Chitin, a major structural component of the fungal cell wall, synthesized via the activityof the enzyme chitin synthase (chs), has become a high-profifile target for investigating the effect on morphology, development and pathogenicity of filamentous fungi. Besides, disruption of chitin biosynthesis can modify the mycelial morphology of filamentous fungi and regulate the biosynthesis of the target metabolites during submerged fermentation. Thus, we summarize the classifification, structure and function of chs enzymes, the biosynthetic pathway of chitin of filamentous fungi.
- chitin
- mycelia morphology
- filamentous fungi
- development
1. Structure and Function of Chitin
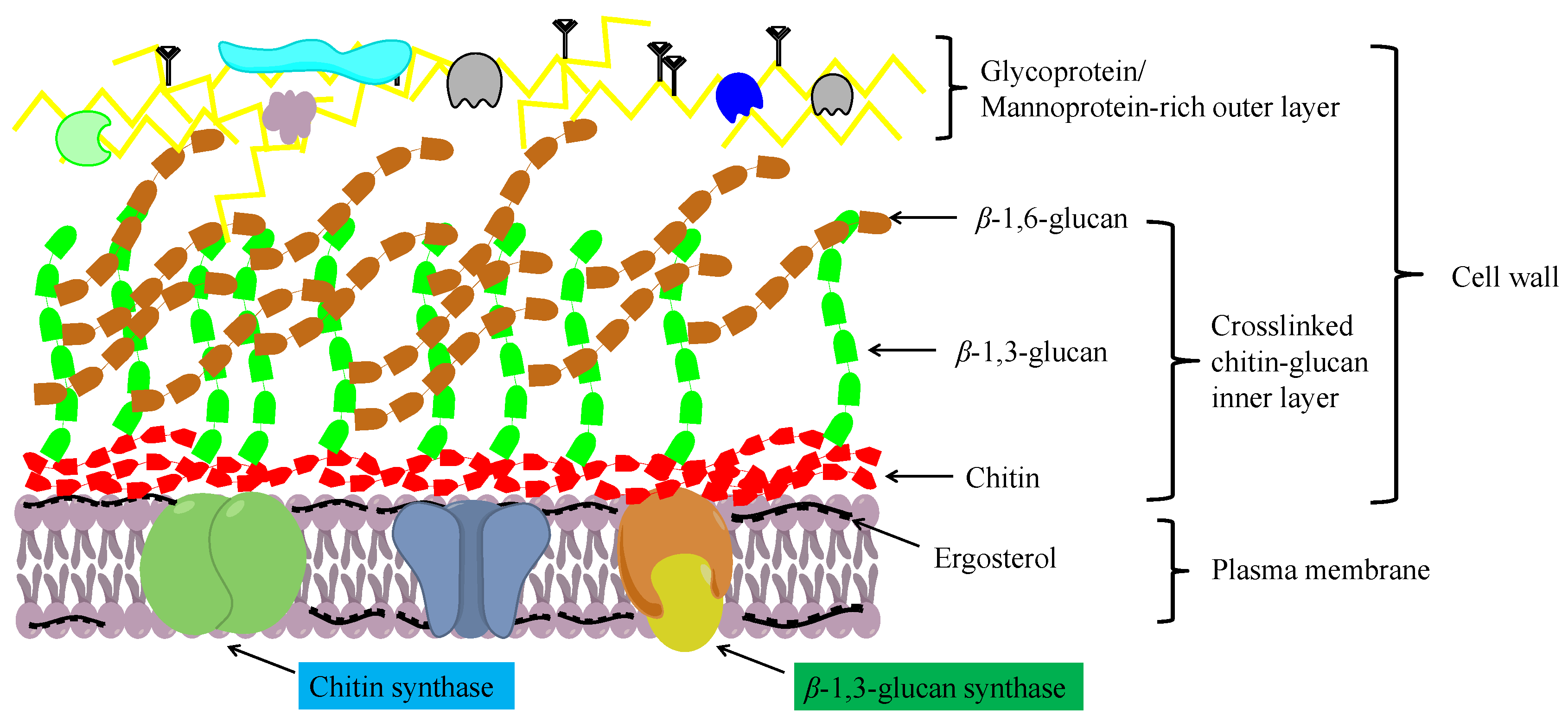
2. The Chitin Biosynthetic Pathway
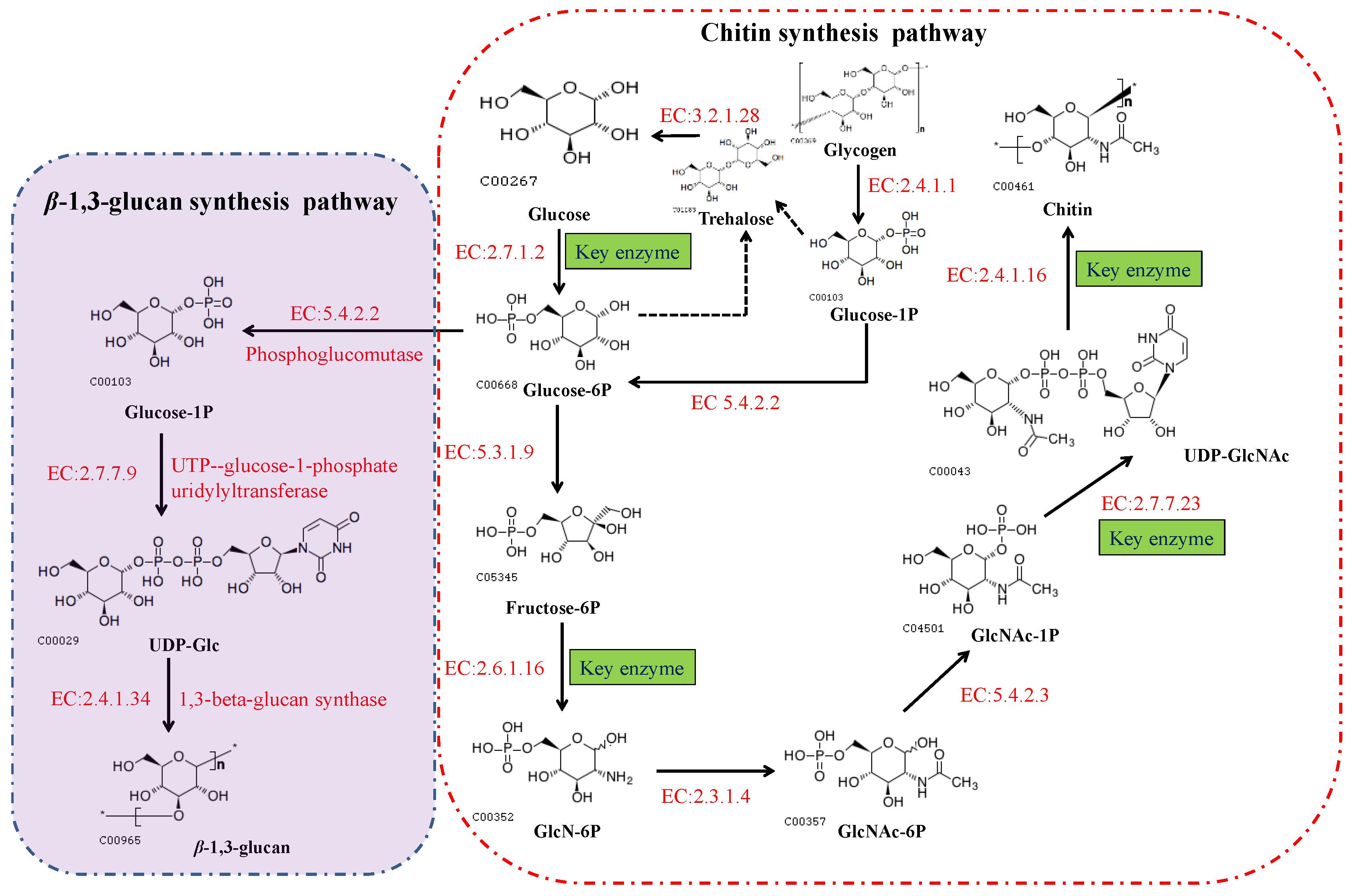
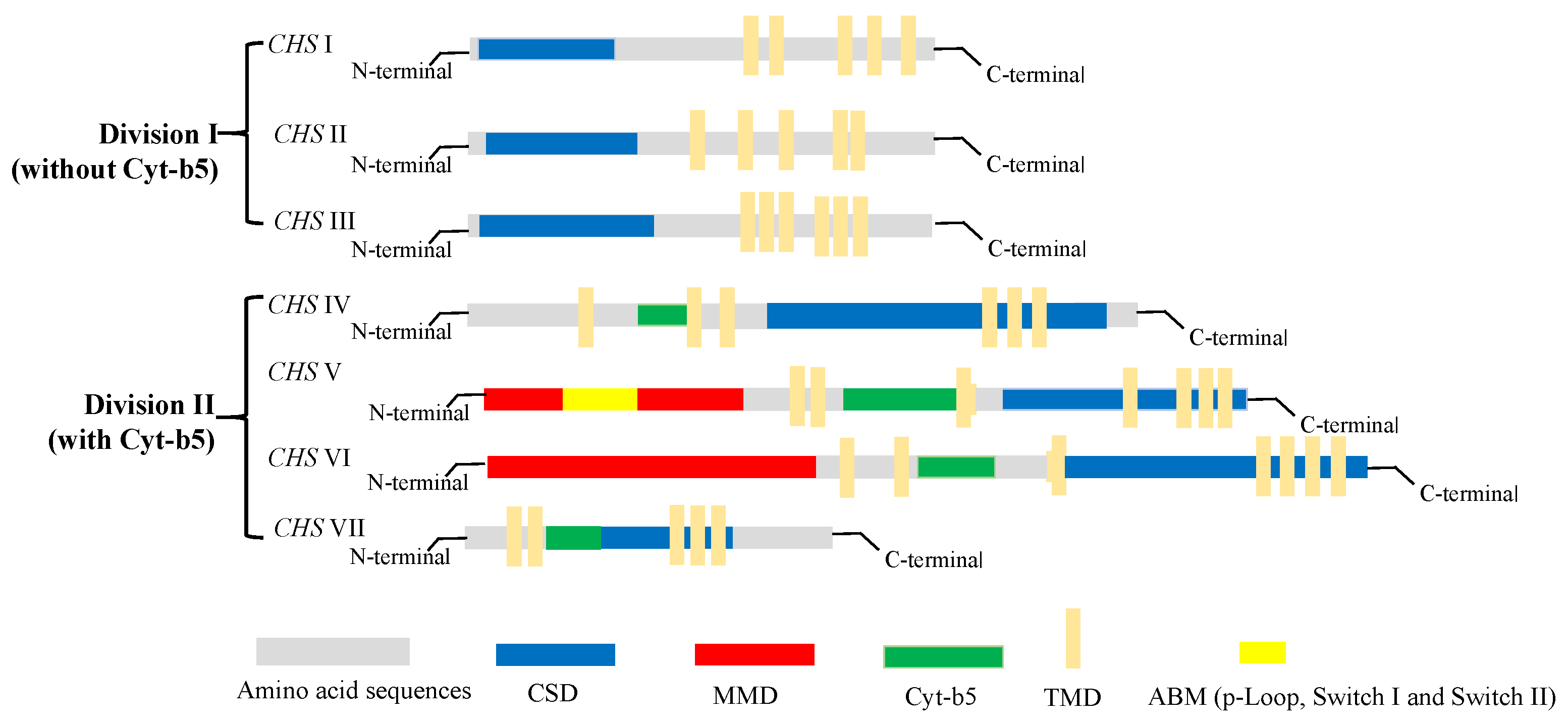
3. Classification of Chitin Synthase
| Organism | T-Number | The Members of Chitin Synthase | Number of Genes |
|||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Saccharomyces cerevisiae S288c | T00005 | YBR023C, chs 3 | YBR038W, chs 2 | YNL192W, chs 1 | YLR330W, chs 5 | YJL099W, chs 6 |
YHR142W, no KO assigned | (RefSeq) chs7; chs 7p |
6 | ||||
| Lodderomyces elongisporus NRRLYB-4239 | T01116 | LELG_05384, chs 2 | LELG_05013, chs 1 | LELG_02210, chs 2 | LELG_00298, chs 3 | LELG_00300, chs 3 | 5 | |||||
| Candida tropicalis MYA-3404 | T01115 | CAALFM_ C113110CA, chs 3 |
CAALFM_ C300710WA, chs 8 |
