A group of green algae in the order of Volvocales provides an ideal model system for studying the transition from unicellular to differentiated multicellularity. This group—known as the volvocine algae—evolved multicellularity relatively recently (~240 million years ago) and contains extant relatives that span a range of complexities from unicellularity, to undifferentiated multicellularity, to differentiated multicellularity. The regA-like gene family within the volvocine algae serves as a model for the evolution of the genetic basis of cellular differentiation.
- multicellularity
- cellular differentiation
- life history
- individuality
- gene co-option
- Volvox
- Chlamydomonas
- volvocine algae
- regA
1. The Volvocine Model System
2. regA Gene Structure and Function
Early investigations into cellular differentiation in V. carteri identified a class of mutants called “somatic regenerators” in which somatic cells appear to first develop normally but then dedifferentiate and become reproductive [44,45][22][23]. Linkage analysis found that all such regenerator mutants map onto a single locus which was named regA (from “regenerator”) [36,46,47][14][24][25]. Huskey & Griffin [47][25] originally described a second regB locus based on linkage group analysis of regenerator mutants, but reexamination of regB mutants by members of the same research lab determined that they are not regenerator mutants and have a different mutant phenotype [48][26]. Thus, in retrospect, all regenerator mutants can be mapped onto the regA locus [36][14]. However, it is worth noting that the annotation “RegA” or “Reg genes” has been used multiple times independently in species from other groups (e.g., bacteria and animals) to refer to different genes coding for distinct unrelated proteins. Such similarities in name are due to historical and linguistic coincidence rather than any shared function or homology. In this entreview, wey, researchers are strictly discussing the regA gene and its gene family that is restricted to volvocine algae. Based on the link between somatic regeneration and the regA locus, the regA gene was deemed the master regulatory gene that controls somatic cell development in V. carteri [36,37,49][14][15][27]. Kirk et al. [49][27] used transposon tagging to identify the regA gene and went on to determine that the RegA protein is localized in the nuclei of somatic cells. In V. carteri f. nagariensis, regA is expressed exclusively in somatic progenitor cells, with its transcription beginning early in development shortly after inversion [49,50,51,52][27][28][29][30]. regA transcript levels appear to persist and fluctuate throughout the life cycle [49][27], but see the study by König and Nedelcu [24][31] for an alternative possibility and discussion. The functional role of RegA, its amino acid composition, and the presence of a DNA-binding SAND domain in the RegA protein [53][32] helped establish the current working model in which RegA acts as a transcriptional repressor of genes needed for gonidial development [37][15]. A long-standing hypothesis is that regA suppresses the expression of nuclear-encoded chloroplast proteins required for chloroplast biogenesis and turn-over [54,55,56][33][34][35]. These negative effects on the chloroplasts would be reflected in the inability of the somatic cells to photosynthesize, grow, and divide. However, Matt and Umen [52][30] cast some doubt on this idea. They used whole transcriptome analysis to compare the expression profiles of germ cells and somatic cells. While photosynthetic genes were expressed at around two-fold higher levels in germ cells, photosynthetic genes were nevertheless highly abundant in somatic cells as well. Matt and Umen [52][30] propose that both germ cells and somatic cells maintain active photosynthesis, but germ cells are specialized in anabolic processes such as starch, fatty acid, and amino acid biosynthesis, while somatic cells break down starch and lipids to provide the substrates needed to synthesize ECM glycoproteins. Therefore, while it remains plausible that regA downregulates photosynthetic genes, it is also possible that regA downregulates other genes related to germ cell growth such as starch synthesis. The structure of the regA gene has been well described for V. carteri and serves as the basic template for the gene structures of many other homologs of regA in the VARL (volvocine algae regA-like) gene family. The minimal promoter of regA consists of only 42 nt found directly upstream of the transcription start site with a plausible TATA box with the sequence TAATTGA beginning at −28 and an initiator region with the sequence CACTCAT beginning -1 relative to the transcription start site [57][36]. The transcriptional unit of regA is 12,477 nt long and contains 7 introns and 8 exons. After the introns are spliced out, the mature regA mRNA is 6725 nt long and consists of a 940 nt 5′UTR (exons 1–5), a 3147 nt coding region (exons 5–8), and a 2638 nt 3′UTR with a UGUAA polyadenylation signal [49][27]. However, a splice variant that retains