ThiRes articleearchers presents a comprehensive survey on the diagnosis of colon cancer. This covers many aspects related to colon cancer, such as its symptoms and grades as well as the available imaging modalities (particularly, histopathology images used for analysis) in addition to common diagnosis systems. Furthermore, the most widely used datasets and performance evaluation metrics are discussed. WeResearchers provide a comprehensive review of the current studies on colon cancer, classified into deep-learning (DL) and machine-learning (ML) techniques, and weresearchers identify their main strengths and limitations. These techniques provide extensive support for identifying the early stages of cancer that lead to early treatment of the disease and produce a lower mortality rate compared with the rate produced after symptoms develop. In addition, these methods can help to prevent colorectal cancer from progressing through the removal of pre-malignant polyps, which can be achieved using screening tests to make the disease easier to diagnose. Finally, the existing challenges and future research directions that open the way for future work in this field are presented.
- colon cancer diagnosis
- imaging modalities
- deep-learning techniques
1. Introduction
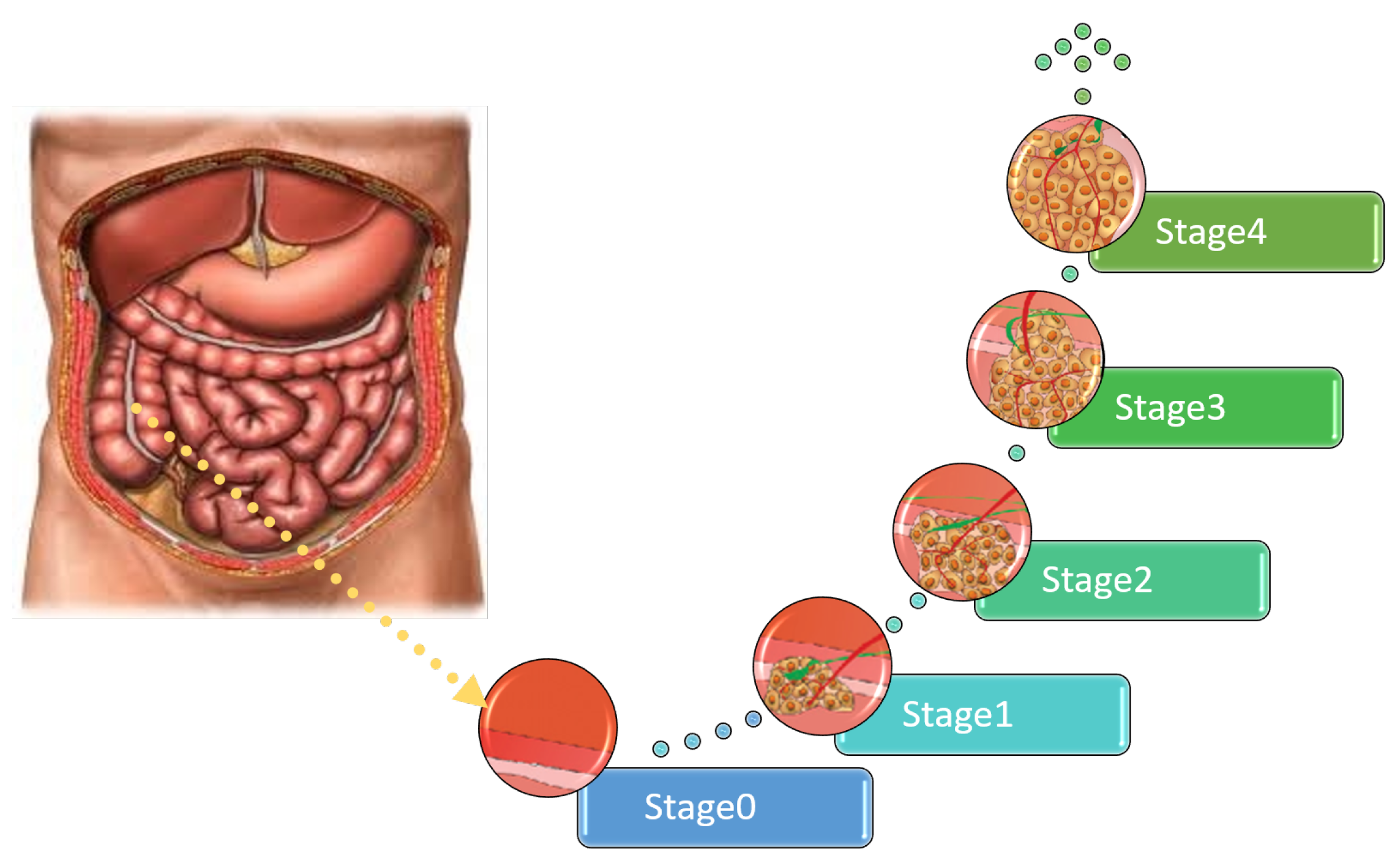
| No. | Spread to Various Organs | Symptoms |
|---|---|---|
| 1 | Liver |
|
| 2 | Lung |
|
| 3 | Bone |
|
| 4 | Lymph nodes |
|
2. Colon Cancer Diagnosis
Before going in depth and reviewing the current work on colon cancer diagnosis, many aspects related to the diagnosis process should be taken into consideration, such as the image modality used, type of diagnosis system, the dataset used, and the metrics used for evaluation. Therefore, in the following subsections, researchers discuss these aspects.2.1. Imaging Modalities
