The CHD (chromodomain-helicase-DNA binding) family consists of nine members (CHD1–CHD9). Their structure is characterised by two consecutive chromodomains in the N-terminal region and an SNF2-like (sucrose-non-fermenting (SNF)) ATP-dependent helicase domain positioned in the central region. They recognise and bind nucleosomes to contribute, in many cases, to the formation of heterochromatin typically marked by the presence of methylated histones and other repressive chromatin markers. CHD proteins can bind to methylated histones through their chromodomains and use their helicase activity to remodel the chromatin and contribute to the formation of heterochromatin, a fundamental feature of chromosomes that ensures genomic stability.
- cochlea
- hair cells
- epigenetic
- chromatin remodellers
- CHD
1. CHD Family
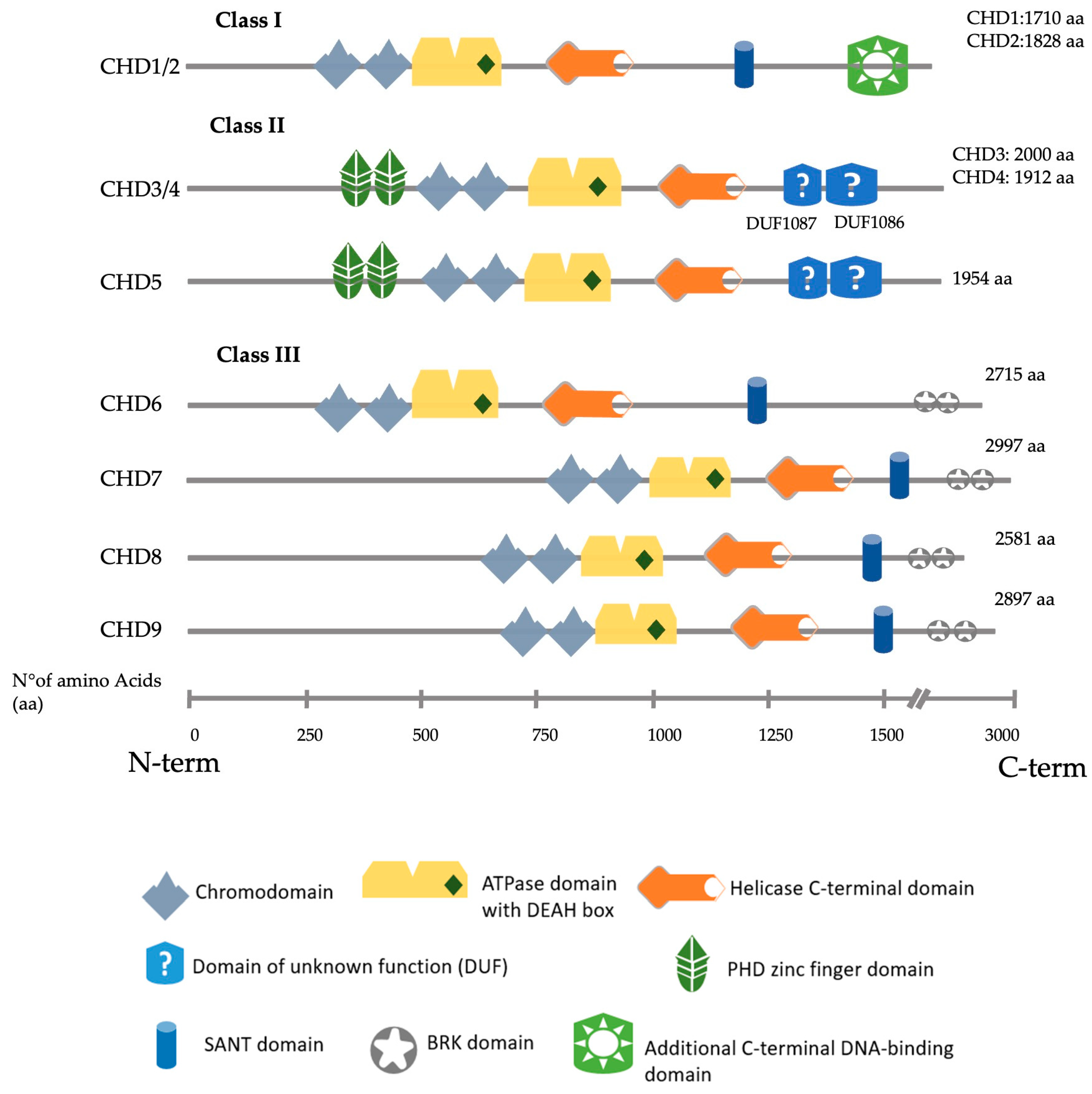
2. CHD3/4
CHD3 and CDH4 share 71.6% of their amino acid sequences. Both proteins form distinct nucleosome remodelling deacetylase (NuRD) complexes with different but overlapping functionalities [9]. Mass spectrometry data show that CHD3 and CHD4 may form heterodimers within NuRD complexes [10]. CHD3 and CHD4 are the core subunit of the NuRD complex [11]. Their binding to this complex is mutually exclusive [12]. Normal NuRD complexes are essential for many developmental and cell differentiation processes, including maintaining embryonic stem (ES) cell pluripotency and regulating progenitor cell development in numerous organs such as the brain, hematopoietic system or kidney [13][14]. Snijders Blok–Campeau syndrome is a recently described syndrome that encompasses patients with variants in CHD3, characterised by intellectual disability, developmental delay (especially speech delay) and several dysmorphic features [15], with most of the mutations localising within the ATPase or the helicase domain [16]. A minority of these patients manifest neurosensory hearing loss (3 out of 24). More investigations are needed to understand further the implications of hearing deficit in this syndrome. Mutations in the CHD4 protein result in Sifrim–Hitz–Weiss (SIHIWES) syndrome, an autosomal dominant neurodevelopmental disease. Symptoms identified include global developmental delay, mild to moderate intellectual disability, brain anomalies, congenital heart defects, dysmorphic features, macrocephaly and conductive and/or sensorineural hearing loss in 55% of the cases [17]. The mutations identified in hearing loss patients affect the ATPase domain, the PHD domain or the DUF1087 domain (Figure 2) [17][18].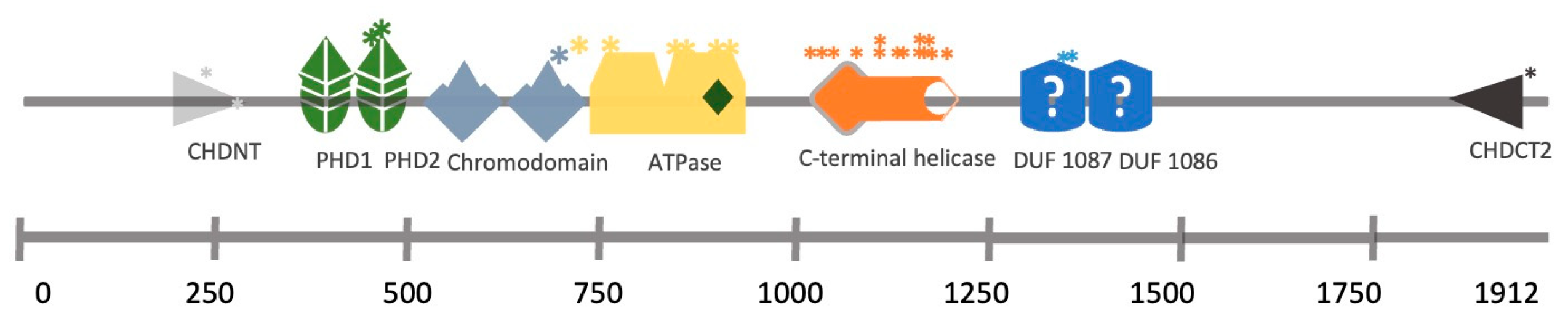
3. CHD7
