Title: Exploitation of Heterosis in Pearl Millet: A Review
by Rakesh K. Srivastava *, Srikanth Bollam, Vijayalakshmi Pujarula, Madhu Pusuluri, Ram B. Singh, Gopi Potupureddi and Rajeev Gupta *
International Crops Research Institute for the Semi-Arid Tropics (ICRISAT), Hyderabad TS 502324, India
*
Plants 2020, 9(7), 807; https://doi.org/10.3390/plants9070807
Received: 6 March 2020 / Revised: 23 June 2020 / Accepted: 23 June 2020 / Published: 27 June 2020
Abstract
The phenomenon of heterosis has fascinated plant breeders ever since it was first described by Charles Darwin in 1876 in the vegetable kingdom and later elaborated by George H Shull and Edward M East in maize during 1908. Heterosis is the phenotypic and functional superiority manifested in the F1 crosses over the parents. Various classical complementation mechanisms gave way to the study of the underlying potential cellular and molecular mechanisms responsible for heterosis. In cereals, such as maize, heterosis has been exploited very well, with the development of many single-cross hybrids that revolutionized the yield and productivity enhancements. Pearl millet (Pennisetum glaucum (L.) R. Br.) is one of the important cereal crops with nutritious grains and lower water and energy footprints in addition to the capability of growing in some of the harshest and most marginal environments of the world. In this highly cross-pollinating crop, heterosis was exploited by the development of a commercially viable cytoplasmic male-sterility (CMS) system involving a three-lines breeding system (A-, B- and R-lines). The first set of male-sterile lines, i.e., Tift 23A and Tift18A, were developed in the early 1960s in Tifton, Georgia, USA. These provided a breakthrough in the development of hybrids worldwide, e.g., Tift 23A was extensively used by Punjab Agricultural University (PAU), Ludhiana, India, for the development of the first single-cross pearl millet hybrid, named Hybrid Bajra 1 (HB 1), in 1965. Over the past five decades, the pearl millet community has shown tremendous improvement in terms of cytoplasmic and nuclear diversification of the hybrid parental lines, which led to a progressive increase in the yield and adaptability of the hybrids that were developed, resulting in significant genetic gains. Lately, the whole genome sequencing of Tift 23D2B1 and re-sequencing of circa 1000 genomes by a consortium led by the International Crops Research Institute for the Semi-Arid Tropics (ICRISAT) has been a significant milestone in the development of cutting-edge genetic and genomic resources in pearl millet. Recently, the application of genomics and molecular technologies has provided better insights into genetic architecture and patterns of heterotic gene pools. Development of whole-genome prediction models incorporating heterotic gene pool models, mapped traits and markers have the potential to take heterosis breeding to a new level in pearl millet. This review discusses advances and prospects in various fronts of heterosis for pearl millet.
Keywords: heterosis; hybrid vigor; pearl millet; genome sequence; heterotic gene pools; genomic selection
The phenomenon of heterosis has fascinated plant breeders ever since it was first described by Charles Darwin in 1876 in the vegetable kingdom and later elaborated by George H Shull and Edward M East in maize during 1908. Heterosis is the phenotypic and functional superiority manifested in the F1 crosses over the parents. Various classical complementation mechanisms gave way to the study of the underlying potential cellular and molecular mechanisms responsible for heterosis. In cereals, such as maize, heterosis has been exploited very well, with the development of many single-cross hybrids that revolutionized the yield and productivity enhancements.
