Pancreatic ductal adenocarcinoma (PDAC) has an extremely poor prognosis due to the lack of methods or biomarkers for early diagnosis and its resistance to conventional treatment modalities, targeted therapies, and immunotherapies. PDACs are a heterogenous group of malignant epithelial neoplasms with various histomorphological patterns and complex, heterogenous genetic/molecular landscapes. The newly proposed molecular classifications of PDAC based on extensive genomic, transcriptomic, proteomic and epigenetic data have provided significant insights into the molecular heterogeneity and aggressive biology of this deadly disease. Recent sStudies characterizing the tumor microenvironment (TME) have shed light on the dynamic interplays between the tumor cells and the immunosuppressive TME of PDAC, which is essential to disease progression, as well as its resistance to chemotherapy, newly developed targeted therapy and immunotherapy. There is a critical need for the development of predictive markers that can be clinically utilized to select effective personalized therapies for PDAC patients.
- pancreatic ductal adenocarcinoma
- molecular pathology
- predictive marker
- tumor microenvironment
- targeted therapy and immunotherapy
1. Introduction
2. Histology and Morphologic Heterogeneity of PDAC
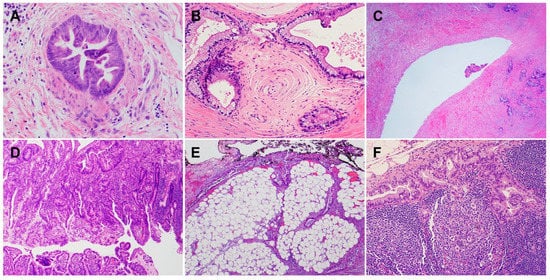
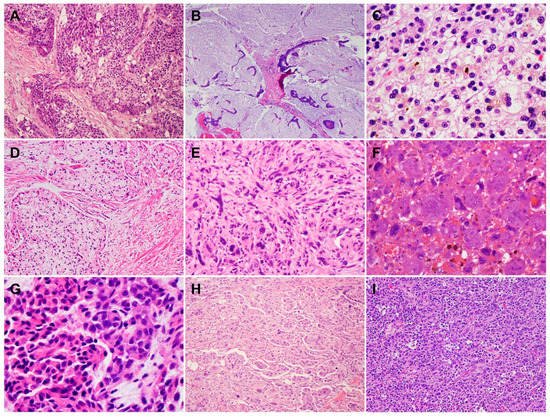
PDACs are a heterogeneous group of malignant pancreatic epithelial neoplasms. Conventional PDACs are characterized by dense desmoplastic stroma intermixed with angulated glands, small nests of malignant epithelial cells and/or single tumor cells (Figure 1A). PDACs often show a spectrum of differentiation ranging from well-differentiated to poorly differentiated adenocarcinomas within the same tumor and show significant inter- and intra-tumoral heterogeneity in histomorphological patterns, such as complex interconnecting tumoral glands embedded in desmoplastic stroma (Figure 1B), a large duct type (Figure 1C), poorly differentiated carcinoma with eosinophilic or clear cells (Figure 1D–F), complex intraluminal micropapillae formation (Figure 1G), cribriform histology with foamy cells (Figure 1H), and the pagetoid involvement of pancreatic duct/intraductal carcinoma (Figure 1I). Aggressive histological features, such as lymphovascular invasion, tumor invasion into peripancreatic soft tissue, large vessels and adjacent organ(s)/structure(s), perineural invasion, the involvement of resection margins, and lymph node metastasis, are frequently present in resected PDAC specimens (Figure 2A–F). The presence of these aggressive histological features is associated with an increased risk of post-operative tumor recurrence/distant metastasis and poor survival in PDAC patients [9,10,11,12][9][10][11][12].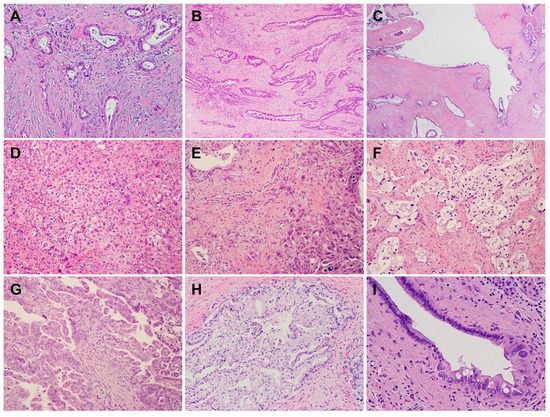
3. Genetic Alterations and Molecular Subtypes of PDAC
PDAC is characterized by a handful of inherited (germline) and recurring somatic mutations. The first whole exome sequencing of human PDAC samples was reported in 2008 by Jones et al. [24]. In that study, 20,661 protein-coding genes in 24 PDAC samples were sequenced, and more than 1500 somatic mutations in 1007 genes were identified. Thise study was followed by several landmark, large-scale whole exome sequencing and comprehensive molecular profiling of human PDAC samples, which provided uresearchers with the in-depth understanding of the heterogeneous molecular landscapes of PDAC [25,26,27][25][26][27]. These studies identified four “mountains” (the genes mutated at the greatest frequency): oncogenic mutations of Kirsten rat sarcoma (KRAS), loss-of-function mutations and/or deletions of the TP53 tumor suppressor genes, mothers against decapentaplegic homolog 4 (SMAD4), and the cyclin dependent kinase inhibitor 2A (CDKN2A) [24]. The data from genetically engineered mouse models have