De novo heterozygous mutations in the CHD7 gene are the leading causes of CHARGE syndrome, a complex neurodevelopmental disorder characterised by ocular Coloboma, Heart defects, Atresia of the choanae, Retarded growth and development, Genital hypoplasia and Ear abnormalities, including mixed conductive/sensorineural hearing loss [21][22]. More than 500 heterozygous CHD7 mutations have been identified in CHARGE syndrome patients, most of which are nonsense, frameshift indels, splice site and missense mutations [23]. In mice, Chd7 is highly expressed throughout the developing otocyst starting at E9.5. Later during development, Chd7 is present in the cochlear epithelium both in hair cells (HCs) and SCs and in spiral ganglion neurons. Chd7 is still present postnatally but is decreased in the organ of Corti [24][25]. A Chd7 heterozygous mouse model was generated from Chd7-deficient, gene-trapped lacZ reporter ES cells (hereafter Chd7Gt/+ mice) that expresses β-galactosidase under the control of the Chd7 promoter [26]. Homozygous Chd7Gt/Gt are embryonic lethal. HC formation and cochlear innervation were normal in Chd7Gt/+ mice. However, Chd7Gt/+ animals present mild hearing loss with elevated auditory brainstem recordings (ABR) and reduced distortion product otoacoustic emissions (DPOAE) [26]. Recently, a re-evaluation of Chd7Gt/+ suggests a new function in the development and maintenance of satellite glial cells in the spiral ganglia, as well as in the regulation of myelin sheaths surrounding type I spiral ganglion neurons that innervate the sensory epithelium of the cochlea [27]. In two ENU-induced mutations in Chd7—looper and trooper strains—having a nonsense mutation (c.5690C>A, p.S1897X) or a cryptic splice site mutation (c.3219-18T>A), respectively, middle ear and vestibular defects were found to be a prominent phenotype with early onset hearing loss [28][29]. Deficiency in Chd7 during development has also been reported to specifically impact neuronal development in the inner ear. In a Foxg1-Cre-driven conditional Chd7 knockout mouse, the otocyst had a reduced vestibulocochlear ganglion size. This smaller neuronal number is due to the reduced expression of the neuronal fate-specific genes Ngn1, Otx2 and Fgf10, leading to reduced neuronal cell proliferation [24][30]. A recent study has shown that Chd7 deficiency leads to IHC and auditory neurons being vulnerable to ototoxic stress, rapid postnatal degeneration and profound hearing loss [31]. Modulating retinoic acid signalling prevents inner ear defects caused by Chd7 deficiency [32]. Indeed, loss of the retinoic acid (RA) synthetic enzyme Aldh1a3 or treatment with citral—an inhibitor of retinoic acid (RA) synthesis—partially rescued the semicircular canal deformation phenotype in Chd7Gt/+ mice [33]. Upregulation or