CAALFM_ C702770WA, chs 1 |
CAALFM_ CR09020CA, chs 2 |
4 | ||||||
| Candida orthopsilosis Co 90-125 |
T02488 | CORT_0A01870, chs 3 | CORT_0D06430, chs 8 | CORT_0G01660, chs 2 | CORT_0H01960, chs 1 | CORT_0H01970, chs 1 | 5 | |||||
| Sugiyamaella lignohabitans CBS 10342 | T05270 | AWJ20_11, chs 6 | AWJ20_12, chs 3 | AWJ20_13, chs 3 | AWJ20_1163, chs 2 | AWJ20_1500, chs 2 | AWJ20_3769, chs 1 | AWJ20_4861, chs 3 | AWJ20_4948, chs 3 | 8 | ||
| Xenopus laevis (African clawed frog) | T01010 | 108717413, chs 2 | 108716131, chs 2 |
2 | ||||||||
| Xenopus tropicalis (tropical clawed frog) | T01011 | 105947355, chs 2-like isoform X1 | 1 | |||||||||
| Carassius auratus (goldfish) | T07313 | 113057339 CHS 2-like | 113061218 CHS 1-like | 113061224 CHS 1-like | 113061225 CHS 1-like | 113061526 CHS 1 | 113061527 CHS 1-like | 113113123 CHS 2-like |
7 | |||
| Pyricularia oryzae 70-15 | T01027 | MGG_09962, chs 4 |
MGG_06064, chs D |
MGG_09551, chs 3 |
MGG_13013, chs 8 |
MGG_13014, CHS V |
MGG_01802, chs1 |
MGG_04145, chs 2 |
7 | |||
| Fusarium graminearum | T01038 | FGSG_01272, chs 4 |
FGSG_01949, chs D |
fgr:FGSG_12039, chs 6 |
fgr:FGSG_01964, hypothetical protein |
fgr:FGSG_02483, chs 2 |
fgr:FGSG_10116, chs 1 |
fgr:FGSG_10327, chs 3 |
fgr:FGSG_10619, hypothetical protein |
fgr:FGSG_03418, chs 1 |
fgr:FGSG_06550, hypothetical protein |
10 |
| Purpureocillium lilacinum | T05029 | VFPFJ_00650, chs D |
VFPFJ_00666, chs 6 |
VFPFJ_00667, chs 6 |
VFPFJ_03324, chs D |