intron 7 (1194 bp) is expressed at low levels in V. carteri f. nagariensis as well. The donor splice site of intron 7 is GC instead of the typical GU, which may explain the variation in splicing. Remarkably, intron 7 encodes an ORF in the same frame as the rest of the regA coding region and, therefore, is likely to be translated, resulting in two different RegA protein products. However, experiments using modified regA transformation constructs to alter the splicing and translation of intron 7 have demonstrated that the presence or absence of intron 7 splicing has no detectable effect on the phenotypic rescue of regenerator mutants, despite the retention of intron 7 adding nearly 400 more amino acid residues to the RegA protein [57][36]. Interestingly, the homologous region to intron 7 is not spliced out in the closely related V. carteri f. kawasakiensis, and protein-level homology has been described in the intron 7 region across a wide variety of volvocine algae species [25,53][12][32]. Thus, it appears likely that splicing out intron 7 is a quirk specific to V. carteri f. nagariensis, while homologous regions are exonic in other species. In addition to the promoter, the differential transcription of regA is regulated by two enhancers found in introns 3 and 5 and a silencer found in intron 7 [57][36]. Eight possible AUG start codons are found in the 5′UTR of mature regA mRNA and are thought to be bypassed via a ribosome shunting mechanism so that translation begins at the ninth AUG sequence of the mRNA [58][37]. Following translation, the predicted RegA protein is 1049-amino-acids-long without the inclusion of intron 7 or 1447 with intron 7 and contains a high proportion of glutamine, alanine, and proline residues [49,57][27][36]. A key structural region within the RegA protein is the VARL domain, which is the distinguishing feature of the VARL gene family [53,59][32][38]. The VARL domain is located between amino acids 444 and 558 in the RegA of V. carteri f. nagariensis and is composed of a highly conserved core VARL region (sites 484–558), a short but highly conserved N-terminal extension region (sites 444–455), and a less conserved linker region between these two [25,53,59][12][32][38]. In addition, two short motifs of high amino acid conservation have been identified that are shared across the predicted RegA proteins of numerous volvocine algae species: a “LALRP” motif upstream of the VARL domain and an “FLQ” motif found within the intron 7 region downstream of the VARL domain [25][12]. The core VARL domain appears to encode a DNA-binding SAND domain [53][32]. The SAND domain (IPR000770/PF01342)—named after Sp100, AIRE-1, NucP41/75, and DEAF-1—is a DNA-binding domain found in animal and plant proteins that function in chromatin-dependent transcriptional control or bind-specific DNA sequences (e.g., [60][39]). SAND-containing proteins are involved in multiple distinct processes, both general and lineage/tissue-specific. However, most of the SAND-containing proteins with known functions are involved in multicellular development, including cell differentiation, cell proliferation, tissue homeostasis, and organ formation. For instance, DEAF-1 (Deformed Epidermal Autoregulatory Factor-1) is involved in breast epithelial cell differentiation in mammals [61][40] and is necessary for embryonic development in Drosophila melanogaster [62][41]. GMEB (Glucocorticoid Modulatory Element Binding) regulates neural apoptosis in the nematode Caenorhabditis elegans [63][42]. Spe44 (Speckled protein 44 kDa) is a master switch for germ cell fate in C. elegans and, like the mammalian AIRE1 (Autoimmune Regulator 1), plays a role in sperm cell differentiation [64,65,66][43][44][45]. In land plants, SAND domains are associated with ATX (the Arabidopsis homolog of trithorax) and ULTRAPETALA (ULT) proteins, which are involved in cell proliferation, cell differentiation, and tissue patterning. Specifically, ATX1 in Arabidopsis thaliana is required for root, leaf, and floral development through its histone methyltransferase activity [67][46], and ULT is a negative regulator that influences shoot and floral meristem size by controlling cell accumulation [68,69,70][47][48][49].3. regA-like Gene Family Evolution
The VARL gene family is defined by the presence of a homologous VARL domain within the predicted protein (note that volvocine algae possess additional SAND-containing proteins outside the VARL family). Although all VARL genes contain the VARL domain, the sequence level conservation outside of the VARL domain is very low. Thus, entire gene sequences cannot be aligned and used for phylogenetic analyses. The VARL domain itself is very short (~86 amino acids) and not highly conserved, such that its utility for inferring evolutionary relationships between the members of the VARL gene family is also limited. Nevertheless, information from gene synteny, sequence signatures outside of the VARL domain, and the locations of conserved introns can help draw more robust conclusions regarding the evolution of the VARL family [25].