downregulation of RA signalling during embryogenesis leads to developmental defects similar to those observed in CHARGE. RNA-seq and qRT-PCR combined with ChIP experiments in inner ears demonstrate that Chd7 acts upstream of Aldh1a3 via direct binding and repression of Aldh1a3 [33]. To evaluate the Chd7 deficiency at single-cell resolution, Durruthy-Durruthy et al. performed single-cell multiplexed qPCR in Pax2Cre; mT/mGFP otocysts cells from Chd7+/+ and Chd7Gt/+ mice for 192 inner ear specific genes and reported a cellular identity shift towards neuroblasts in E10.5 otocysts [34]. This cell fate decision may arise through disruption in the pro-sensory and pro-neurogenic gene expression network and Notch signalling pathways. To model the inner ear phenotypes in CHARGE syndrome in a human context, Nie et al. adopted a previously published human induced pluripotent stem (hiPS) cell/human ES cell-derived otic organoid model [35]. The study reported a complete loss of inner ear HC formation in the CHD7 knock-out line (CHD7−/−) or CHD7 patient-specific missense mutation (CHD7S834F/S834F) d70 otic organoids [36]. In contrast, the haploinsufficient/heterozygous otic organoids (CHD7S834F/+ or CHD7+/−) could generate HCs. To understand the molecular pathogenesis, the CHD7 mutations were created in a WA25 PAX2nGFP hESC line, which allows for the study of the transcriptome of PAX2+ otic progenitors from both CHD7+/+ and CHD7−/− d20 organoids. Single-cell RNA-sequencing (scRNA-seq) transcriptome data on those PAX2+ cells revealed a downregulation of several essential inner ear morphogenesis and developmental genes such as TBX1, LMX1A, SOX10, DLX5 and SIX1 and disruption in FGF, BMP, Notch, TGF-β and Wnt signalling pathways. These effects on PAX2 otic progenitors were reflected in HC development. Indeed, scRNA-seq on POU4F3-nd tomato hair cells from CHD7+/− d70 organoids showed numerous dysregulated deafness genes compared with the WT counterparts. SIX1, USH1C and STRC were downregulated, giving potential explanations for hearing loss observed in patients with CHARGE syndrome. Co-differentiating both WT and CHD7−/− hESCs in a chimeric organoid system partially restored the expression of otherwise severely downregulated essential otic genes FBXO2, SOX10 and DLX5 but failed to generate any hair cells in the CHD7−/− background [36]. Altogether, this study described the critical role of CHD7 in otic lineage differentiation and hair cell development and, more importantly, in an in-vitro humanised experimental model.4. CHD8