VFPFJ_04443, chs A |
VFPFJ_08553, chs G |
VFPFJ_08866, chs A |
VFPFJ_11040, chs |
8 | ||
| Pestalotiopsis fici W106-1 | T04924 | PFICI_01118, chs 1 | PFICI_01446, chs 4 | PFICI_04362, hypothetical protein | PFICI_04363, hypothetical protein | PFICI_05017, chs D | PFICI_05238, chs 2 | PFICI_06085, chs 3 | PFICI_07201, chs 1 | PFICI_12982, hypothetical protein | PFICI_13513, chs 1 | 10 |
| Botrytis cinerea B05.10 | T01072 | BCIN_01g02520, CHS IIIb | BCIN_01g03790, CHS IV | BCIN_04g03120, CHS IIIa | BCIN_07g01300, CHS VII | BCIN_09g01210, CHS I | BCIN_12g01380, CHS II | BCIN_12g05360, CHS VI | BCIN_12g05370, CHS V | 8 | ||
| Aspergillus fumigatus Af293 | T01017 | AFUA_4G04180, chs B | AFUA_8G05630, chs F | AFUA_5G00760, chs C | AFUA_2G01870, chs A | AFUA_1G12600, chs D | AFUA_3G14420, chs G | AFUA_2G13430, chs | AFUA_2G13440, chs E | 8 | ||
| Aspergillus niger CBS 513.88 | T01030 | ANI_1_316024, chs | ANI_1_2332024, chs | ANI_1_1542034, chs C | ANI_1_684064, chs C | ANI_1_1986074, chs D | ANI_1_252084, chs D | ANI_1_498084, chs B | ANI_1_1214104, chs C | ANI_1_120124, chs A | 9 | |
| Aspergillus nidulans FGSC A4 | T01016 | AN1555.2, CHS V (chs D) | AN2523.2, chs B | AN4367.2, hypothetical protein | AN4566.2, hypothetical protein | AN6317.2, hypothetical protein | AN6318.2, hypothetical protein | AN7032.2, hypothetical protein | 7 | |||
| Neurospora crassa | T01034 | NCU09324, chs 4 | NCU04352, chs 5 | NCU04350, chs 6 | NCU05268, chs 6; | NCU05239, chs A | NCU03611, chs 1 | NCU04251, chs 3 | 7 | |||