As mentioned before, ourthe main goal is the automatic diagnosis of colon cancer with high detection accuracy and without manual intervention. In this section, rResearchers take an in-depth look at the different imaging modalities recently applied for MIA, including Computed Tomography (CT), Endorectal Ultrasound (ERUS), virtual Computed Tomography Colonoscopy (CTC), and Magnetic Resonance Imaging (MRI) [22][19] in addition to other modalities, such as Histopathological Imaging (HI) and Positron Emission Tomography (PET) [23][20].2.2. Common Diagnosis Systems Based on HI Analysis
In tThis paper, oure main focus is on colon cancer diagnosis based on HI analysis. In general, most of the stages of HI analysis depend mainly on the basic concepts of mathematics. Figure 72 presents the main stages of a typical HI Analysis pipeline [44][21].
2.3. Datasets
Dramatically increasing the dataset size needed for testing training is a critical challenge [54,55][24][25]. There are public datasets in the electronic pathology course, including manual observations for HIs. These are helpful in the review process. Image artifacts (e.g., the zoom level and image resolution) and slide problems (e.g., smudges) have similarity ratios. However, all of these datasets are expected only in specific states of tumors, and there are several tasks that the existing databases do not handle.-
CRC Grading Dataset
-
PanNuke Dataset
-
The Warwick-QU Dataset
-
CoNSeP Dataset
-
ETIS-LARIB
-
CRCHistoPhenotypes–Labeled Cell Nuclei Dataset
-
Kent Integrated Dataset (KID)
-
CVC-ColonDB and CVC-ClinicDB
-
Colonoscopy Dataset
-
ASU-Mayo ClinicCurrently, there are numerous research programs based on co-funded acceleration, seed research, and team science grants [65][35]. This means that more than 20–30 cohorts of senior nursing students in their clinical training by Mayo Clinic nursing faculty on the Mayo campus are expected to be completed. Due to this effort and cooperation, the seed grant program has added joint, cutting-edge research collaborations, a host of dual degree opportunities, and others. In 2016 and in the summer of 2010, the relationships of the Mayo Clinic became enterprise-wide, and the ASU Alliance for Health Care was formed.