CHD8 is most prominently known for its association with autism spectrum disorders (ASD) [37]. CHD8 has two isoforms: CHD8L, a full-length version of the protein weighing 280 kDa, and CHD8S (Duplin), a 110 kDa protein of the NH2-terminal chromodomain region. However, the latter does not have a counterpart in humans [38]. CHD8 interacts with CHD7 and FAM124B (family with sequence similarity 124B) as a complex [39][40]. However, the functional implication of this interaction in the context of inner ear development and deafness remains elusive. Chd8 has been found in the proteome of HCs from P4–P7 organs of Corti derived from a Pou4f3-eGFP transgenic mouse model [41]. It has been suggested as a candidate deafness gene (DFNA53) with a possible functional role in HCs. In another study, RNA-seq data showed that CHD8 knock-out hES cell-derived neuroectodermal cells are enriched in genes for “inner ear morphogenesis” gene-ontology terms such as PAX8, FGF9 and MYO15A [42]. This observed increase in “inner ear” genes might arise since CHD8 has been shown to interact directly with β-catenin and be recruited to the promoter regions of numerous β-catenin-responsive genes acting as a repressor of both β-catenin and the Wnt pathway [43][44]. Several identified genes (PAX8, DLX6, FGF9) are upstream or downstream effectors of β-catenin/Wnt pathway modulation [45][46]. In a recent study, the deletion of Chd8 in mice under the control of the Atoh1 promoter—expressed explicitly in HCs and spiral ganglion neurons—did not affect ABR [47]. Moreover, the development of HCs and auditory neurons seemed normal. Therefore, Chd8 does not appear to be necessary for cochlea development and hearing function in mice, while it seems essential in humans. The functional role of CHD8 in inner ear development and hearing loss remains to be established.References
- Woodage, T.; Basrai, M.A.; Baxevanis, A.D.; Hieter, P.; Collins, F.S. Characterization of the CHD family of proteins. Proc. Natl. Acad. Sci. USA 1997, 94, 11472–11477.
- Marfella, C.G.; Imbalzano, A.N. The Chd family of chromatin remodelers. Mutat. Res. 2007, 618, 30–40.
- Liu, X.R.; Ye, T.T.; Zhang, W.J.; Guo, X.; Wang, J.; Huang, S.P.; Xie, L.S.; Song, X.W.; Deng, W.W.; Li, B.M.; et al. CHD4 variants are associated with childhood idiopathic epilepsy with sinus arrhythmia. CNS Neurosci. Ther. 2021, 27, 1146–1156.
- Allen, M.D.; Religa, T.L.; Freund, S.M.; Bycroft, M. Solution structure of the BRK domains from CHD7. J. Mol. Biol. 2007, 371, 1135–1140.
- Alendar, A.; Berns, A. Sentinels of chromatin: Chromodomain helicase DNA-binding proteins in development and disease. Genes Dev. 2021, 35, 1403–1430.
- UniProt, C. UniProt: The universal protein knowledgebase in 2021. Nucleic Acids Res. 2021, 49, D480–D489.
- Letunic, I.; Bork, P. 20 years of the SMART protein domain annotation resource. Nucleic Acids Res. 2018, 46, D493–D496.
