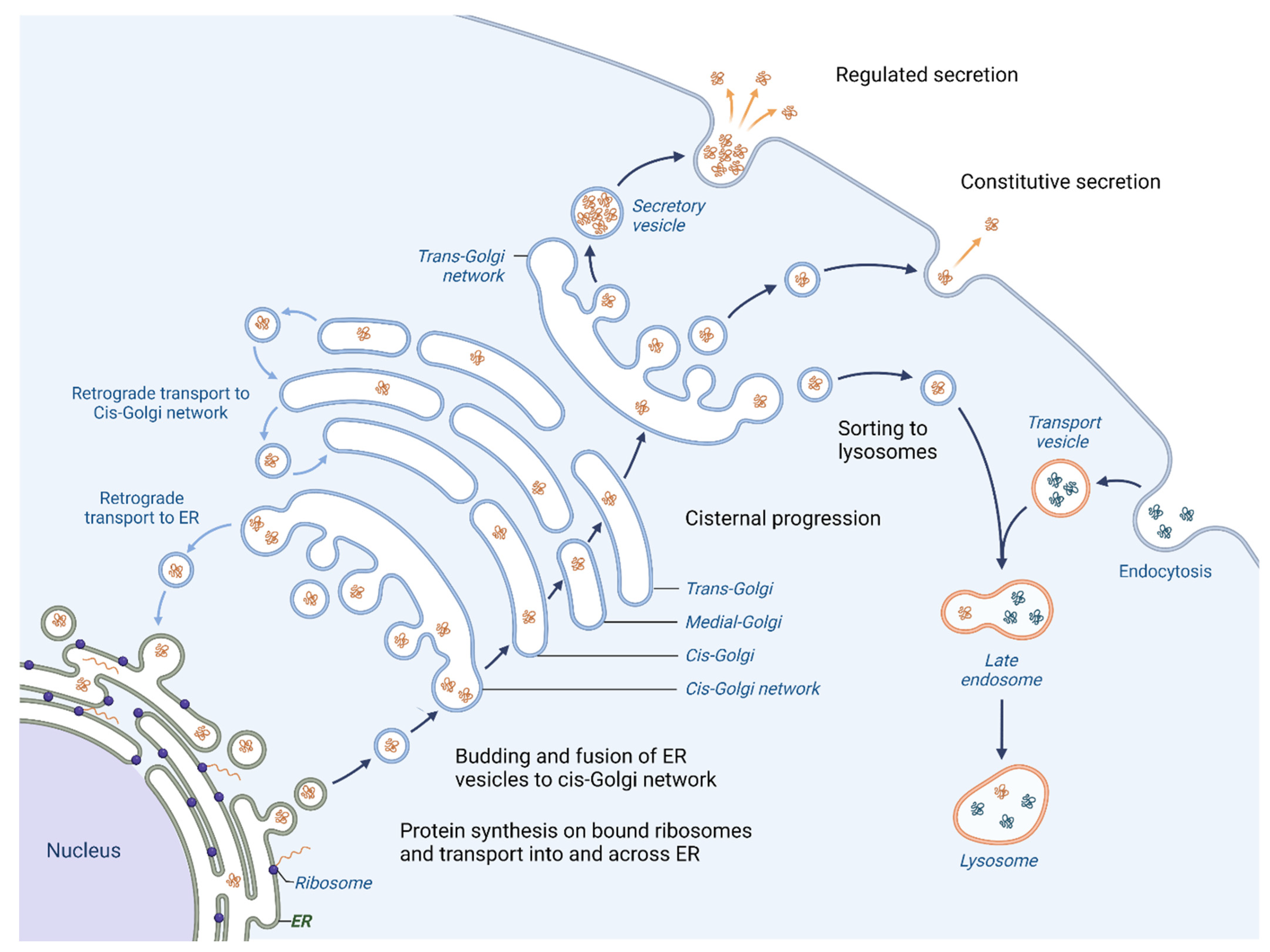
One of the primary roles of the Golgi apparatus is to sort and pack proteins for transport to their final destination within the cell or for secretion outside the cell. Proteins that are created in the Endoplasmic reticulum (ER) follow an anterograde pathway through the Golgi apparatus, passing through the cis-, medial- and trans-Golgi compartments (Figure 1).