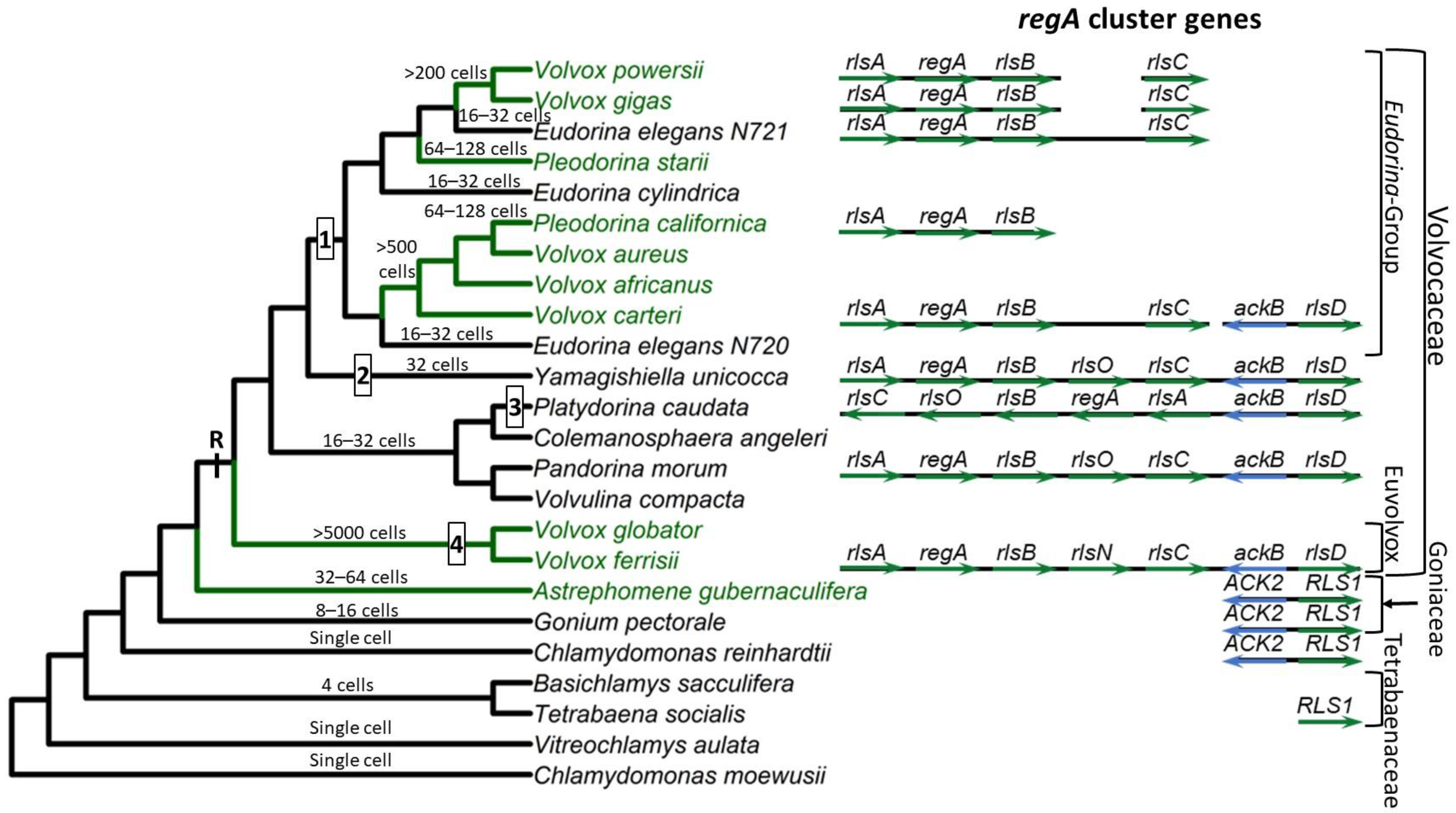
Based on currently available whole genome sequence data, the VARL gene family contains 12 members in C. reinhardtii [59][38], 8 in G. pectorale [32][8] and T. socialis [72][50], 6 in A. gubernaculifera [33][9], and 14 in V. carteri [59][38]. With the exception of regA orthologs (when present), all other regA homologs are known as regA-like sequences, annotated as RLS1-12 in Chlamydomonas and Goniaceae or rlsA-O in Volvocaceae. C. reinhardtii and other volvocine algae outside the Volvocaceae lack orthologs of any of the regA cluster genes. The closest homolog to the regA cluster genes found in these species is RLS1. This gene is an ortholog of the Volvocaceaen rlsD, which is the closest rls paralog of the regA cluster. Currently it is thought that the VARL gene family comprising several paralogs including RLS1/rlsD was already present in the common ancestor of all volvocine green algae. RLS1/rlsD underwent one or more duplication events in the common ancestor of the Volvocaceae family to give rise to a five-gene regA gene cluster comprising rlsA, regA, rlsB, rlsO, and rlsC. After the lineage leading to V. ferrisii diverged from the rest of the Volvocaceae, its rlsO gene gained a second VARL domain and evolved into rlsN. Meanwhile, the common ancestor of the Eudorina group lost rlsO. In addition, Y. unicocca lost two internal regA cluster genes (regA, rlsB, or rlsO) but restored the five-gene cluster via gene duplication, and the regA cluster of P. caudata became inverted relative to nearby syntenic markers (Figure 1).