- heterosis
- hybrid vigor
- pearl millet
- genome sequence
- heterotic gene pools
- genomic selection
1. Introduction
1. Introduction
Heterosis (syn hybrid vigor) is a natural phenomenon whereby hybrid (first filial generation, i.e., F
) offsprings of genetically diverse individuals exhibit improved physical and functional features relative to their parents [
,
]. Heterosis has been studied in most eukaryotic organisms, including plants, animals, and fungi. When two homozygous inbred lines (true breeding line derived from recurrent inbreeding) with distinct genetic constitutions are hybridized together, the resultant hybrids have greater height, weight, fertility, robustness, and constitutional vigor than either of the parents and their self-pollinated counterparts [
]. Naturally cross-pollinating species like pearl millet, rye, maize, and other grasses typically exhibit a higher degree of heterosis than the self-pollinating crop plants such as rice, barley, wheat, and oats. Nevertheless, many hybrid cultivars have also been developed in self-pollinating plant species [
]. Heterosis can manifest by virtue of improvement of several traits during plant growth and development. In pearl millet, considerable growth differences between hybrids and their parents can be monitored during different stages of growth and development. Post-germination, two root architectural traits such as primary root length and lateral root density show differences at early as well as later stages. During late development stages, the F
hybrids display relatively more luxurious growth, greater biomass accumulation, and higher seed setting than their parent genotypes. It is noteworthy that the degree of heterosis can vary considerably among various traits. For example, grain iron (Fe) and zinc (Zn) contents in pearl millet F
hybrids do not improve over their parental lines and require high levels of micronutrients in both parents. The maximum level of heterosis is monitored in the F
derived from cross-pollination between diverse genotypes. This demonstrates the involvement of several alleles or genetic loci with diverse interactions at cellular and molecular levels. While the superiority of the progenies over their parents is progressively reduced in the successive generations developed from self-pollination. Heterosis has been exploited in many cereals, oilseeds, vegetables, fruits, pulses, etc., through the development of F
hybrids globally [
The Evolution of Understanding of Heterosis
].
2. Molecular Bases of Heterosis
The perceived greater vigor in the heterotic phenotype than its inbred parental lines is the cumulative result of the coding of genetic information by transcription, translation and their levels of regulation. With the advent of next-generation sequencing (NGS) technologies, genome-wide single-nucleotide polymorphisms (SNPs), insertion or deletions (indels), large structural variations in the form of presence and absence variations (PAVs), and copy number variations (CNVs) that contribute to the phenotypic diversity can be easily deciphered [