shown that these mutations play an essential role in the development and/or progression of PDAC [24,28,29][24][28][29]. In addition to the four “mountains”, a large number of less common “hills” (genes mutated at low frequencies) have been detected in PDACs. For example, amplifications of other less frequent oncogenes such as CMYC (on chromosome 8q), MYB (chromosome 6q), AIB1/NCOA3 (chromosome 20q), EGFR (chromosome 7p), and GATA6, as well as recurrent chromosomal amplifications, have also been identified [7]. One or more somatic/germline mutations of the genes involved in DNA damage repair (DDR), such as BRCA2, BRCA1, PALB2, ATM, CHEK2, RAD51C, and RAD51D mutations, may be detected in 10–20% of PDAC patients [30,31,32,33][30][31][32][33]. These less common genetic alterations may represent valuable targets or serve as predictive biomarkers for PDAC patients. For example, defects in DDR pathway in PDACs represent a unique subset of patients who may benefit from platinum-based chemotherapy (e.g., cisplatin) or the newly approved poly (ADP-ribose) polymerase (PARP) inhibitors such as olaparib [34,35][34][35]. Multiple studies have reported on the molecular subtypes of PDAC based on the whole exome sequencing data and/or integrated analyses of the genomic, transcriptomic, proteomic, and epigenetic profiles of human PDAC samples [36,37,38,39][36][37][38][39]. Collisson et al. reported three molecular subtypes of PDAC: the classical, quasi-mesenchymal (QM), and exocrine-like based on the analysis of 27 microdissected PDAC samples [38]. The classical subtype showed a higher expression of adhesion-associated and epithelial genes and a higher expression of KRAS and GATA6, an essential gene for pancreatic development and PDAC progression, compared with the QM subtype. The QM subtype was found to have a high expression of mesenchyme-associated genes. The exocrine-like subtype showed the relatively high expression of tumor cell-derived digestive enzyme genes. These molecular subtypes were found to be significantly correlated with patient survival in that the QM subtype had the worst survival. They also demonstrated that PDAC cell lines of the QM subtype were more sensitive to gemcitabine, whereas the classical subtype cell lines were more sensitive to erlotinib (an EGFR antagonist) [38]. Using non-negative matrix factorization to digitally dissect the tumor and stromal gene signature of primary and metastatic PDAC, Moffitt et al. identified two tumor-specific subtypes: the basal-like subtype, which is molecularly similar to the basal-like carcinoma of breast and bladder, and the classical subtype, and two stromal subtypes: normal and activated. The basal-like subtype had a worse prognosis but a superior response to adjuvant therapy compared with the classical subtype [39]. Via RNA sequencing, they showed that the KRASG12D mutation was significantly overrepresented in the basal-like subtype, KRASG12V was isolated to the classical subtype, and SMAD4 expression was significantly higher in the classical subtype compared with the basal-like subtype, which is consistent with the observation that SMAD4 loss confers a more aggressive tumor behavior [39]. The normal stroma showed the relatively high expression of known markers for pancreatic stellate cells (desmin, smooth muscle actin, and vimentin), whereas the activated stroma was characterized by a gene set associated with macrophages (integrin ITGAM and the chemokine ligands CCL13 and CCL18) and other genes that are reported to promote tumor progression (SPARC, WNT2, WNT5A, MMP9, and MMP11) [39,40][39][40]. Patients with the classical subtype and activated stroma had worse survival compared with those with the classical subtype and normal stroma. Stromal subtypes were not associated with survival in patients with basal-like subtype PDAC [39]. Moffitt’s classification of PDACs into basal-like