Based on its role in suppressing reproduction in somatic cells, it has been hypothesized that regA evolved from a gene that was involved in trading off reproduction for survival (i.e., a life history trade-off gene) in the single-celled ancestors of V. carteri. Specifically, such a gene could have been co-opted by changing its expression from a temporal context (in response to an environmental cue) into a spatial context (in response to a developmental cue) [17][51].The common ancestor of V. carteri and C. reinhardtii likely had several VARL gene family members, one of which was RLS1. The RLS1 gene duplicated several times to give rise to the regA gene cluster in the common ancestor of the Volvocaceae, setting the stage for the functional co-option of regA during the evolution of cellular differentiation as well as other lineage-specific changes to regA cluster genes (Figure 1). The co-option of RLS1′s functions into a regA-like gene responsible for somatic cell differentiation likely involved the simulation of the ancestral environmentally induced signal in a developmental context.
References
- Coleman, A.W. A Comparative Analysis of the Volvocaceae (Chlorophyta). J. Phycol. 2012, 48, 491–513.
- Umen, J.G. Volvox and volvocine green algae. EvoDevo 2020, 11, 7–15.
- Herron, M.D.; Michod, R.E. Evolution of complexity in the volvocine algae: Transitions in individuality through Darwin’s eye. Evol. Int. J. Org. Evol. 2008, 62, 436–451.
- Kirk, D.L. A twelve-step program for evolving multicellularity and a division of labor. BioEssays News Rev. Mol. Cell. Dev. Biol. 2005, 27, 299–310.