- Blum, M.; Chang, H.Y.; Chuguransky, S.; Grego, T.; Kandasaamy, S.; Mitchell, A.; Nuka, G.; Paysan-Lafosse, T.; Qureshi, M.; Raj, S.; et al. The InterPro protein families and domains database: 20 years on. Nucleic Acids Res. 2021, 49, D344–D354.
- Hoffmeister, H.; Fuchs, A.; Erdel, F.; Pinz, S.; Grobner-Ferreira, R.; Bruckmann, A.; Deutzmann, R.; Schwartz, U.; Maldonado, R.; Huber, C.; et al. CHD3 and CHD4 form distinct NuRD complexes with different yet overlapping functionality. Nucleic Acids Res. 2017, 45, 10534–10554.
- Kloet, S.L.; Baymaz, H.I.; Makowski, M.; Groenewold, V.; Jansen, P.W.; Berendsen, M.; Niazi, H.; Kops, G.J.; Vermeulen, M. Towards elucidating the stability, dynamics and architecture of the nucleosome remodeling and deacetylase complex by using quantitative interaction proteomics. FEBS J. 2015, 282, 1774–1785.
- Lai, A.Y.; Wade, P.A. Cancer biology and NuRD: A multifaceted chromatin remodelling complex. Nat. Rev. Cancer 2011, 11, 588–596.
- Tong, J.K.; Hassig, C.A.; Schnitzler, G.R.; Kingston, R.E.; Schreiber, S.L. Chromatin deacetylation by an ATP-dependent nucleosome remodelling complex. Nature 1998, 395, 917–921.
- Zhao, H.; Han, Z.; Liu, X.; Gu, J.; Tang, F.; Wei, G.; Jin, Y. The chromatin remodeler Chd4 maintains embryonic stem cell identity by controlling pluripotency- and differentiation-associated genes. J. Biol. Chem. 2017, 292, 8507–8519.
- Hirota, A.; Nakajima-Koyama, M.; Ashida, Y.; Nishida, E. The nucleosome remodeling and deacetylase complex protein CHD4 regulates neural differentiation of mouse embryonic stem cells by down-regulating p53. J. Biol. Chem. 2019, 294, 195–209.
- Snijders Blok, L.; Rousseau, J.; Twist, J.; Ehresmann, S.; Takaku, M.; Venselaar, H.; Rodan, L.H.; Nowak, C.B.; Douglas, J.; Swoboda, K.J.; et al. CHD3 helicase domain mutations cause a neurodevelopmental syndrome with macrocephaly and impaired speech and language. Nat. Commun. 2018, 9, 4619.
- Drivas, T.G.; Li, D.; Nair, D.; Alaimo, J.T.; Alders, M.; Altmuller, J.; Barakat, T.S.; Bebin, E.M.; Bertsch, N.L.; Blackburn, P.R.; et al. A second cohort of CHD3 patients expands the molecular mechanisms known to cause Snijders Blok-Campeau syndrome. Eur. J. Hum. Genet. 2020, 28, 1422–1431.
- Weiss, K.; Lazar, H.P.; Kurolap, A.; Martinez, A.F.; Paperna, T.; Cohen, L.; Smeland, M.F.; Whalen, S.; Heide, S.; Keren, B.; et al. The CHD4-related syndrome: A comprehensive investigation of the clinical spectrum, genotype-phenotype correlations, and molecular basis. Genet. Med. 2020, 22, 389–397.
- Farnung, L.; Ochmann, M.; Cramer, P. Nucleosome-CHD4 chromatin remodeler structure maps human disease mutations. Elife 2020, 9, e56178.
- Layman, W.S.; Sauceda, M.A.; Zuo, J. Epigenetic alterations by NuRD and PRC2 in the neonatal mouse cochlea. Hear. Res. 2013, 304, 167–178.
- Nitarska, J.; Smith, J.G.; Sherlock, W.T.; Hillege, M.M.; Nott, A.; Barshop, W.D.; Vashisht, A.A.; Wohlschlegel, J.A.; Mitter, R.; Riccio, A. A Functional Switch of NuRD Chromatin Remodeling Complex Subunits Regulates Mouse Cortical Development. Cell Rep. 2016, 17, 1683–1698.