and classical subtypes was validated by Puleo et al., who classified PDACs into five subtypes: pure basal-like, stroma-activated, desmoplastic, pure classical, and immune classical based on features of cancer cells and the tumor microenvironment [41]. Thus, targeting the tumor-promoting genes in activated stroma may represent a potential strategy for PDAC patients. The integrated genomic analysis of 456 PDAC samples by Bailey et al. identified 32 recurrent mutated genes in 10 pathways—KRAS, TGF-β, WNT, NOTCH, ROBO/SLIT signaling, G1/S transition, SWI-SNF, chromatin modification, DNA repair, and RNA processing—and four molecular subtypes of PDACs—pancreatic progenitor, squamous, aberrantly differentiated endocrine exocrine (ADEX), and immunogenic [36]. The pancreatic progenitor subtype (19%) was characterized by the transcriptional factors involved in early pancreatic development (PDX1, MNX1, HNF1A, HNF1B, HNF4A, HNF4G, FOXA2, FOXA3, and HES1) and metabolic pathways such as fatty acid oxidation and drug metabolism [42]. The squamous subtype (31%) was enriched with TP53 and KDM6A mutations, as well as the hypermethylation of genes governing pancreatic endodermal cell-fate determination (e.g., PDX1, MNX1, GATA6, and HNF1B). The ADEX subtype (21%) showed the upregulation of genes that regulate networks involved in KRAS activation. This subtype included two gene programs, with one focused on exocrine function (NR5A2, MIST1 and RBPJL) and the other related to endocrine differentiation (NEUROD1, MODY, INS and NKX2–2). The immunogenic subtype (29%) contained a family of genes related to immune cell function including B cell signaling, Toll-like receptor signaling, antigen presentation, and the infiltration of CD8+ and CD4+ T cells, with the upregulation of the immune inhibitor PD-1 and cytotoxic T-lymphocyte-associated protein-4 (CTLA-4) [36,42][36][42]. These molecular subtypes correlated with histopathological features in that the squamous subtype represented ASC, progenitor and immunogenic represented colloid carcinomas and carcinomas arising from IPMN, and ADEX represented rare acinar cell carcinomas [36]. Among these molecular subtypes, the squamous subtype had the worst survival, while the other three subtypes showed similar survival rates [36]. Another integrated analysis of the mRNA, miRNA, lncRNA and DNA methylation profiling of 150 PDAC samples by the Cancer Genome Atlas Research Network identified two subtypes of PDACs: SNF-1 and SNF-2. The SNF-1 subtype represented most of the basal-like subtype in Moffitt’s classification, the squamous subtype in Bailey’s classification, and the QM subtype in Collison’s classification [37]. More recently, Chan-Seng-Yue et al. performed whole genome and transcriptome analysis of purified tumor cells from 314 primary and metastatic PDAC patients to generate tumor-specific expression signatures [43]. They classified PDACs into five molecular subtypes: basal-like A and B for the previously defined basal-like subtype, hybrid, and classical A and B for the previously defined previously defined classical subtype. The hybrid subtype was inconsistently classified by previous classification systems due to the presence of multiple expression signatures. Patients with basal-like A PDAC often present with advanced disease and show the worst response to gemcitabine-based chemotherapies and FOLFIRINOX. In contrast, patients with basal-like B and hybrid tumors often present with resectable disease. Therefore, the ability to distinguish the basal-like A, basal-like B, and hybrid subtypes from the group formerly classified as basal-like allows for the more accurate prediction of chemotherapy response. Classical A/B tumors were found to be associated with an increased frequency of GATA6 amplification and complete SMAD4 loss, whereas basal-like A/B tumors showed the complete loss of CDKN2A and a higher