- Nozaki, H.; Misawa, K.; Kajita, T.; Kato, M.; Nohara, S.; Watanabe, M.M. Origin and evolution of the colonial volvocales (Chlorophyceae) as inferred from multiple, chloroplast gene sequences. Mol. Phylogenet. Evol. 2000, 17, 256–268.
- Arakaki, Y.; Kawai-Toyooka, H.; Hamamura, Y.; Higashiyama, T.; Noga, A.; Hirono, M.; Olson, B.J.S.C.; Nozaki, H. The simplest integrated multicellular organism unveiled. PLoS ONE 2013, 8, e81641.
- Nozaki, H.; Itoh, M.; Watanabe, M.M.; Kuroiwa, T. Ultrastructure of the vegetative colonies and systematic position of basichlamys (Volvocales, Chlorophyta). Eur. J. Phycol. 1996, 31, 67–72.
- Hanschen, E.R.; Marriage, T.N.; Ferris, P.J.; Hamaji, T.; Toyoda, A.; Fujiyama, A.; Neme, R.; Noguchi, H.; Minakuchi, Y.; Suzuki, M.; et al. The Gonium pectorale genome demonstrates co-option of cell cycle regulation during the evolution of multicellularity. Nat. Commun. 2016, 7, 11370.
- Yamashita, S.; Yamamoto, K.; Matsuzaki, R.; Suzuki, S.; Yamaguchi, H.; Hirooka, S.; Minakuchi, Y.; Miyagishima, S.-Y.; Kawachi, M.; Toyoda, A.; et al. Genome sequencing of the multicellular alga Astrephomene provides insights into convergent evolution of germ-soma differentiation. Sci. Rep. 2021, 11, 22231.
- Nozaki, H.; Mahakham, W.; Athibai, S.; Yamamoto, K.; Takusagawa, M.; Misumi, O.; Herron, M.D.; Rosenzweig, F.; Kawachi, M. Rediscovery of the species of ‘ancestral Volvox’: Morphology and phylogenetic position of Pleodorina sphaerica (Volvocales, Chlorophyceae) from Thailand. ’ Phycologia 2017, 56, 469–475.
- Lindsey, C.R.; Rosenzweig, F.; Herron, M.D. Phylotranscriptomics points to multiple independent origins of multicellularity and cellular differentiation in the volvocine algae. BMC Biol. 2021, 19, 182.
- Grochau-Wright, Z.I.; Hanschen, E.R.; Ferris, P.J.; Hamaji, T.; Nozaki, H.; Olson, B.J.S.C.; Michod, R.E. Genetic Basis for Soma is Present in Undifferentiated Volvocine Green Algae. J. Evol. Biol. 2017, 30, 1205–1218.
- Herron, M.D.; Hackett, J.D.; Aylward, F.O.; Michod, R.E. Triassic origin and early radiation of multicellular volvocine algae. Proc. Natl. Acad. Sci. USA 2009, 106, 3254–3258.
- Kirk, D.L. Volvox: Molecular-Genetic Origins of Multicellularity and Cellular Differentiation; Cambridge University Press: Cambridge, UK, 1998.
- Kirk, D.L. Germ-soma differentiation in Volvox. Dev. Biol. 2001, 238, 213–223.
- Koufopanou, V. The Evolution of Soma in the Volvocales. Am. Nat. 1994, 143, 907–931.
- Kirk, M.M.; Ransick, A.; McRae, S.E.; Kirk, D.L. The Relationship between Cell Size and Cell Fate in Volvox carteri. J. Cell Biol. 1993, 123, 191–208.
- Grochau-Wright, Z.I. The Origin and Evolution of the Reg Cluster in the Volvocine Green Algae: A Model System for the Evolution of Cellular Differentiation. Ph.D. Dissertation, University of Arizona, Tucson, AZ, USA, 2019.
- Grochau-Wright, Z.I.; Ferris, P.J.; Tumberger, J.; Jiménez-Marin, B.; Olson, B.J.S.C.; Michod, R.E. Characterization and Transformation of reg Cluster Genes in Volvox powersii Enable Investigation of Convergent Evolution of Cellular Differentiation in Volvox. Protist 2021, 172, 125834.