,
,
]. To decode the underlying mechanisms determining the degree of vigor differences between inbreds and hybrids, molecular studies were advanced to assess their transcriptome, proteome, epigenome and other regulatory related mechanisms [
]. At a glance,
gives a basic idea on the molecular bases of heterosis.
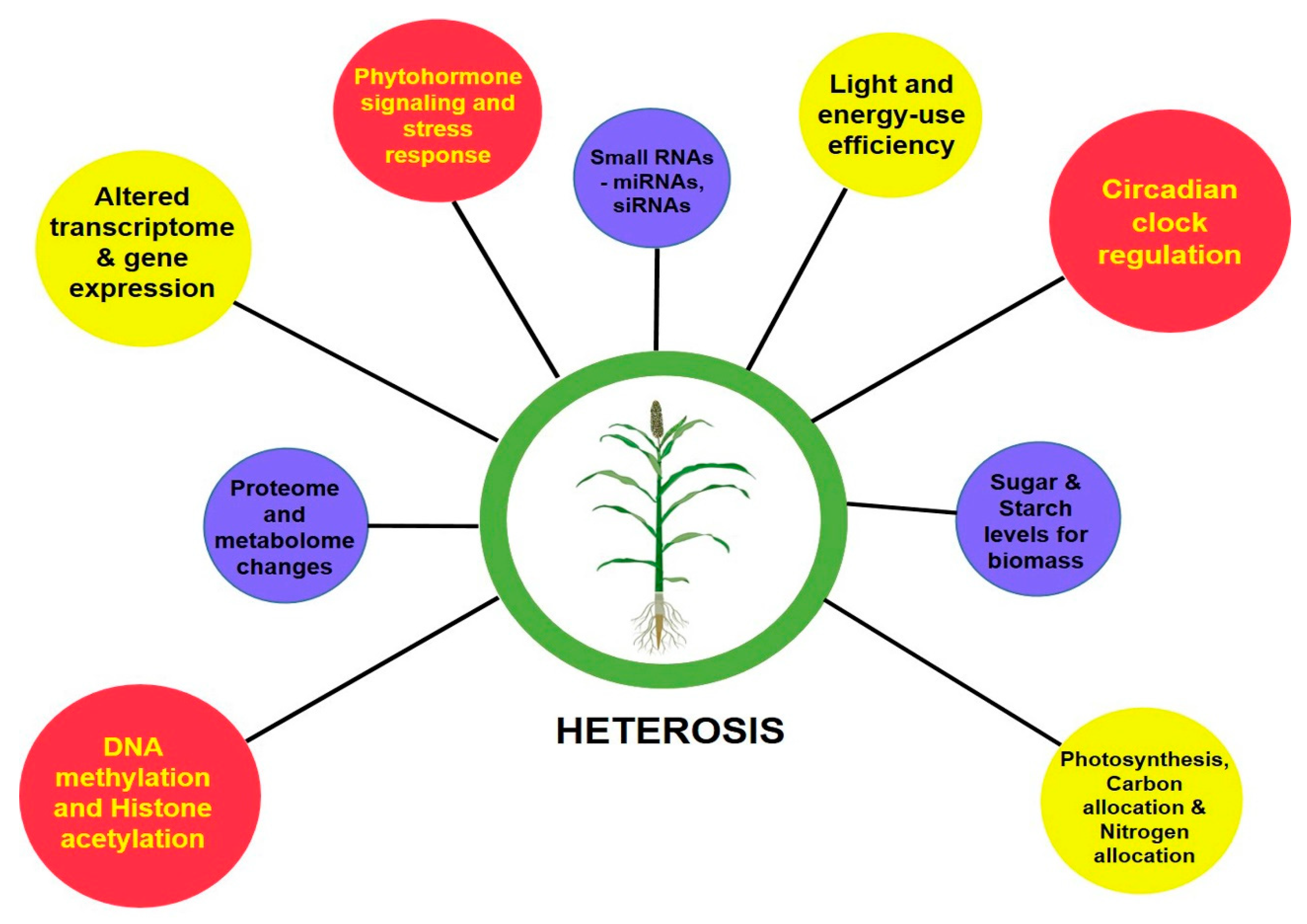
2.1. Transcriptomics View on Heterosis
Hybridization of the inbred parental lines leads to interactions between the nucleus and cytoplasm and the resulting changes at the cellular and molecular level leads to the altered patterns of gene expression. Genome-wide modifications in the gene expression levels and their mechanism of actions in the hybrids vis-à-vis its inbreds have been documented in several hybrids of maize [
,
,
], rice [
], wheat [
], cotton [
],
[
,
,
], etc. Transcriptomics and its potential in heterosis can best be viewed as a transitional phase in between the genetic information and the plant phenotype which specifically measures the relative contribution of each allele in a hybrid [
]. To identify the genes involved in heterosis using transcriptome analyses, technologies like microarray, RNA-sequencing (RNA-seq), etc., are being used to compare inbred parental lines with their F
progenies. From the initial transcriptomic studies on several crop species, it was believed that favorable gene expression levels in hybrids are predominant when compared to its inbred parents [
,
,
,
]. However, it is important to note that differential gene expression levels may not directly correspond to the protein activity between inbred parental lines and hybrids, or to the observed heterosis in the hybrids, and post transcription/translation regulations need to be taken into account [
,
].
2.2. Proteomics View on Heterosis
Studying the role of proteins in determining heterosis is important, as changes at the level of transcripts may not always reflect at the protein level due to various post-transcriptional and translational regulatory mechanisms [
]. The generalized concept is that, in the inbred parental lines protein metabolism is higher due to unstable protein levels which require more energy to quench. Consequently, less energy remains for its vegetative growth, biomass, and yield in the end. The genetic basis for this condition in inbred parental lines is due to the lack of allelic choice in their homozygous condition with only two alleles, whereas hybrids in polyploidy condition will have more alleles and thus display faster growth rates due to increased cell divisions [
].