frequency of TP53 mutations. At single-cell resolution, the authors also showed that basal-like and classical subtypes can co-exist in the same tumor, which highlighted the intra-tumoral molecular heterogeneity [43]. The most recent molecular subclassification of PDACs was reported by Hwang et al. in 2022. Using the single-nucleus RNA sequencing and whole digital spatial transcriptome profiling of 43 primary PDAC samples (18 untreated and 25 treated), they identified three distinct subtypes: classical, squamoid–basaloid, and treatment-enriched. Their study uncovered that the neural-like progenitor (NRP) malignant cell program was enriched in residual carcinoma after chemoradiation therapy. The NRP cells were associated with treatment resistance and poor survival in PDAC patients via the regulation of genes involved in drug efflux, the negative regulation of cell death, and resistance to chemotherapy (e.g., ABCB1, BCL2, PDGFD and SPP1), tumor–nerve crosstalk (e.g., SEMA3E, RELN and SEMA5A), and metastasis (NFIB) [44]. These molecular classifications of PDAC provide rich and comprehensive datasets to better understand pancreatic tumorigenesis, genetic/molecular landscapes, intra- and inter-tumoral heterogeneity, tumor progression, and drug resistance. More importantly, the molecular subtyping of PDAC may provide useful information for more effective subtype-tailored therapies for PDAC patients. However, due to their complexity, these classifications of PDAC have not been utilized in daily pathologic diagnosis or clinical practice.4. Heterogeneous Response of PDAC to Neoadjuvant Therapy
Neoadjuvant therapy is routinely used to treat PDAC patients with borderline resectable and high-risk resectable disease, as well as selected patients with locally advanced disease [45]. Pathologic studies of the post-therapy pancreatectomy specimens have shown that only 12.6% to 18.6% PDAC patients demonstrate a complete or near complete pathologic response to neoadjuvant therapy, which is associated with better survival, while the majority of PDAC patients (>80%) demonstrate a moderate or minimal response to neoadjuvant therapy and poor survival [46,47,48,49][46][47][48][49]. These data highlight not only the fact that vast majority of PDACs are resistant to neoadjuvant chemotherapy with or without radiation but also the inter-tumoral heterogeneity in tumor response among PDAC patients. It is also not uncommon to observe significant intra-tumoral, heterogeneous response to neoadjuvant therapy in different areas of the same treated tumor, with some areas of the tumor showing complete or near complete response and other areas showing minimal or no response (Figure 4A,B). Occasionally, differential responses to neoadjuvant therapy are also observed between the primary PDAC and the metastatic carcinoma in lymph node(s) in the same patient (Figure 4C,D). Currently, there are limited data on the molecular correlation with tumor response to neoadjuvant therapies. The molecular mechanisms underlying the inter- and intra-tumoral heterogeneity in response to neoadjuvant therapy is not clear. It is possible that the cellular and molecular/genetic heterogeneity and the heterogeneity in the TME contribute to the heterogeneous response in PDAC patients. The recent molecular profiling of treated PDACs suggested that the NRP malignant program was enriched in residual carcinoma after chemoradiation therapy and plays a role in tumor resistance to therapy [44]. Future biomarker-driven clinical trials of neoadjuvant therapy are needed to address this important question.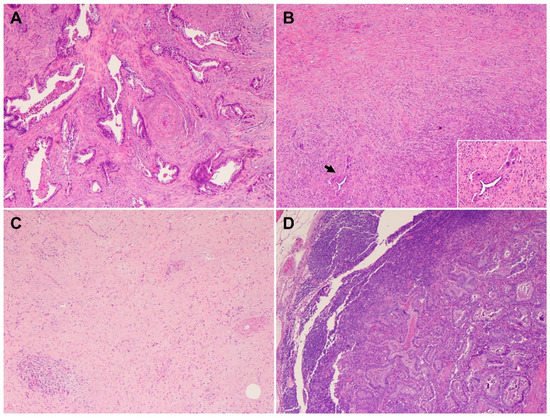
References
- Sung, H.; Ferlay, J.; Siegel, R.L.; Laversanne, M.; Soerjomataram, I.; Jemal, A.; Bray, F. Global Cancer Statistics 2020: GLOBOCAN Estimates of Incidence and Mortality Worldwide for 36 Cancers in 185 Countries. CA Cancer J. Clin. 2021, 71, 209–249.