- Ransick, A. Reproductive cell specification during Volvox obversus development. Dev. Biol. 1991, 143, 185–198.
- Ransick, A. Specification of reproductive cells in Volvox. In Evolutionary Conservation of Developmental Mechanisms, Proceedings of the 50th Symposium of the Society for Developmental Biology, Marquette University, Milwaukee, WI, USA, 20–23 June 1991; Spradling, A., Ed.; Wiley-Liss: Hoboken, NJ, USA, 1993; pp. 55–70.
- Sessoms, A.H.; Huskey, R.J. Genetic Control of Development in Volvox: Isolation and Characterization of Morphogenetic Mutants. Proc. Natl. Acad. Sci. USA 1973, 70, 1335–1338.
- Starr, R.C. Control of differentiation in Volvox. Dev. Biol. 1970, 4, 59–100.
- Huskey, R.J.; Griffin, B.E.; Cecil, P.O.; Callahan, A.M. A Preliminary Genetic Investigation of Volvox carteri. Genetics 1979, 91, 229–244.
- Huskey, R.J.; Griffin, B.E. Genetic Control of Somatic Cell Differentiation in Volvox. Dev. Biol. 1979, 72, 226–235.
- Baran, G. Analysis of Somatic Cell Differentiation in Volvox carteri f. nagariensis; University of Virginia: Charlottesville, VA, USA, 1984.
- Kirk, M.M.; Stark, K.; Miller, S.M.; Müller, W.; Taillon, B.E.; Gruber, H.; Schmitt, R.; Kirk, D.L. regA, a Volvox gene that plays a central role in germ-soma differentiation, encodes a novel regulatory protein. Development 1999, 126, 639–647.
- Harryman, A. Investigating the Roles of regA and Related Genes in the Evolution of Multicellularity in the Volvocine Green Algae. Ph.D. Thesis, University of Maryland, Baltimore, MD, USA, 2012.
- Klein, B.; Wibberg, D.; Hallmann, A. Whole transcriptome RNA-Seq analysis reveals extensive cell type-specific compartmentalization in Volvox carteri. BMC Biol. 2017, 15, 1–22.
- Matt, G.Y.; Umen, J.G. Cell-Type Transcriptomes of the Multicellular Green Alga Volvox carteri Yield Insights into the Evolutionary Origins of Germ and Somatic Differentiation Programs. G3 Genes|Genomes|Genet. 2018, 8, 531–550.
- König, S.G.; Nedelcu, A.M. The genetic basis for the evolution of soma: Mechanistic evidence for the co-option of a stress-induced gene into a developmental master regulator. Proc. R. Soc. B Biol. Sci. 2020, 287, 20201414.
- Duncan, L.; Nishii, I.; Howard, A.; Kirk, D.; Miller, S.M. Orthologs and paralogs of regA, a master cell-type regulatory gene in Volvox carteri. Curr. Genet. 2006, 50, 61–72.
- Choi, G.; Przybylska, M.; Straus, D. Three abundant germ line-specific transcripts in Volvox carteri encode photosynthetic proteins. Curr. Genet. 1996, 30, 347–355.
- Meissner, M.; Stark, K.; Cresnar, B.; Kirk, D.L.; Schmitt, R. Volvox germline-specific genes that are putative targets of RegA repression encode chloroplast proteins. Curr. Genet. 1999, 36, 363–370.
- Tam, L.; Kirk, D.L. Identification of Cell-Type-Specific Characterization of their Expression Genes of Volvox carteri and during the Asexual Life Cycle. Dev. Biol. 1991, 145, 51–66.
- Stark, K.; Kirk, D.L.; Schmitt, R. Two enhancers and one silencer located in the introns of regA control somatic cell differentiation in Volvox carteri. Genes Dev. 2001, 15, 1449–1460.