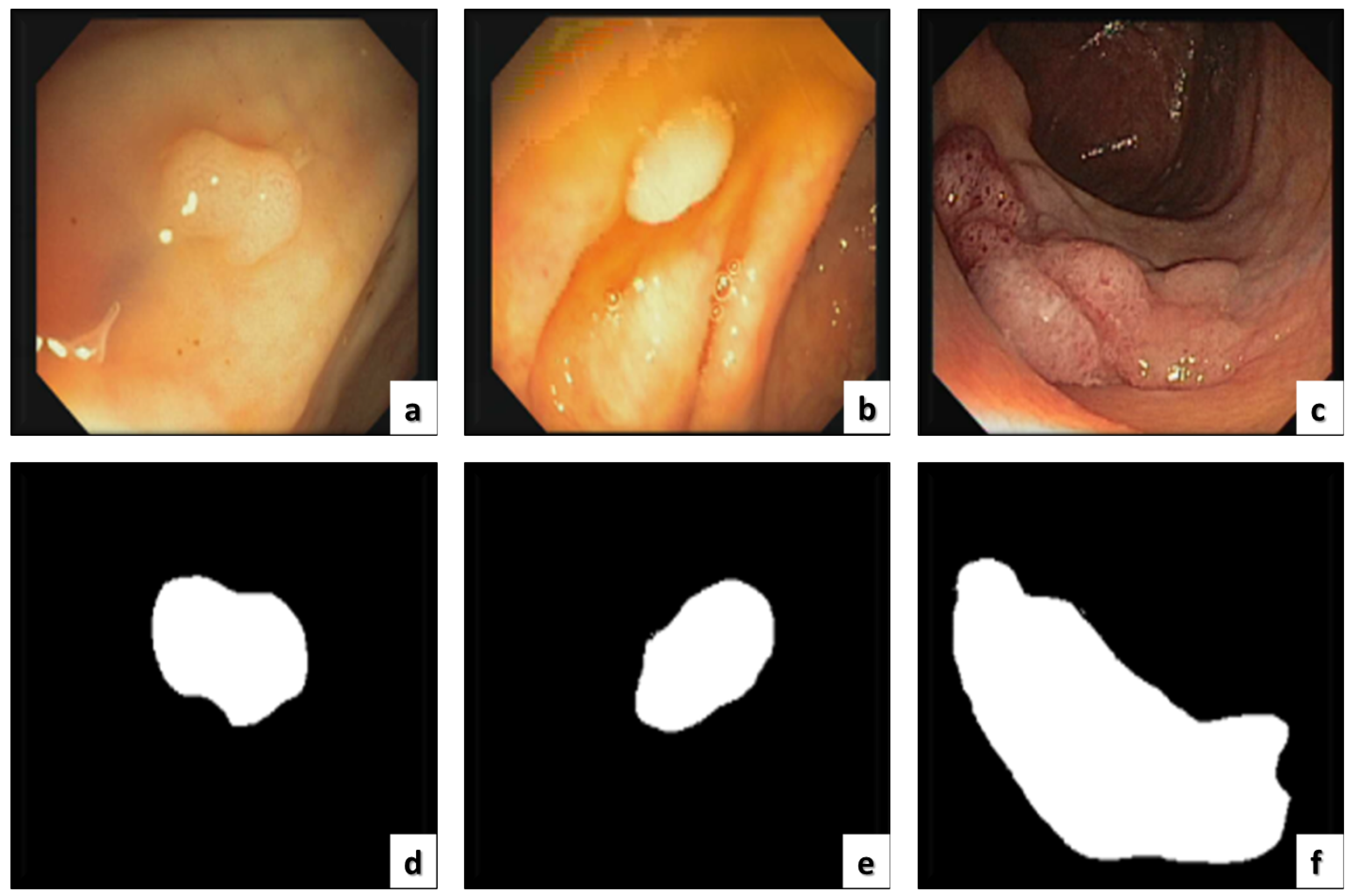
3.4. Performance Evaluation Metrics
Metrics of evaluation are utilized to measure the quality of models of machine learning. One can evaluate whether the DL algorithm of training is effective on new data by using these metrics of evaluation. Many different evaluation metrics can be used for testing a model. More accurate results can be found using multiple metrics for evaluating the quality of a trained model because each model performing using a metric of evaluation differs from the same model using another evaluation metric. The factors of correctly used evaluation metrics are critical as these describe whether the trained model is performing well or not. In the following section, rResearchers show some formulas and an explanation of the evaluation metrics utilized by academic papers. True Positive (TP) is when a method classifies the correct category correctly, while False Positive (FP) is when a method classifies the correct category incorrectly. On the other hand, True Negative (TN) is when a method classifies the negative category correctly, while False Negative (FN) is when a method classifies the negative category incorrectly. researchers can customize these values in the medical field of cancer detection. An example is that, if the image includes cancerous cells, then the trained model predicts the malignant cells successfully, and thus this case is called TP, while if the trained model predicts that it is not a malignant cell, then this case is called FP. On the other hand, if the image includes no malignant cells, and the model predicts that the image does not contain cancerous cells, then this case is called TN. If the image includes no malignant cells, and the trained model predicts it as a malignant cell, then this case is called FN. In the next section, researchers present an explanation and description for formulas that are related to the common evaluation metrics.-
The Accuracy measures the proportion of true observations to the number of samples measured, which can be calculated as:
-
The Rate of Error shows the proportion of inaccurate observations to the number of measured samples, which can be calculated as:
-
The Precision measures the true classified positive estimates of the total classified estimates in a correct category, which can be calculated as:
-
The Recall is employed for measuring the ratio of correct estimates that are correctly predicted. This can be calculated as:
-
The Specificity is presented for measuring the positive observations rate of false samples and can be calculated as:
-
The Sensitivity measures the number of correct samples that are classified as true and can be calculated as:
-
The ROC curve presents the ratio of false positives to the ratio of TPs by showing the performances of the possible threshold values used and can be calculated as:
References
- Allison, J.E. Colorectal cancer screening guidelines: The importance of evidence and transparency. Gastroenterology 2010, 138, 1648–1652.
- An, F.P.; Liu, J.E. Medical Image Segmentation Algorithm Based on Optimized Convolutional Neural Network-Adaptive Dropout Depth Calculation. Complexity 2020, 2020, 1645479.
- Araghi, M.; Soerjomataram, I.; Jenkins, M.; Brierley, J.; Morris, E.; Bray, F.; Arnold, M. Global trends in colorectal cancer mortality: Projections to the year 2035. Int. J. Cancer 2019, 144, 2992–3000.
- Barish, M.A.; Soto, J.A.; Ferrucci, J.T. Consensus on current clinical practice of virtual colonoscopy. Am. J. Roentgenol. 2005, 184, 786–792.
- Thun, M.J.; Calle, E.E.; Namboodiri, M.M.; Flanders, W.D.; Coates, R.J.; Byers, T.; Boffetta, P.; Garfinkel, L.; Heath, C.W., Jr. Risk factors for fatal colon cancer in a large prospective study. JNCI J. Natl. Cancer Inst. 1992, 84, 1491–1500.
- Rathore, S.; Hussain, M.; Ali, A.; Khan, A. A recent survey on colon cancer detection techniques. IEEE/ACM Trans. Comput. Biol. Bioinform. 2013, 10, 545–563.
- Baxter, N.N.; Goldwasser, M.A.; Paszat, L.F.; Saskin, R.; Urbach, D.R.; Rabeneck, L. Association of colonoscopy and death from colorectal cancer. Ann. Intern. Med. 2009, 150, 1–8.
- Bera, K.; Schalper, K.A.; Rimm, D.L.; Velcheti, V.; Madabhushi, A. Artificial intelligence in digital pathology—New tools for diagnosis and precision oncology. Nat. Rev. Clin. Oncol. 2019, 16, 703–715.
- Kitayama, J.; Nagawa, H.; Tsuno, N.; Osada, T.; Hatano, K.; Sunami, E.; Saito, H.; Muto, T. Laminin mediates tethering and spreading of colon cancer cells in physiological shear flow. Br. J. Cancer 1999, 80, 1927–1934.