- Choo, D.I.; Tawfik, K.O.; Martin, D.M.; Raphael, Y. Inner ear manifestations in CHARGE: Abnormalities, treatments, animal models, and progress toward treatments in auditory and vestibular structures. Am. J. Med. Genet. C Semin Med. Genet. 2017, 175, 439–449.
- Van Ravenswaaij-Arts, C.; Martin, D.M. New insights and advances in CHARGE syndrome: Diagnosis, etiologies, treatments, and research discoveries. Am. J. Med. Genet. C Semin Med. Genet. 2017, 175, 397–406.
- Basson, M.A.; van Ravenswaaij-Arts, C. Functional Insights into Chromatin Remodelling from Studies on CHARGE Syndrome. Trends Genet. 2015, 31, 600–611.
- Balendran, V.; Skidmore, J.M.; Ritter, K.E.; Gao, J.; Cimerman, J.; Beyer, L.A.; Hurd, E.A.; Raphael, Y.; Martin, D.M. Chromatin remodeler CHD7 is critical for cochlear morphogenesis and neurosensory patterning. Dev. Biol. 2021, 477, 11–21.
- Bosman, E.A.; Penn, A.C.; Ambrose, J.C.; Kettleborough, R.; Stemple, D.L.; Steel, K.P. Multiple mutations in mouse Chd7 provide models for CHARGE syndrome. Hum. Mol. Genet. 2005, 14, 3463–3476.
- Hurd, E.A.; Capers, P.L.; Blauwkamp, M.N.; Adams, M.E.; Raphael, Y.; Poucher, H.K.; Martin, D.M. Loss of Chd7 function in gene-trapped reporter mice is embryonic lethal and associated with severe defects in multiple developing tissues. Mamm. Genome 2007, 18, 94–104.
- Ritter, K.E.; Lynch, S.M.; Gorris, A.M.; Beyer, L.A.; Kabara, L.; Dolan, D.F.; Raphael, Y.; Martin, D.M. Loss of the chromatin remodeler CHD7 impacts glial cells and myelination in the mouse cochlear spiral ganglion. Hear. Res. 2022, 426, 108633.
- Ogier, J.M.; Arhatari, B.D.; Carpinelli, M.R.; McColl, B.K.; Wilson, M.A.; Burt, R.A. An intronic mutation in Chd7 creates a cryptic splice site, causing aberrant splicing in a mouse model of CHARGE syndrome. Sci. Rep. 2018, 8, 5482.
- Ogier, J.M.; Carpinelli, M.R.; Arhatari, B.D.; Symons, R.C.; Kile, B.T.; Burt, R.A. CHD7 deficiency in “Looper”, a new mouse model of CHARGE syndrome, results in ossicle malformation, otosclerosis and hearing impairment. PLoS ONE 2014, 9, e97559.
- Hurd, E.A.; Poucher, H.K.; Cheng, K.; Raphael, Y.; Martin, D.M. The ATP-dependent chromatin remodeling enzyme CHD7 regulates pro-neural gene expression and neurogenesis in the inner ear. Development 2010, 137, 3139–3150.
- Ahmed, M.; Moon, R.; Prajapati, R.S.; James, E.; Basson, M.A.; Streit, A. The chromatin remodelling factor Chd7 protects auditory neurons and sensory hair cells from stress-induced degeneration. Commun. Biol. 2021, 4, 1260.
- Micucci, J.A.; Layman, W.S.; Hurd, E.A.; Sperry, E.D.; Frank, S.F.; Durham, M.A.; Swiderski, D.L.; Skidmore, J.M.; Scacheri, P.C.; Raphael, Y.; et al. CHD7 and retinoic acid signaling cooperate to regulate neural stem cell and inner ear development in mouse models of CHARGE syndrome. Hum. Mol. Genet. 2014, 23, 434–448.
- Yao, H.; Hill, S.F.; Skidmore, J.M.; Sperry, E.D.; Swiderski, D.L.; Sanchez, G.J.; Bartels, C.F.; Raphael, Y.; Scacheri, P.C.; Iwase, S.; et al. CHD7 represses the retinoic acid synthesis enzyme ALDH1A3 during inner ear development. JCI Insight 2018, 3, e97440.