With the advent of 2-D gel electrophoresis (2-DGE) along with mass spectrometry (MS), numerous studies have been reported to identify the differentially expressed proteins (DEPs) determining heterosis. In addition, MS-based protein detection and quantification depend upon the two isobaric labeling reagents like tandem mass tags (TMT) and isobaric tags for relative and absolute quantification (iTRAQ), which are employed to detect altered proteins or DEPs in heterotic phenotypes [
,
]. To date, the majority of the DEPs determining heterosis have been identified in major crop species like maize, wheat and rice from the tissue samples of the embryo, root, and leaf [
,
,
]. It is apparent from these studies that the majority of the DEPs identified between inbreds and hybrids are due to the non-additive gene effects, and it has also been reported that these DEPs belong to the pathways of signal transduction, glycolysis, photosynthesis, disease resistance, carbon metabolism, protein, amino acid metabolism, etc. [
]. These results indicate that the degree of heterosis was dependent upon the incidence of protein isoforms or modifications [
].
2.3. Epigenomics View on Heterosis
DNA methylation: The resulting vigor in hybrids after the hybridization of two distant inbred parental lines is often linked with epigenetic modifications, viz., DNA methylation [
], chromatin structure modification [
], histone acetylation [
], or small RNA induced regulations, etc. [
]. Genome activity and the cellular development of any crop species are indeed regulated by DNA methylation. In most plant species, DNA methylation takes place on the 5’ position of cytosine residues at CG, CHH and CHG regions (where H may be A, T or C) by DNA methyltransferases [
,
]. The degree of DNA methylation change in the hybrids depends on the diversity among inbred parents [
]. Using bisulfite and siRNA sequencing, differential methylated loci were identified in the rice hybrids (Nipponbare and Indica) and their parents to decipher the role of DNA methylation in causing epigenetic heritability [
]. In the allopolyploids of
, genes with cis- but not trans-regulatory changes were augmented in loci that were hypo-methylated or hyper-methylated [
]. The manifestation of heterosis by DNA methylation is mainly mediated by the suppression of the transcription process of the regulatory genes involved in enhancing inbreeding depression or by promoting the expression of genes for heterosis [
]. Greaves et al. [
] suggested that DNA methylation sites in hybrids were frequently associated with regions that are differentially methylated in their inbred parents. In particular, methylated regions in the inbred parents were usually covered by the siRNA levels, and this implies that RNA-directed DNA methylation (RdDM) pathway may induce the remodeling of DNA methylation sites in hybrids to manifest heterosis [
].
Apart from DNA methylation, another epigenetic system involved in hybrid vigor is histone modification. These modifications can occur at the post-translational level for the amino acids in histone proteins at the N-terminal tails in the form of methylation, phosphorylation, and acetylation of certain residues [
]. Usual modifications will occur with the histones like H3K9ac and H3K4me3 found in euchromatin regions with active gene expression. Meanwhile, histones H3K27me3 and H3K4me3 were found in the pericentromeric, heterochromatin and transposable element (TE) regions with low transcript levels [
]. He et al. [
] compared differential expression patterns of H3K4me3 and H3K27me3 between hybrids and parents of rice subspecies and found that H3K4me3 is transcriptionally active and H3K27me3 is inactive. In maize F
hybrid endosperm-derived transcriptomes, a histone alternate HTA112 showed significant variations in expression when compared to its parental lines [
]. These studies, though limited, raised the possibility of inducing heterosis through epigenetic histone modifications.