Figure 1. Vesicular trafficking in cell. Initially, the proteins are packaged in vesicles within the ER. A part of them joins together to form the cis-cisterna of the Golgi apparatus. Enzymes within the organelle modify the proteins as they progress through the Golgi, with the specific modifications acting as a signal to determine the final destination of the proteins. The proteins are then sorted within the trans-Golgi network and packaged into vesicles that are transported to specific locations. Certain proteins are directed towards lysosomes for breakdown (proteases, lipases and glycosidases), whereas others are released outside of the cell where they aid in creating the extracellular matrix (ECM), engage with or modify the ECM or facilitate communication with other cells. Moreover, the Golgi apparatus is responsible for retrograde transport of proteins; hence the recycling of proteins back to their origins is possible.
The Golgi apparatus is also responsible for post-translational modifications (PTMs) of proteins. PTMs are chemical modifications that occur after a protein has been synthesized by the ribosome and are essential for proper protein function, one of the most significant being the glycosylation of proteins. This process involves the addition of oligosaccharides to particular proteins or lipids, resulting in the formation of glycoconjugates (glycoproteins, proteoglycans, glycosphingolipids, etc.). It occurs within the ER and Golgi apparatus, and its primary mechanisms are N-glycosylation and O-glycosylation. N-glycosylation is a process where a carbohydrate chain is covalently attached to the amide group of an asparagine residue (N) in the protein. The initial stages of N-glycosylation commence in the ER, where a pre-formed oligosaccharide is moved from a dolichol lipid carrier to an Asn residue located within the N-X-S/T consensus sequence (where X is any amino acid except proline and S/T is serine or threonine). The oligosaccharide is then modified and further processed as the protein moves through the Golgi apparatus
[3][4]. O-glycosylation, on the other hand, is a process in which a carbohydrate chain is attached to the hydroxyl group of a serine or threonine residue (S/T) in the protein. There are various types, including mucin-type O-glycosylation, O-GlcNAcylation and O-fucosylation, that occur in the Golgi apparatus and/or other compartments within the cell
[3][5]. Furthermore, there are other types of pathways that belong to the process of glycosylation. Sialylation represents the terminal addition of sialic acid, a negatively charged monosaccharide, to a glycan chain on a protein or lipid. It plays a crucial role in numerous biological processes such as inflammation, immune response and cell–cell recognition. Sialylation mainly occurs in the Golgi apparatus and is facilitated by a group of enzymes known as sialyltransferases
[6][7]. In addition, fucosylation is also a biochemical process in which fucosyltransferases (FUT) enzymatically transfer a fucose molecule from GDP-fucose to glycan structures
[8]. So, the Golgi apparatus is responsible for the final processing and sorting of glycoproteins, which involves the addition, removal and modification of carbohydrate groups.
Researchers need to fathom how this organelle interferes when the tumoral cell’s request for proteins increases and overwhelms the capacity of the Golgi in cancer, leading this way to the insufficiency (or stress) of the function
[9]. When the organelle is exposed to stress, it triggers a cascade of molecular events that are collectively referred to as the Golgi stress response, a cellular mechanism that is activated when the functionality of the organelle is impaired, aiding in the restoration of its normal function. The pathways of the Golgi stress response, as they are for ER, do not seem very tangled. There is a group of sensor molecules that identify when the Golgi apparatus is not performing its tasks adequately. This, in turn, activates a transcription factor that encourages the transcription of genes related to the Golgi apparatus. The transcriptional enhancer element is where the aforementioned factor binds, and it targets genes that encode Golgi-related proteins, with a noteworthy mention of the glycosylation enzymes
[10]. Only five stress response pathways have been described—transcription factor E3 (TFE3), cyclic AMP-responsive element-binding protein 3 (CREB3), proteoglycan (PG), mitogen-activated protein kinase MAPK-ETS and heat shock protein 4 (HSP47)—although many of the other elements remain to be clarified
[11]. So, the Golgi stress response is an important cellular mechanism that helps to maintain the function and integrity of the Golgi apparatus in tumoral cells, making it worthy of further studies leading probably to therapies that counter the pathological effects of them.