- Burdan, F.; Sudol-Szopinska, I.; Staroslawska, E.; Kolodziejczak, M.; Klepacz, R.; Mocarska, A.; Caban, M.; Zelazowska-Cieslinska, I.; Szumilo, J. Magnetic resonance imaging and endorectal ultrasound for diagnosis of rectal lesions. Eur. J. Med. Res. 2015, 20, 1–14.
- Bychkov, D.; Linder, N.; Turkki, R.; Nordling, S.; Kovanen, P.E.; Verrill, C.; Walliander, M.; Lundin, M.; Haglund, C.; Lundin, J. Deep learning based tissue analysis predicts outcome in colorectal cancer. Sci. Rep. 2018, 8, 3395.
- Levin, B.; Lieberman, D.A.; McFarland, B.; Andrews, K.S.; Brooks, D.; Bond, J.; Dash, C.; Giardiello, F.M.; Glick, S.; Johnson, D.; et al. Screening and surveillance for the early detection of colorectal cancer and adenomatous polyps, 2008: A joint guideline from the American Cancer Society, the US Multi-Society Task Force on Colorectal Cancer, and the American College of Radiology. Gastroenterology 2008, 134, 1570–1595.
- Chaddad, A.; Tanougast, C.; Dandache, A.; Al Houseini, A.; Bouridane, A. Improving of colon cancer cells detection based on Haralick’s features on segmented histopathological images. In Proceedings of the 2011 IEEE International Conference on Computer Applications and Industrial Electronics (ICCAIE), Penang, Malaysia, 21–22 May 2011; pp. 87–90.
- Hur, C.; Chung, D.C.; Schoen, R.E.; Gazelle, G.S. The management of small polyps found by virtual colonoscopy: Results of a decision analysis. Clin. Gastroenterol. Hepatol. 2007, 5, 237–244.
- Gunduz-Demir, C.; Kandemir, M.; Tosun, A.B.; Sokmensuer, C. Automatic segmentation of colon glands using object-graphs. Med. Image Anal. 2010, 14, 1–12.
- Wang, D.; Foran, D.J.; Ren, J.; Zhong, H.; Kim, I.Y.; Qi, X. Exploring automatic prostate histopathology image gleason grading via local structure modeling. In Proceedings of the 2015 37th Annual International Conference of the IEEE Engineering in Medicine and Biology Society (EMBC), Milano, Italy, 25–29 August 2015; pp. 2649–2652.
- Dal Molin, M.; Matthaei, H.; Wu, J.; Blackford, A.; Debeljak, M.; Rezaee, N.; Wolfgang, C.L.; Butturini, G.; Salvia, R.; Bassi, C.; et al. Clinicopathological correlates of activating GNAS mutations in intraductal papillary mucinous neoplasm (IPMN) of the pancreas. Ann. Surg. Oncol. 2013, 20, 3802–3808.
- Masud, M.; Sikder, N.; Nahid, A.A.; Bairagi, A.K.; AlZain, M.A. A machine learning approach to diagnosing lung and colon cancer using a deep learning-based classification framework. Sensors 2021, 21, 748.
- Elazab, N.; Soliman, H.; El-Sappagh, S.; Islam, S.; Elmogy, M. Objective Diagnosis for Histopathological Images Based on Machine Learning Techniques: Classical Approaches and New Trends. Mathematics 2020, 8, 1863.
- Bar-Shalom, R.; Valdivia, A.Y.; Blaufox, M.D. PET imaging in oncology. Semin. Nucl. Med. 2000, 30, 150–185.
- Moroz, M.A.; Kochetkov, T.; Cai, S.; Wu, J.; Shamis, M.; Nair, J.; De Stanchina, E.; Serganova, I.; Schwartz, G.K.; Banerjee, D.; et al. Imaging colon cancer response following treatment with AZD1152: A preclinical analysis of fluoro-2-deoxyglucose and fluorothymidine imaging. Clin. Cancer Res. 2011, 17, 1099–1110.