Small RNAs: Small RNAs, mediating gene expression, and epigenetic regulation refer to the class of microRNAs (miRNAs), small interfering RNAs (siRNAs), and
-acting siRNAs (tasiRNAs) [
]. Usually, siRNAs will maintain the genome stability by affecting the genes tagged with transposable elements (TEs) and also by retaining the stable inheritance of repeat-associated siRNAs, whereas miRNAs and tasiRNAs are involved in controlling morphological and developmental traits [
]. The combination of distant inbred parental siRNAs in the hybrids will exert both
and
-acting effects on TEs and TE-coding genes which causes genomic instability thereby leading to the abortive embryo and endosperm formations termed as hybrid lethality [
]; on the contrary, some small RNAs also play positive regulatory roles in terms of protection from the genomic shock, ultimately leading to hybrid vigor. Small RNA gene expression studies in the hybrids of rice,
and maize revealed the downregulated levels of 24-nt siRNAs when compared to their inbred parental lines. The 24-nt siRNAs that are involved in the RdDM mechanism will finally lead to transcriptional repression by stimulating the DNA methylation process. Hence, the reduction of these siRNA levels may lead to the ample expression of protein-coding genes in hybrids [
,
]. The differential expression of siRNAs in hybrids relative to their parents is due to the differences in the promoter regions of small RNA coding genes. Therefore, the epigenetically developed differences at the gene expression level of 24-nt siRNAs in hybrids may contribute to heterosis [
,
,
].
2.4. Genes Associated with Heterosis
3. Unifying Theory for Heterosis
4. Exploitation of Heterosis in Cereals

5. Pearl Millet Introduction and Importance
5.1. Climate Resilience
5.2. Nutritional Aspects
6. History of Hybrid Development in Pearl Millet
6.1. Development of Male Sterile Lines
6.2. Development of CMS System—A1, A2, A4 and A5 Systems
6.3. Work Done in India on Pearl Millet Hybrid Breeding
6.4. Development of Restorers and Maintainers
7. Heterosis and Genomics
8. Development of Heterotic Gene Pools

9. Development of Whole Genome Prediction Models
10. Future Prospects
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Coors, C.G.; Pandey, S. (Eds.) The Genetics and Exploitation of Heterosis in Crops; American Society of Agronomy: Madison, WI, USA, 1999; pp. 99–118. [Google Scholar]
- Shull, G.H. What is heterosis? Genetics 1948, 33, 439–446. [Google Scholar]
- Darwin, C.R. The Effects of Cross and Self Fertilization in the Vegetable Kingdom; John Murray: London, UK, 1876; p. 482. [Google Scholar]
- Rajendrakumar, P.; Hariprasanna, K.; Seetharama, N. Prediction of Heterosis in Crop Plants—Status and Prospects. Am. J. Exp. Agric. 2015, 9, 1–16. [Google Scholar] [CrossRef]
- Mendel, G. Experiments in Plant Hybridization; Harvard University Press: Cambridge, MA, USA, 1977. [Google Scholar]
- Zhou, G.; Chen, Y.; Yao, W.; Zhang, C.; Xie, W.; Hua, J.; Xing, Y.; Xiao, J.; Zhang, Q. Genetic composition of yield heterosis in an elite rice hybrid. Proc. Natl. Acad. Sci. USA 2012, 109, 15847–15852. [Google Scholar] [CrossRef] [PubMed]