| Penicillium digitatum Pd1 | T04849 | PDIP_79230, chs E | PDIP_62350, hypothetical protein | PDIP_46630, chs G | PDIP_26990, chs D | PDIP_24450, chs G | PDIP_15450, chs B | PDIP_07640, chs A | PDIP_03360, chs F | 9 | ||
| Coccidioides immitis RS | T01114 | CIMG_05021, CHS V | CIMG_05598, chs C | CIMG_05647, chs G | CIMG_05022, chs 5 | CIMG_08766, chs 4 | CIMG_08655, chs 2 | CIMG_06862, CHS VI | 8 | |||
| Monascus purpureus HQ1 | TQB77221.1, CHS V | TQB75461.1, CHS III | TQB73913.1, CHS I | TQB72986.1, CHS VII | TQB70564.1, CHS II | TQB69157.1, CHS II | TQB73548.1, hypothetical protein | TQB73973.1, hypothetical protein | TQB73547.1, hypothetical protein | 9 | ||
| Monascus purpureus LQ-6 | monascus_02563, chs2 | monascus_02508, chs3 | monascus_05,161 chs 4 | monascus_05162, chs 6 | monascus_02870, chs activator | monascus_02765, chs 5 | monascus_02400, chs G | monascus_04382, chs A | 8 | |||
| Monascus purpureus M183 | g872, chs 2 | g920, chs F | g3077, chsE | g3078, chs | g2747, chs 3 | g5275, chs 3 | g4739, chs B | g5640, chs A | 8 | |||
1. Al Shaqsi, N.H.K.; Al Hoqani, H.A.S.; Hossain, M.A.; Al Sibani, M.A. Optimization of the demineralization process for the extraction of chitin from Omani Portunidae segnis. Biochem. Biophys. Rep. 2020, 23, 100779.
2. Poerio, A.; Petit, C.; Jehl, J.P.; Arab-Tehrany, E.; Mano, J.F.; Cleymand, F. Extraction and Physicochemical Characterization of Chitin from Cicada orni Sloughs of the South-Eastern French Mediterranean Basin. Molecules 2020, 25, 2543.