- Rahib, L.; Wehner, M.R.; Matrisian, L.M.; Nead, K.T. Estimated Projection of US Cancer Incidence and Death to 2040. JAMA Netw. Open 2021, 4, e214708.
- National Cancer Institute SRP, Cancer Statistics Branch. Surveillance Epidemiology and End Results (SEER) (1975–2018). 2021. Available online: http://seer.cancer.gov (accessed on 7 August 2022).
- Kamisawa, T.; Wood, L.D.; Itoi, T.; Takaori, K. Pancreatic cancer. Lancet 2016, 388, 73–85.
- Maitra, A.; Hruban, R.H. Pancreatic cancer. Annu. Rev. Pathol. 2008, 3, 157–188.
- Garcea, G.; Dennison, A.R.; Pattenden, C.J.; Neal, C.P.; Sutton, C.D.; Berry, D.P. Survival following curative resection for pancreatic ductal adenocarcinoma. A systematic review of the literature. JOP 2008, 9, 99–132.
- Hong, S.M.; Park, J.Y.; Hruban, R.H.; Goggins, M. Molecular signatures of pancreatic cancer. Arch. Pathol. Lab. Med. 2011, 135, 716–727.
- Campbell, P.J.; Yachida, S.; Mudie, L.J.; Stephens, P.J.; Pleasance, E.D.; Stebbings, L.A.; Morsberger, L.A.; Latimer, C.; McLaren, S.; Lin, M.L.; et al. The patterns and dynamics of genomic instability in metastatic pancreatic cancer. Nature 2010, 467, 1109–1113.
- Chatterjee, D.; Katz, M.H.; Rashid, A.; Wang, H.; Iuga, A.C.; Varadhachary, G.R.; Wolff, R.A.; Lee, J.E.; Pisters, P.W.; Crane, C.H.; et al. Perineural and intraneural invasion in posttherapy pancreaticoduodenectomy specimens predicts poor prognosis in patients with pancreatic ductal adenocarcinoma. Am. J. Surg. Pathol. 2012, 36, 409–417.
- Chatterjee, D.; Rashid, A.; Wang, H.; Katz, M.H.; Wolff, R.A.; Varadhachary, G.R.; Lee, J.E.; Pisters, P.W.; Gomez, H.F.; Abbruzzese, J.L.; et al. Tumor invasion of muscular vessels predicts poor prognosis in patients with pancreatic ductal adenocarcinoma who have received neoadjuvant therapy and pancreaticoduodenectomy. Am. J. Surg. Pathol. 2012, 36, 552–559.
- Fischer, L.K.; Katz, M.H.; Lee, S.M.; Liu, L.; Wang, H.; Varadhachary, G.R.; Wolff, R.A.; Lee, J.E.; Maitra, A.; Roland, C.L.; et al. The number and ratio of positive lymph nodes affect pancreatic cancer patient survival after neoadjuvant therapy and pancreaticoduodenectomy. Histopathology 2016, 68, 210–220.
- Liu, L.; Katz, M.H.; Lee, S.M.; Fischer, L.K.; Prakash, L.; Parker, N.; Wang, H.; Varadhachary, G.R.; Wolff, R.A.; Lee, J.E.; et al. Superior Mesenteric Artery Margin of Posttherapy Pancreaticoduodenectomy and Prognosis in Patients with Pancreatic Ductal Adenocarcinoma. Am. J. Surg. Pathol. 2015, 39, 1395–1403.
- International Agency for Research on Cancer. WHO 2019 Classification of Tumors, Digestive System Tumours, 5th ed.; International Agency for Research on Cancer: Lyon, France, 2019; Volume 1, pp. 295–332.
- Adsay, N.V.; Merati, K.; Nassar, H.; Shia, J.; Sarkar, F.; Pierson, C.R.; Cheng, J.D.; Visscher, D.W.; Hruban, R.H.; Klimstra, D.S. Pathogenesis of colloid (pure mucinous) carcinoma of exocrine organs: Coupling of gel-forming mucin (MUC2) production with altered cell polarity and abnormal cell-stroma interaction may be the key factor in the morphogenesis and indolent behavior of colloid carcinoma in the breast and pancreas. Am. J. Surg. Pathol. 2003, 27, 571–578.