- Groszmann, M.; Greaves, I.K.; Albertyn, Z.I.; Scofield, G.N.; Peacock, W.J.; Dennis, E.S. Changes in 24-nt siRNA levels in Arabidopsis hybrids suggest an epigenetic contribution to hybrid vigor. Proc. Natl. Acad. Sci. USA 2011, 108, 2617–2622. [Google Scholar] [CrossRef] [PubMed]
- Duvick, D.N.; Coors, J.G.; Pandey, S. Heterosis: Feeding People and Protecting Natural Resources. Soil Surv. Land Use Plan. 2015, 1, 19–29. [Google Scholar]
- Just, T.; Reed, H.S.; Verdoorn, F. A Short History of the Plant Sciences. Am. Midl. Nat. 1943, 30, 810. [Google Scholar] [CrossRef]
- Bruce, A.B. The Mendelian Theory of Heredity and the Augmentation of Vigor. Science 1910, 32, 627–628. [Google Scholar] [CrossRef]
- Jones, D.F. Dominance of Linked Factors as a Means of Accounting for Heterosis. Proc. Natl. Acad. Sci. USA 1917, 3, 310–312. [Google Scholar] [CrossRef]
- East, E.M. Heterosis. Genetics 1936, 21, 375–397. [Google Scholar]
- Chen, Z.J. Molecular mechanisms of polyploidy and hybrid vigor. Trends Plant. Sci. 2010, 15, 57–71. [Google Scholar] [CrossRef]
- Fu, D.; Xiao, M.; Hayward, A.; Fu, Y.; Liu, G.; Jiang, G.; Zhang, H. Utilization of crop heterosis: A review. Euphytica 2014, 197, 161–173. [Google Scholar] [CrossRef]
- Charlesworth, B.; Charlesworth, D. The genetic basis of inbreeding depression. Genet. Res. 1999, 74, 329–340. [Google Scholar] [CrossRef] [PubMed]
- McClintock, B. The significance of responses of the genome to challenge. Science 1984, 226, 792–801. [Google Scholar] [CrossRef] [PubMed]
- Ha, M.; Lu, J.; Tian, L.; Ramachandran, V.; Kasschau, K.D.; Chapman, E.J.; Carrington, J.C.; Chen, X.; Wang, X.-J.; Chen, Z.J. Small RNAs serve as a genetic buffer against genomic shock in Arabidopsis interspecific hybrids and allopolyploids. Proc. Natl. Acad. Sci. USA 2009, 106, 17835–17840. [Google Scholar] [CrossRef]
- Lippman, Z.B.; Cohen, O.; Alvarez, J.P.; Abu-Abied, M.; Pekker, I.; Paran, I.; Eshed, Y.; Zamir, D. The Making of a Compound Inflorescence in Tomato and Related Nightshades. PLoS Biol. 2008, 6, e288. [Google Scholar] [CrossRef]
- Velu, G.; Rai, K.N.; Muralidharan, V.; Longvah, T.; Crossa, J. Gene effects and heterosis for grain iron and zinc density in pearl millet (Pennisetum glaucum (L.) R. Br). Euphytica 2011, 180, 251–259. [Google Scholar] [CrossRef]
- Díaz, A.; Zikhali, M.; Turner, A.S.; Isaac, P.; Laurie, D.A. Copy number variation affecting the photoperiod-B1 and vernalization-A1 genes is associated with altered flowering time in wheat (Triticum aestivum). PLoS ONE 2012, 7, 1–11. [Google Scholar] [CrossRef]
- Zmienko, A.; Samelak-Czajka, A.; Kozlowski, P.; Figlerowicz, M. Copy number polymorphism in plant genomes. Theor. Appl. Genet. 2013, 127, 1–18. [Google Scholar] [CrossRef]
- Saxena, R.K.; Edwards, D.; Varshney, R.K. Structural variations in plant genomes. Brief. Funct. Genom. 2014, 13, 296–307. [Google Scholar] [CrossRef]
- Kaeppler, S. Heterosis: Many Genes, Many Mechanisms—End the Search for an Undiscovered Unifying Theory. Isrn Bot. 2012, 2012, 1–12. [Google Scholar] [CrossRef]