2. Golgi Effects in a Cancer Cell Invasion
Firstly, the Golgi apparatus is involved in the secretion of proteins and lipids from the cell; hence, various alterations can affect the composition of the secretome—the collection of all the proteins and other molecules secreted by a cell—and the extracellular matrix (ECM), leading to numerous diseases, including obesity, diabetes and cancer
[12]. Several gene-encoding proteins involved in Golgi-associated processes have been found to be frequently mutated in cancer. In addition, mutations in these genes are associated with enhanced metastasis and lower patient survival in several types of cancer, including breast and lung
[13].
Increased secretion and deposition of ECM components can contribute to cancer development and progression. Matrix metalloproteinases (MMPs) can degrade components of the ECM, allowing cancer cells to invade and migrate through tissues, and can also promote angiogenesis by releasing VEGF and other signaling molecules from the ECM. Furthermore, some MMPs are important for the degradation of the basement membrane, this process being necessary during the metastatic cascade
[8][12].
The Golgi apparatus performs a vital function in releasing immune components such as cytokines and chemokines, which take part in the formation of the tumor micro-environment (TME). In many types of cancer, alterations in the Golgi apparatus can lead to an increased secretion of immune factors that promote an immunosuppressive TME, which can contribute to cancer development and progression. For example, Phosphatidylinositol 4-kinase III beta (PI4KIIIβ) is a lipid kinase primarily localized to the Golgi apparatus, where it has been shown to regulate the activity of several signaling pathways that are involved in cancer development and progression
[13]. In addition, GOLPH3 (Golgi phosphoprotein 3) and CKAP4 (cytoskeleton-associated protein 4) are two proteins localized in the Golgi apparatus and upregulated in several types of cancer by enhancing the secretion of exosomal WNT3A
[14]. Furthermore, inflammatory cells secrete cytokines (TNF-α, IL-6, etc.) which can promote chronic inflammation within the TME, leading to cancer cell proliferation, survival and migration
[15].
As mentioned before, the Golgi complex is also responsible for the well-mediated trafficking of the secreted proteins within or outside the cell. Dysregulation of this process in cancer can result in altered protein localization, increased protein secretion and the activation of oncogenic signaling pathways
[12][16]. In particular, RAB and ARF proteins (ADP-ribosylation factor) are both involved in the regulation of Golgi-mediated trafficking, but they play distinct roles in this process: RAB proteins are involved in transport from the ER to the Golgi, intra-Golgi and from the Golgi to the other organelles or the plasma membrane, while ARF proteins regulate vesicle formation and budding from membranes. A dysregulation of both GTPases leads to disrupted protein trafficking and contributes to the development and progression of cancer
[17][18]. Moreover, SNARE and SNAP proteins, which are critical for the regulation of membrane fusion events in cells, are adjusted in cancer cells to promote invasion
[19]. Golgi-specific proteins also can act as oncoproteins when their expression is upregulated or their function is altered in cancer cells—GOLPH3’s increased secretion causes exocytosis of pro-metastatic factors
[5][14]. Hence, Golgi’s role in protein trafficking is significant in tumoral cells because of the great number of molecules used for survival of cancer.
Secondly, the orientation of the Golgi can alter its function and contribute to cell polarity and migration in the front–rear polarity model
[20]. The Golgi can dynamically reorient itself in response to migratory cues, such as chemokine gradients or extracellular matrix (ECM) stiffness, to guide vesicular flow toward the leading edge of the cell. Out of consideration for this, in migratory cells, this directional flow can facilitate the localized secretion of pro-migratory factors, such as MMPs, growth factors and cytokines, which can promote cell migration and invasion
[21]. So, the orientation of the Golgi apparatus regulates exocytosis and directs the movement of vesicles toward the front edge of the cell, promoting directional migration that can facilitate cancer metastasis.