- Adsay, N.V.; Pierson, C.; Sarkar, F.; Abrams, J.; Weaver, D.; Conlon, K.C.; Brennan, M.F.; Klimstra, D.S. Colloid (mucinous noncystic) carcinoma of the pancreas. Am. J. Surg. Pathol. 2001, 25, 26–42.
- Poultsides, G.A.; Reddy, S.; Cameron, J.L.; Hruban, R.H.; Pawlik, T.M.; Ahuja, N.; Jain, A.; Edil, B.H.; Iacobuzio-Donahue, C.A.; Schulick, R.D.; et al. Histopathologic basis for the favorable survival after resection of intraductal papillary mucinous neoplasm-associated invasive adenocarcinoma of the pancreas. Ann. Surg. 2010, 251, 470–476.
- Seidel, G.; Zahurak, M.; Iacobuzio-Donahue, C.; Sohn, T.A.; Adsay, N.V.; Yeo, C.J.; Lillemoe, K.D.; Cameron, J.L.; Hruban, R.H.; Wilentz, R.E. Almost all infiltrating colloid carcinomas of the pancreas and periampullary region arise from in situ papillary neoplasms: A study of 39 cases. Am. J. Surg. Pathol. 2002, 26, 56–63.
- Brody, J.R.; Costantino, C.L.; Potoczek, M.; Cozzitorto, J.; McCue, P.; Yeo, C.J.; Hruban, R.H.; Witkiewicz, A.K. Adenosquamous carcinoma of the pancreas harbors KRAS2, DPC4 and TP53 molecular alterations similar to pancreatic ductal adenocarcinoma. Mod. Pathol. 2009, 22, 651–659.
- Le, D.T.; Uram, J.N.; Wang, H.; Bartlett, B.R.; Kemberling, H.; Eyring, A.D.; Skora, A.D.; Luber, B.S.; Azad, N.S.; Laheru, D.; et al. PD-1 Blockade in Tumors with Mismatch-Repair Deficiency. N. Engl. J. Med. 2015, 372, 2509–2520.
- Wilentz, R.E.; Goggins, M.; Redston, M.; Marcus, V.A.; Adsay, N.V.; Sohn, T.A.; Kadkol, S.S.; Yeo, C.J.; Choti, M.; Zahurak, M.; et al. Genetic, immunohistochemical, and clinical features of medullary carcinoma of the pancreas: A newly described and characterized entity. Am. J. Pathol. 2000, 156, 1641–1651.
- Calhoun, E.S.; Jones, J.B.; Ashfaq, R.; Adsay, V.; Baker, S.J.; Valentine, V.; Hempen, P.M.; Hilgers, W.; Yeo, C.J.; Hruban, R.H.; et al. BRAF and FBXW7 (CDC4, FBW7, AGO, SEL10) mutations in distinct subsets of pancreatic cancer: Potential therapeutic targets. Am. J. Pathol. 2003, 163, 1255–1260.
- Goggins, M.; Offerhaus, G.J.; Hilgers, W.; Griffin, C.A.; Shekher, M.; Tang, D.; Sohn, T.A.; Yeo, C.J.; Kern, S.E.; Hruban, R.H. Pancreatic adenocarcinomas with DNA replication errors (RER+) are associated with wild-type K-ras and characteristic histopathology. Poor differentiation, a syncytial growth pattern, and pushing borders suggest RER+. Am. J. Pathol. 1998, 152, 1501–1507.
- Yamamoto, H.; Itoh, F.; Nakamura, H.; Fukushima, H.; Sasaki, S.; Perucho, M.; Imai, K. Genetic and clinical features of human pancreatic ductal adenocarcinomas with widespread microsatellite instability. Cancer Res. 2001, 61, 3139–3144.
- Jones, S.; Zhang, X.; Parsons, D.W.; Lin, J.C.; Leary, R.J.; Angenendt, P.; Mankoo, P.; Carter, H.; Kamiyama, H.; Jimeno, A.; et al. Core signaling pathways in human pancreatic cancers revealed by global genomic analyses. Science 2008, 321, 1801–1806.
- Mehlen, P.; Delloye-Bourgeois, C.; Chedotal, A. Novel roles for Slits and netrins: Axon guidance cues as anticancer targets? Nat. Rev. Cancer 2011, 11, 188–197.