- Swanson-Wagner, R.A.; Jia, Y.; DeCook, R.; Borsuk, L.A.; Nettleton, D.S.; Schnable, P.S. All possible modes of gene action are observed in a global comparison of gene expression in a maize F1 hybrid and its inbred parents. Proc. Natl. Acad. Sci. USA 2006, 103, 6805–6810. [Google Scholar] [CrossRef] [PubMed]
- Guo, M.; Rupe, M.A.; Yang, X.; Crasta, O.; Zinselmeier, C.; Smith, O.S.; Bowen, B. Genome-wide transcript analysis of maize hybrids: Allelic additive gene expression and yield heterosis. Theor. Appl. Genet. 2006, 113, 831–845. [Google Scholar] [CrossRef] [PubMed]
- Stupar, R.M.; Springer, N.M. Cis-transcriptional Variation in Maize Inbred Lines B73 and Mo17 Leads to Additive Expression Patterns in the F1Hybrid. Genetics 2006, 173, 2199–2210. [Google Scholar] [CrossRef] [PubMed]
- He, G.; Zhu, X.; Elling, A.A.; Chen, L.; Wang, X.; Guo, L.; Liang, M.; He, H.; Zhang, H.; Chen, F.; et al. Global Epigenetic and Transcriptional Trends among Two Rice Subspecies and Their Reciprocal Hybrids. Plant. Cell 2010, 22, 17–33. [Google Scholar] [CrossRef] [PubMed]
- Wang, Z.; Ni, Z.; Wu, H.; Nie, X.; Sun, Q. Heterosis in root development and differential gene expression between hybrids and their parental inbreds in wheat (Triticum aestivum L.). Theor. Appl. Genet. 2006, 113, 1283–1294. [Google Scholar] [CrossRef]
- Flagel, L.; Udall, J.A.; Nettleton, D.S.; Wendel, J.F. Duplicate gene expression in allopolyploid Gossypium reveals two temporally distinct phases of expression evolution. BMC Biol. 2008, 6, 16. [Google Scholar] [CrossRef]
- Shen, H.; He, H.; Li, J.; Chen, W.; Wang, X.; Guo, L.; Peng, Z.; He, G.; Zhong, S.; Qi, Y.; et al. Genome-Wide Analysis of DNA Methylation and Gene Expression Changes in Two Arabidopsis Ecotypes and Their Reciprocal Hybrids. Plant. Cell 2012, 24, 875–892. [Google Scholar] [CrossRef]
- Fujimoto, R.; Taylor, J.; Shirasawa, S.; Peacock, W.J.; Dennis, E.S. Heterosis of Arabidopsis hybrids between C24 and Col is associated with increased photosynthesis capacity. Proc. Natl. Acad. Sci. USA 2012, 109, 7109–7114. [Google Scholar] [CrossRef]
- 32. Fujimoto, R; Taylor, J.M; Sasaki, T; Kawanabe, T; Dennis, E.S. Genome wide gene expression in artificially synthesized amphidiploids of Arabidopsis. Plant Mol. Biol. 2011, 77, 419–431. [CrossRef]
- Schnable, P.S.; Springer, N.M. Progress Toward Understanding Heterosis in Crop Plants. Annu. Rev. Plant. Biol. 2013, 64, 71–88. [Google Scholar] [CrossRef]
- Comings, D.E.; MacMurray, J.P. Molecular Heterosis: A Review. Mol. Genet. Metab. 2000, 71, 19–31. [Google Scholar] [CrossRef] [PubMed]
- Baranwal, V.K.; Mikkilineni, V.; Zehr, U.B.; Tyagi, A.K.; Kapoor, S. Heterosis: Emerging ideas about hybrid vigour. J. Exp. Bot. 2012, 63, 6309–6314. [Google Scholar] [CrossRef] [PubMed]
- Stupar, R.M.; Gardiner, J.; Oldre, A.; Haun, W.J.; Chandler, V.L.; Springer, N.M. Gene expression analyses in maize inbreds and hybrids with varying levels of heterosis. BMC Plant. Biol. 2008, 8, 33. [Google Scholar] [CrossRef] [PubMed]
- Fujimoto, R.; Uezono, K.; Ishikura, S.; Osabe, K.; Peacock, W.J.; Dennis, E.S. Recent research on the mechanism of heterosis is important for crop and vegetable breeding systems. Breed. Sci. 2018, 68, 145–158. [Google Scholar] [CrossRef]