3. Rinaudo, M. Chitin and chitosan: Properties and applications. Prog. Polym. Sci. 2006, 31, 603–632.
4. Klis, F.M.; Mol, P.; Hellingwerf, K.; Brul, S. Dynamics of cell wall structure in Saccharomyces cerevisiae. FEMS Microbiol. Rev. 2002,26, 239–256.
5. Lesage, G.; Bussey, H. Cell wall assembly in Saccharomyces cerevisiae. Microbiol. Mol. Biol. Rev. 2006, 70, 317–343.
6. Garcia-Rubio, R.; de Oliveira, H.C.; Rivera, J.; Trevijano-Contador, N. The Fungal Cell Wall: Candida, Cryptococcus, and Aspergillus Species. Front. Microbiol. 2019, 10, 2993.
7. Chattaway, F.W.; Holmes, M.R.; Barlow, A.J. Cell wall composition of the mycelial and blastospore forms of Candida albicans. J.Gen. Microbiol. 1968, 51, 367–376.
8. Kanetsuna, F.; Carbonell, L.M.; Moreno, R.E.; Rodriguez, J. Cell wall composition of the yeast and mycelial forms of Paracoccidioides brasiliensis. J. Bacteriol. 1969, 97, 1036–1041.
9. Kang, M.S.; Elango, N.; Mattia, E.; Au-Young, J.; Robbins, P.W.; Cabib, E. Isolation of chitin synthetase from Saccharomyces cerevisiae. Purifification of an enzyme by entrapment in the reaction product. J. Biol. Chem. 1984, 259, 14966–14972.
10. El Knidri, H.; Belaabed, R.; Addaou, A.; Laajeb, A.; Lahsini, A. Extraction, chemical modifification and characterization of chitin and chitosan. Int. J. Biol. Macromol. 2018, 120, 1181–1189.