- Durruthy-Durruthy, R.; Sperry, E.D.; Bowen, M.E.; Attardi, L.D.; Heller, S.; Martin, D.M. Single Cell Transcriptomics Reveal Abnormalities in Neurosensory Patterning of the Chd7 Mutant Mouse Ear. Front. Genet. 2018, 9, 473.
- Koehler, K.R.; Nie, J.; Longworth-Mills, E.; Liu, X.P.; Lee, J.; Holt, J.R.; Hashino, E. Generation of inner ear organoids containing functional hair cells from human pluripotent stem cells. Nat. Biotechnol. 2017, 35, 583–589.
- Nie, J.; Ueda, Y.; Solivais, A.J.; Hashino, E. CHD7 regulates otic lineage specification and hair cell differentiation in human inner ear organoids. Nat. Commun. 2022, 13, 7053.
- Sugathan, A.; Biagioli, M.; Golzio, C.; Erdin, S.; Blumenthal, I.; Manavalan, P.; Ragavendran, A.; Brand, H.; Lucente, D.; Miles, J.; et al. CHD8 regulates neurodevelopmental pathways associated with autism spectrum disorder in neural progenitors. Proc. Natl. Acad. Sci. USA 2014, 111, E4468–E4477.
- Nishiyama, M.; Oshikawa, K.; Tsukada, Y.; Nakagawa, T.; Iemura, S.; Natsume, T.; Fan, Y.; Kikuchi, A.; Skoultchi, A.I.; Nakayama, K.I. CHD8 suppresses p53-mediated apoptosis through histone H1 recruitment during early embryogenesis. Nat. Cell Biol. 2009, 11, 172–182.
- Batsukh, T.; Schulz, Y.; Wolf, S.; Rabe, T.I.; Oellerich, T.; Urlaub, H.; Schaefer, I.M.; Pauli, S. Identification and characterization of FAM124B as a novel component of a CHD7 and CHD8 containing complex. PLoS ONE 2012, 7, e52640.
- Batsukh, T.; Pieper, L.; Koszucka, A.M.; von Velsen, N.; Hoyer-Fender, S.; Elbracht, M.; Bergman, J.E.; Hoefsloot, L.H.; Pauli, S. CHD8 interacts with CHD7, a protein which is mutated in CHARGE syndrome. Hum. Mol. Genet. 2010, 19, 2858–2866.
- Hickox, A.E.; Wong, A.C.; Pak, K.; Strojny, C.; Ramirez, M.; Yates, J.R., III; Ryan, A.F.; Savas, J.N. Global Analysis of Protein Expression of Inner Ear Hair Cells. J. Neurosci. 2017, 37, 1320–1339.
- Ding, S.; Lan, X.; Meng, Y.; Yan, C.; Li, M.; Li, X.; Chen, J.; Jiang, W. CHD8 safeguards early neuroectoderm differentiation in human ESCs and protects from apoptosis during neurogenesis. Cell Death Dis. 2021, 12, 981.
- Thompson, B.A.; Tremblay, V.; Lin, G.; Bochar, D.A. CHD8 is an ATP-dependent chromatin remodeling factor that regulates beta-catenin target genes. Mol. Cell Biol. 2008, 28, 3894–3904.
- Nishiyama, M.; Skoultchi, A.I.; Nakayama, K.I. Histone H1 recruitment by CHD8 is essential for suppression of the Wnt-beta-catenin signaling pathway. Mol. Cell Biol. 2012, 32, 501–512.
- DeJonge, R.E.; Liu, X.P.; Deig, C.R.; Heller, S.; Koehler, K.R.; Hashino, E. Modulation of Wnt Signaling Enhances Inner Ear Organoid Development in 3D Culture. PLoS ONE 2016, 11, e0162508.
- Wright, K.D.; Mahoney Rogers, A.A.; Zhang, J.; Shim, K. Cooperative and independent functions of FGF and Wnt signaling during early inner ear development. BMC Dev. Biol. 2015, 15, 33.
- Kawamura, A.; Katayama, Y.; Kakegawa, W.; Ino, D.; Nishiyama, M.; Yuzaki, M.; Nakayama, K.I. The autism-associated protein CHD8 is required for cerebellar development and motor function. Cell Rep. 2021, 35, 108932.