- Babinger, K.; Hallmann, A.; Schmitt, R. Translational control of regA, a key gene controlling cell differentiation in Volvox carteri. Development 2006, 133, 4045–4051.
- Duncan, L.; Nishii, I.; Harryman, A.; Buckley, S.; Howard, A.; Friedman, N.R.; Miller, S.M. The VARL gene family and the evolutionary origins of the master cell-type regulatory gene, regA, in Volvox carteri. J. Mol. Evol. 2007, 65, 1–11.
- Bottomley, M.J.J.; Collard, M.W.W.; Huggenvik, J.I.I.; Liu, Z.; Gibson, T.J.J.; Sattler, M. The SAND domain structure defines a novel DNA-binding fold in transcriptional regulation. Nat. Struct. Biol. 2001, 8, 626–633.
- Barker, H.E.; Smyth, G.K.; Wettenhall, J.; Ward, T.A.; Bath, M.L.; Lindeman, G.J.; Visvader, J.E. Deaf-1 regulates epithelial cell proliferation and side-branching in the mammary gland. BMC Dev. Biol. 2008, 8, 94.
- Veraksa, A.; Kennison, J.; McGinnis, W. DEAF-1 function is essential for the early embryonic development of Drosophila. Genesis 2002, 33, 67–76.
- Nakagawa, T.; Tsuruma, K.; Uehara, T.; Nomura, Y. GMEB1, a novel endogenous caspase inhibitor, prevents hypoxia- and oxidative stress-induced neuronal apoptosis. Neurosci. Lett. 2008, 438, 34–37.
- Kulkarni, M.; Shakes, D.C.; Guevel, K.; Smith, H.E. SPE-44 Implements Sperm Cell Fate. PLoS Genet. 2012, 8, e1002678.
- Radhakrishnan, K.; Bhagya, K.P.; Kumar, A.T.; Devi, A.N.; Sengottaiyan, J.; Kumar, P.G. Autoimmune Regulator (AIRE) Is Expressed in Spermatogenic Cells, and It Altered the Expression of Several Nucleic-Acid-Binding and Cytoskeletal Proteins in Germ Cell 1 Spermatogonial (GC1-spg) Cells. Mol. Cell. Proteom. 2016, 15, 2686–2698.
- Schaller, C.E.; Wang, C.L.; Beck-engeser, G.; Goss, L.; Scott, H.S.; Anderson, M.S.; Wabl, M. Expression of Aire and the Early Wave of Apoptosis in Spermatogenesis. J. Immunol. 2008, 180, 1338–1343.
- Chen, L.-Q.; Luo, J.-H.; Cui, Z.-H.; Xue, M.; Wang, L.; Zhang, X.-Y.; Pawlowski, W.P.; He, Y. ATX3, ATX4, and ATX5 Encode Putative H3K4 Methyltransferases and Are Critical for Plant Development. Plant Physiol. 2017, 174, 1795–1806.
- Carles, C.C. ULTRAPETALA1 encodes a SAND domain putative transcriptional regulator that controls shoot and floral meristem activity in Arabidopsis. Development 2005, 132, 897–911.
- Carles, C.C.; Fletcher, J.C. The SAND domain protein ULTRAPETALA1 acts as a trithorax group factor to regulate cell fate in plants. Genes Dev. 2009, 23, 2723–2728.
- Fletcher, J.C. The ULTRAPETALA gene controls shoot and floral meristem size in Arabidopsis. Development 2001, 128, 1323–1333.
- Featherston, J.; Arakaki, Y.; Hanschen, E.R.; Ferris, P.J.; Michod, R.E.; Olson, B.J.S.C.; Nozaki, H.; Durand, P.M. The 4-celled tetrabaena socialis nuclear genome reveals the essential components for genetic control of cell number at the origin of multicellularity in the volvocine lineage. Mol. Biol. Evol. 2018, 35, 855–870.
- Nedelcu, A.M.; Michod, R.E. The evolutionary origin of an altruistic gene. Mol. Biol. Evol. 2006, 23, 1460–1464.