- Sabatier, C.; Plump, A.S.; Le, M.; Brose, K.; Tamada, A.; Murakami, F.; Lee, E.Y.; Tessier-Lavigne, M. The divergent Robo family protein rig-1/Robo3 is a negative regulator of slit responsiveness required for midline crossing by commissural axons. Cell 2004, 117, 157–169.
- Trusolino, L.; Comoglio, P.M. Scatter-factor and semaphorin receptors: Cell signalling for invasive growth. Nat. Rev. Cancer 2002, 2, 289–300.
- Samuel, N.; Hudson, T.J. The molecular and cellular heterogeneity of pancreatic ductal adenocarcinoma. Nat. Rev. Gastroenterol. Hepatol. 2011, 9, 77–87.
- Biankin, A.V.; Waddell, N.; Kassahn, K.S.; Gingras, M.C.; Muthuswamy, L.B.; Johns, A.L.; Miller, D.K.; Wilson, P.J.; Patch, A.M.; Wu, J.; et al. Pancreatic cancer genomes reveal aberrations in axon guidance pathway genes. Nature 2012, 491, 399–405.
- Connor, A.A.; Denroche, R.E.; Jang, G.H.; Timms, L.; Kalimuthu, S.N.; Selander, I.; McPherson, T.; Wilson, G.W.; Chan-Seng-Yue, M.A.; Borozan, I.; et al. Association of Distinct Mutational Signatures with Correlates of Increased Immune Activity in Pancreatic Ductal Adenocarcinoma. JAMA Oncol. 2017, 3, 774–783.
- Lowery, M.A.; Wong, W.; Jordan, E.J.; Lee, J.W.; Kemel, Y.; Vijai, J.; Mandelker, D.; Zehir, A.; Capanu, M.; Salo-Mullen, E.; et al. Prospective Evaluation of Germline Alterations in Patients With Exocrine Pancreatic Neoplasms. J. Natl. Cancer Inst. 2018, 110, 1067–1074.
- Waddell, N.; Pajic, M.; Patch, A.M.; Chang, D.K.; Kassahn, K.S.; Bailey, P.; Johns, A.L.; Miller, D.; Nones, K.; Quek, K.; et al. Whole genomes redefine the mutational landscape of pancreatic cancer. Nature 2015, 518, 495–501.
- Perkhofer, L.; Golan, T.; Cuyle, P.J.; Matysiak-Budnik, T.; Van Laethem, J.L.; Macarulla, T.; Cauchin, E.; Kleger, A.; Beutel, A.K.; Gout, J.; et al. Targeting DNA Damage Repair Mechanisms in Pancreas Cancer. Cancers 2021, 13, 4259.
- Sahin, I.H.; Lowery, M.A.; Stadler, Z.K.; Salo-Mullen, E.; Iacobuzio-Donahue, C.A.; Kelsen, D.P.; O’Reilly, E.M. Genomic instability in pancreatic adenocarcinoma: A new step towards precision medicine and novel therapeutic approaches. Expert Rev. Gastroenterol. Hepatol. 2016, 10, 893–905.
- Singhi, A.D.; George, B.; Greenbowe, J.R.; Chung, J.; Suh, J.; Maitra, A.; Klempner, S.J.; Hendifar, A.; Milind, J.M.; Golan, T.; et al. Real-Time Targeted Genome Profile Analysis of Pancreatic Ductal Adenocarcinomas Identifies Genetic Alterations That Might Be Targeted With Existing Drugs or Used as Biomarkers. Gastroenterology 2019, 156, 2242–2253 e2244.
- Bailey, P.; Chang, D.K.; Nones, K.; Johns, A.L.; Patch, A.M.; Gingras, M.C.; Miller, D.K.; Christ, A.N.; Bruxner, T.J.; Quinn, M.C.; et al. Genomic analyses identify molecular subtypes of pancreatic cancer. Nature 2016, 531, 47–52.
- Cancer Genome Atlas Research Network. Electronic address, a.a.d.h.e.; Cancer Genome Atlas Research, N. Integrated Genomic Characterization of Pancreatic Ductal Adenocarcinoma. Cancer Cell 2017, 32, 185–203 e113.
- Collisson, E.A.; Bailey, P.; Chang, D.K.; Biankin, A.V. Molecular subtypes of pancreatic cancer. Nat. Rev. Gastroenterol. Hepatol. 2019, 16, 207–220.