- Kalkan, H.; Nap, M.; Duin, R.P.; Loog, M. Automated classification of local patches in colon histopathology. In Proceedings of the Proceedings of the 21st International Conference on Pattern Recognition (ICPR2012); Tsukuba, Japan, 11–15 November 2012, pp. 61–64.
- Greenspan, H.; Van Ginneken, B.; Summers, R.M. Guest editorial deep learning in medical imaging: Overview and future promise of an exciting new technique. IEEE Trans. Med. Imaging 2016, 35, 1153–1159.
- Burt, R.W. Strategies for colon cancer screening with considerations of cost and access to care. J. Natl. Compr. Cancer Netw. 2010, 8, 2–5.
- Fakoor, R.; Ladhak, F.; Nazi, A.; Huber, M. Using deep learning to enhance cancer diagnosis and classification. In Proceedings of the International Conference on Machine Learning, New York, NY, USA, 15–20 July 2013; Volume 28.
- Awan, R.; Sirinukunwattana, K.; Epstein, D.; Jefferyes, S.; Qidwai, U.; Aftab, Z.; Mujeeb, I.; Snead, D.; Rajpoot, N. Glandular morphometrics for objective grading of colorectal adenocarcinoma histology images. Sci. Rep. 2017, 7, 1–12.
- Gamper, J.; Koohbanani, N.A.; Benes, K.; Graham, S.; Jahanifar, M.; Khurram, S.A.; Azam, A.; Hewitt, K.; Rajpoot, N. Pannuke dataset extension, insights and baselines. arXiv 2020, arXiv:2003.10778.
- Sirinukunwattana, K.; Snead, D.R.; Rajpoot, N.M. A stochastic polygons model for glandular structures in colon histology images. IEEE Trans. Med. Imaging 2015, 34, 2366–2378.
- Graham, S.; Vu, Q.D.; Raza, S.E.A.; Azam, A.; Tsang, Y.W.; Kwak, J.T.; Rajpoot, N. Hover-net: Simultaneous segmentation and classification of nuclei in multi-tissue histology images. Med. Image Anal. 2019, 58, 101563.
- Jha, D.; Smedsrud, P.H.; Riegler, M.A.; Halvorsen, P.; de Lange, T.; Johansen, D.; Johansen, H.D. Kvasir-seg: A segmented polyp dataset. In Proceedings of the International Conference on Multimedia Modeling, Daejeon, Republic of Korea, 5–8 January 2020; pp. 451–462.
- Sirinukunwattana, K.; Raza, S.E.A.; Tsang, Y.W.; Snead, D.R.; Cree, I.A.; Rajpoot, N.M. Locality sensitive deep learning for detection and classification of nuclei in routine colon cancer histology images. IEEE Trans. Med. Imaging 2016, 35, 1196–1206.
- Lewer, D.; Bourne, T.; George, A.; Abi-Aad, G.; Taylor, C.; George, J. Data Resource: The Kent Integrated Dataset (KID). Int. J. Popul. Data Sci. 2018, 3, 427.
- Mesejo, P.; Pizarro, D.; Abergel, A.; Rouquette, O.; Beorchia, S.; Poincloux, L.; Bartoli, A. Computer-aided classification of gastrointestinal lesions in regular colonoscopy. IEEE Trans. Med. Imaging 2016, 35, 2051–2063.
- Shaban, M.; Awan, R.; Fraz, M.M.; Azam, A.; Tsang, Y.W.; Snead, D.; Rajpoot, N.M. Context-aware convolutional neural network for grading of colorectal cancer histology images. IEEE Trans. Med. Imaging 2020, 39, 2395–2405.
- Pogorelov, K.; Randel, K.R.; Griwodz, C.; Eskeland, S.L.; de Lange, T.; Johansen, D.; Spampinato, C.; Dang-Nguyen, D.T.; Lux, M.; Schmidt, P.T.; et al. Kvasir: A multi-class image dataset for computer aided gastrointestinal disease detection. In Proceedings of the eighth ACM on Multimedia Systems Conference, Taipei, Taiwan, 20–23 June 2017; pp. 164–169.