11. Moussian, B. Chitin: Structure, Chemistry and Biology. Adv. Exp. Med. Biol. 2019, 1142, 5–18.
12. Hassainia, A.; Satha, H.; Boufifi, S. Chitin from Agaricus bisporus: Extraction and characterization. Int. J. Biol. Macromol. 2018, 117,1334–1342.
13. Jang, M.K.; Kong, B.G.; Jeong, Y.I.; Lee, C.H.; Nah, J.W. Physicochemical characterization of α-chitin,β-chitin, and γ-chitin separated from natural resources. J. Polym. Sci. Part A-Polym. Chem. 2004, 42, 3423–3432.
14. Islam, S.; Bhuiyan, M.A.R.; Islam, M.N. Chitin and Chitosan: Structure, Properties and Applications in Biomedical Engineering. Polym. Environ. 2017, 3, 854–866.
15. Benchamas, G.; Huang, G.; Huang, S.; Huang, H. Preparation and biological activities of chitosan oligosaccharides. Trends Food Sci. Technol. 2021,107, 38–44.
16. No, H.K.; Meyers, S.P. Application of chitosan for treatment of wastewaters. Rev. Environ. Contam. Toxicol. 2000, 163, 1–27.
17. Gow, N.A.R.; Latge, J.P.; Munro, C.A.; Heitman, J. The Fungal Cell Wall: Structure, Biosynthesis, and Function. Microbiol. Spectr. 2017, 5, 5.
18. Yang, J.; Zhang, K.Q. Chitin Synthesis and Degradation in Fungi: Biology and Enzymes. Adv. Exp. Med. Biol. 2019, 1142, 153–167.
19. Goldman, D.L.; Vicencio, A.G. The chitin connection. mBio 2012, 3, e00056-12.
20. Merzendorfer, H. The cellular basis of chitin synthesis in fungi and insects: Common principles and differences. Eur. J. Cell Biol. 2011, 90, 759–769.
21. Watanabe, H.; Azuma, M.; Igarashi, K.; Ooshima, H. Analysis of Chitin at the Hyphal Tip of Candida albicans Using Calcoflfluor White. Biosci. Biotechnol. Biochem. 2005, 69, 1798–1801.
22. Roncero, C. The genetic complexity of chitin synthesis in fungi. Curr. Genet. 2002, 41, 367–378.
23. Zhang, J.; Jiang, H.; Du, Y.; Keyhani, N.O.; Xia, Y.; Jin, K. Members of chitin synthase family in Metarhizium acridum differentially affect fungal growth, stress tolerances, cell wall integrity and virulence. PLoS Pathog. 2019, 15, e1007964.
24. Munro, C.A.; Gow, N.A.R. Chitin synthesis in human pathogenic fungi. Med. Mycol. 2001, 39, 41–53.
25. Larson, T.M.; Kendra, D.F.; Busman, M.; Brown, D.W. Fusarium verticillioides chitin synthases CHS5 and CHS7 are required for normal growth and pathogenicity. Curr. Genet. 2011, 57, 177–189.
26. Cui, Z.; Wang, Y.; Lei, N.; Wang, K.; Zhu, T. Botrytis cinerea chitin synthase BcChsVI is required for normal growth and pathogenicity. Curr. Genet. 2013, 59, 119–128.
27. Takeshita, N.; Yamashita, S.; Ohta, A.; Horiuchi, H. Aspergillus nidulans class V and VI chitin synthases CsmA and CsmB, each with a myosin motor-like domain, perform compensatory functions that are essential for hyphal tip growth. Mol. Microbiol. 2006,59, 1380–1394.
28. Shu, M.; Lu, P.; Liu, S.; Zhang, S.; Gong, Z.; Cai, X.; Zhou, B.; Lin, Q.; Liu, J. Disruption of the Chitin Biosynthetic Pathway Results in Signifificant Changes in the Cell Growth Phenotypes and Biosynthesis of Secondary Metabolites of Monascus purpureus. J. Fungi 2022, 8, 910.
References
- Klis, F.M.; Mol, P.; Hellingwerf, K.; Brul, S. Dynamics of cell wall structure in Saccharomyces cerevisiae. FEMS Microbiol. Rev. 2002, 26, 239–256.
- Kang, M.S.; Elango, N.; Mattia, E.; Au-Young, J.; Robbins, P.W.; Cabib, E. Isolation of chitin synthetase from Saccharomyces cerevisiae. Purification of an enzyme by entrapment in the reaction product. J. Biol. Chem. 1984, 259, 14966–14972.