- Moffitt, R.A.; Marayati, R.; Flate, E.L.; Volmar, K.E.; Loeza, S.G.; Hoadley, K.A.; Rashid, N.U.; Williams, L.A.; Eaton, S.C.; Chung, A.H.; et al. Virtual microdissection identifies distinct tumor- and stroma-specific subtypes of pancreatic ductal adenocarcinoma. Nat. Genet. 2015, 47, 1168–1178.
- Cohen, S.J.; Alpaugh, R.K.; Palazzo, I.; Meropol, N.J.; Rogatko, A.; Xu, Z.; Hoffman, J.P.; Weiner, L.M.; Cheng, J.D. Fibroblast activation protein and its relationship to clinical outcome in pancreatic adenocarcinoma. Pancreas 2008, 37, 154–158.
- Puleo, F.; Nicolle, R.; Blum, Y.; Cros, J.; Marisa, L.; Demetter, P.; Quertinmont, E.; Svrcek, M.; Elarouci, N.; Iovanna, J.; et al. Stratification of Pancreatic Ductal Adenocarcinomas Based on Tumor and Microenvironment Features. Gastroenterology 2018, 155, 1999–2013 e1993.
- Torres, C.; Grippo, P.J. Pancreatic cancer subtypes: A roadmap for precision medicine. Ann. Med. 2018, 50, 277–287.
- Chan-Seng-Yue, M.; Kim, J.C.; Wilson, G.W.; Ng, K.; Figueroa, E.F.; O’Kane, G.M.; Connor, A.A.; Denroche, R.E.; Grant, R.C.; McLeod, J.; et al. Transcription phenotypes of pancreatic cancer are driven by genomic events during tumor evolution. Nat. Genet. 2020, 52, 231–240.
- Hwang, W.L.; Jagadeesh, K.A.; Guo, J.A.; Hoffman, H.I.; Yadollahpour, P.; Reeves, J.W.; Mohan, R.; Drokhlyansky, E.; Van Wittenberghe, N.; Ashenberg, O.; et al. Single-nucleus and spatial transcriptome profiling of pancreatic cancer identifies multicellular dynamics associated with neoadjuvant treatment. Nat. Genet. 2022, 54, 1178–1191.
- Tempero, M.A.; Malafa, M.P.; Al-Hawary, M.; Behrman, S.W.; Berson III, A.B.; Cardin, D.B.; Cha, C.; Chiorean, E.G.; Chung, V.; Czito, B.; et al. NCCN Clinical Practice Guidelines in Oncology, Pancreatic Adenocarcinoma (Version 1.2021). Available online: https://www.nccn.org/professionals/physician_gls/PDF/pancreatic.pdf (accessed on 25 January 2021).
- Chatterjee, D.; Katz, M.H.; Rashid, A.; Varadhachary, G.R.; Wolff, R.A.; Wang, H.; Lee, J.E.; Pisters, P.W.; Vauthey, J.N.; Crane, C.; et al. Histologic grading of the extent of residual carcinoma following neoadjuvant chemoradiation in pancreatic ductal adenocarcinoma: A predictor for patient outcome. Cancer 2012, 118, 3182–3190.
- Chou, A.; Ahadi, M.; Arena, J.; Sioson, L.; Sheen, A.; Fuchs, T.L.; Pavlakis, N.; Clarke, S.; Kneebone, A.; Hruby, G.; et al. A Critical Assessment of Postneoadjuvant Therapy Pancreatic Cancer Regression Grading Schemes with a Proposal for a Novel Approach. Am. J. Surg. Pathol. 2021, 45, 394–404.
- Lee, S.M.; Katz, M.H.; Liu, L.; Sundar, M.; Wang, H.; Varadhachary, G.R.; Wolff, R.A.; Lee, J.E.; Maitra, A.; Fleming, J.B.; et al. Validation of a Proposed Tumor Regression Grading Scheme for Pancreatic Ductal Adenocarcinoma After Neoadjuvant Therapy as a Prognostic Indicator for Survival. Am. J. Surg. Pathol. 2016, 40, 1653–1660.
- Nagaria, T.S.; Wang, H.; Chatterjee, D.; Wang, H. Pathology of Treated Pancreatic Ductal Adenocarcinoma and Its Clinical Implications. Arch. Pathol. Lab. Med. 2020, 144, 838–845.