On the other hand, the morphology and dynamics of the Golgi can be altered in cancer, and these changes can contribute to tumor progression and metastasis. In normal cells, the Golgi is typically organized as a compact perinuclear structure with distinct cis, medial and trans compartments. However, in cancer cells, the Golgi can exhibit a range of morphological changes, including fragmentation, expansion and redistribution throughout the cell
[8]. It is generally agreed that a well-organized Golgi structure is essential for precise post-translational modification and sorting of proteins. Thus, Golgi fragmentation, usually triggered by cellular stresses, growth factors and mitotic phosphorylation of Golgi structural proteins, accelerates protein trafficking and cell proliferation
[22].
Aside from the Golgi’s altered morphology and orientation and its influence on the secretome, there is another adaptation that this organelle adopts when it comes to tumoral growths and metastasis. The accumulation of evidence indicates that changes in the glycosylation patterns of carrier proteins—including partial formation of glycan structures, increased expression of complex branched N-glycans and truncated O-glycans (Tn and Sialyl Tn antigen), as well as changes in the expression of sialylated glycans and an increase in the expression of ‘core’ fucosylation—promote the procurement of cellular traits that are essential for the malignant transformation of cells
[23]. Changes in glycosylation appear to have a direct impact on cell growth and survival, as well as promoting tumor-associated immune responses and metastasis.
N-glycosylation is a highly conserved process that occurs via the catalysis of enzyme complexes called Oligosaccharyltransferases (OST), located in the ER membrane. The OST complex operates by binding glycans to specific asparagine sites on newly forming polypeptides within the ER lumen. The mammalian OST complex is composed of various subunits, among which STT3 (STT3A and STT3B) has a crucial function in N-glycosylation processes in mammals. Blocking the STT3A subunit amplifies the unfolded protein response in mammalian cells, leading to detrimental effects such as cancer. Inhibition of OST enzyme activity results in the disruption of proper protein folding in cells, which is a consequence of its high substrate specificity towards glycosylation. This perturbation in the process can lead to various pathological conditions
[24].
However, the abnormal expression of N-acetylgalactosaminyltransferase 3 (GALNT3), a gene that encodes UDP-GalNAc transferase 3, enhances the O-glycosylation of MUC1. This process leads to increased stabilization of the E-cadherin and β-catenin complex, ultimately stimulating cell migration and proliferation in ovarian cancer cells
[25]. Therefore, suppression of cell proliferation and invasion is observed when MUC1 destabilization occurs via GALNT3 inhibition
[26]. Moreover, it was shown that the interaction between MUC1 and β-catenin leads to the destabilization of adherent junctions, the disturbance of cytoskeletal architecture and an increased cell invasion in breast cancer cells
[5]. MUC16, a transmembrane protein, is involved in promoting cell adhesion contact among epithelial cells via its interaction with the actin cytoskeleton within the extracellular matrix (ECM). Disruption of MUC16 genetic expression interferes with the binding between the actin cytoskeleton and the cytosolic domain of MUC16, thereby leading to increased migration and invasion of epithelial cells
[27].
Numerous transformed tumor cells exhibit distinctive alterations in their sialylation patterns. It has been reported that pancreatic cancer cells with an increased malignant potential are correlated with the overexpression of sialyl Tn (STn) antigen
[28][29][30].
Along with this, abnormal fucosylation is associated with multiple cancers. Physiologically, this process is catalyzed by thirteen types of fucosyltransferases (FUTs) of which FUT1–11 are known to catalyze N-linked fucosylation and be located in the Golgi apparatus, while the other two types exist in the ER and catalyze O-linked fucosylation. Among the fucosylated glycans, the most well-known are ABO blood group antigens
[31]. There were several reports about pathological manifestations that occurred due to dysfunctions of fucosylation: FUT8 was identified as a prognostic marker in patients with colorectal cancer (CRC); changes in the p53 gene alter the predictive relationships between the expression of FUT8 and the prognosis of individuals with CRC.
[32]. In addition, a biomarker for detecting colorectal carcinogenesis was found to be an increase in fucosylation of N-glycans
[33]. Furthermore, the association between high levels of FUT6 and colorectal cancer progression has been confirmed
[34].
This entry is adapted from the peer-reviewed paper 10.3390/cells12111499