- El Knidri, H.; Belaabed, R.; Addaou, A.; Laajeb, A.; Lahsini, A. Extraction, chemical modification and characterization of chitin and chitosan. Int. J. Biol. Macromol. 2018, 120, 1181–1189.
- Moussian, B. Chitin: Structure, Chemistry and Biology. Adv. Exp. Med. Biol. 2019, 1142, 5–18.
- Hassainia, A.; Satha, H.; Boufi, S. Chitin from Agaricus bisporus: Extraction and characterization. Int. J. Biol. Macromol. 2018, 117, 1334–1342.
- Jang, M.K.; Kong, B.G.; Jeong, Y.I.; Lee, C.H.; Nah, J.W. Physicochemical characterization of α-chitin,β-chitin, and γ-chitin separated from natural resources. J. Polym. Sci. Part A-Polym. Chem. 2004, 42, 3423–3432.
- Rinaudo, M. Chitin and chitosan: Properties and applications. Prog. Polym. Sci. 2006, 31, 603–632.
- Islam, S.; Bhuiyan, M.A.R.; Islam, M.N. Chitin and Chitosan: Structure, Properties and Applications in Biomedical Engineering. J. Polym. Environ. 2017, 3, 854–866.
- Benchamas, G.; Huang, G.; Huang, S.; Huang, H. Preparation and biological activities of chitosan oligosaccharides. Trends Food Sci. Technol. 2021, 107, 38–44.
- No, H.K.; Meyers, S.P. Application of chitosan for treatment of wastewaters. Rev. Environ. Contam. Toxicol. 2000, 163, 1–27.
- Gow, N.A.R.; Latge, J.P.; Munro, C.A.; Heitman, J. The Fungal Cell Wall: Structure, Biosynthesis, and Function. Microbiol. Spectr. 2017, 5, 5.
- Yang, J.; Zhang, K.Q. Chitin Synthesis and Degradation in Fungi: Biology and Enzymes. Adv. Exp. Med. Biol. 2019, 1142, 153–167.
- Merzendorfer, H. The cellular basis of chitin synthesis in fungi and insects: Common principles and differences. Eur. J. Cell Biol. 2011, 90, 759–769.
- Watanabe, H.; Azuma, M.; Igarashi, K.; Ooshima, H. Analysis of Chitin at the Hyphal Tip of Candida albicans Using Calcofluor White. Biosci. Biotechnol. Biochem. 2005, 69, 1798–1801.
- Roncero, C. The genetic complexity of chitin synthesis in fungi. Curr. Genet. 2002, 41, 367–378.
- Zhang, J.; Jiang, H.; Du, Y.; Keyhani, N.O.; Xia, Y.; Jin, K. Members of chitin synthase family in Metarhizium acridum differentially affect fungal growth, stress tolerances, cell wall integrity and virulence. PLoS Pathog. 2019, 15, e1007964.
- Munro, C.A.; Gow, N.A.R. Chitin synthesis in human pathogenic fungi. Med. Mycol. 2001, 39, 41–53.
- Larson, T.M.; Kendra, D.F.; Busman, M.; Brown, D.W. Fusarium verticillioides chitin synthases CHS5 and CHS7 are required for normal growth and pathogenicity. Curr. Genet. 2011, 57, 177–189.
- Cui, Z.; Wang, Y.; Lei, N.; Wang, K.; Zhu, T. Botrytis cinerea chitin synthase BcChsVI is required for normal growth and pathogenicity. Curr. Genet. 2013, 59, 119–128.
- Takeshita, N.; Yamashita, S.; Ohta, A.; Horiuchi, H. Aspergillus nidulans class V and VI chitin synthases CsmA and CsmB, each with a myosin motor-like domain, perform compensatory functions that are essential for hyphal tip growth. Mol. Microbiol. 2006, 59, 1380–1394.
- Shu, M.; Lu, P.; Liu, S.; Zhang, S.; Gong, Z.; Cai, X.; Zhou, B.; Lin, Q.; Liu, J. Disruption of the Chitin Biosynthetic Pathway Results in Significant Changes in the Cell Growth Phenotypes and Biosynthesis of Secondary Metabolites of Monascus purpureus. J. Fungi 2022, 8, 910.