- Xing, J.; Sun, Q.; Ni, Z. Proteomic patterns associated with heterosis. Biochim. Biophys. Acta (Bba)—Proteins Proteom. 2016, 1864, 908–915. [Google Scholar] [CrossRef]
- Goff, S.A. A unifying theory for general multigenic heterosis: Energy efficiency, protein metabolism, and implications for molecular breeding. New Phytol. 2010, 189, 923–937. [Google Scholar] [CrossRef]
- Wang, J.; Yu, Q.; Xiong, H.; Wang, J.; Chen, S.; Yang, Z.; Dai, S. Proteomic Insight into the Response of Arabidopsis Chloroplasts to Darkness. PLoS ONE 2016, 11, e0154235. [Google Scholar] [CrossRef]
- Guo, B.; Chen, Y.; Zhang, G.; Xing, J.; Hu, Z.; Feng, W.; Yao, Y.; Peng, H.; Du, J.; Zhang, Y.; et al. Comparative Proteomic Analysis of Embryos between a Maize Hybrid and Its Parental Lines during Early Stages of Seed Germination. PLoS ONE 2013, 8, e65867. [Google Scholar] [CrossRef]
- Song, X.; Ni, Z.; Yao, Y.; Xie, C.; Li, Z.; Wu, H.; Zhang, Y.; Sun, Q. Wheat (Triticum aestivum L.) root proteome and differentially expressed root proteins between hybrid and parents. Proteomics 2007, 7, 3538–3557. [Google Scholar] [CrossRef]
- Zhang, C.; Yin, Y.; Zhang, A.; Lu, Q.; Wen, X.; Zhu, Z.; Zhang, L.; Lu, C. Comparative proteomic study reveals dynamic proteome changes between super hybrid rice LYP9 and its parents at different developmental stages. J. Plant. Physiol. 2012, 169, 387–398. [Google Scholar] [CrossRef]
- Marcon, C.; Schutzenmeister, A.; Schutz, W.; Madlung, J.; Piepho, H.P.; Hochholdinger, F. Non-additive protein accumulation patterns in maize (Zea mays L) hybrids during embryo development. J. Prot. Res. 2010, 9, 6511–6522. [Google Scholar] [CrossRef] [PubMed]
- Parisod, C.; Salmon, A.; Zerjal, T.; Tenaillon, M.; Grandbastien, M.-A.; Ainouche, M. Rapid structural and epigenetic reorganization near transposable elements in hybrid and allopolyploid genomes in Spartina. New Phytol. 2009, 184, 1003–1015. [Google Scholar] [CrossRef] [PubMed]
- Moghaddam, A.M.B.; Colot, V.; Mette, F.; Houben, A. Heterosis and chromatin structure: Does intraspecific hybridization trigger epigenetic changes? Chrom. Res. 2007, 15, 23. [Google Scholar]
- Tanabata, T.; Taguchi-Shiobara, F.; Kishimoto, N.; Chechetka, S.; Shinomura, T.; Habu, Y. A phenomics approach detected differential epigenetic growth regulation between inbreds and their hybrid in Oryza sativa. Mol. Breed. 2010, 26, 729–734. [Google Scholar] [CrossRef]
- Groszmann, M.; Greaves, I.K.; Albert, N.; Fujimoto, R.; Helliwell, C.A.; Dennis, E.S.; Peacock, W.J. Epigenetics in plants—Vernalization and hybrid vigour. Biochim. Biophys. Acta 2011, 1809, 427–437. [Google Scholar] [CrossRef] [PubMed]
- Law, J.A.; Jacobsen, S.E. Establishing, maintaining and modifying DNA methylation patterns in plants and animals. Nat. Rev. Genet. 2010, 11, 204–220. [Google Scholar] [CrossRef] [PubMed]
- Fernie, A.R.; Chen, B.; Wang, X.; Li, X.; Li, J.; He, H.; Yang, M.; Lu, L.; Qi, Y.; Wang, X.; et al. Conservation and divergence of transcriptomic and epigenomic variation in maize hybrids. Genome Biol. 2013, 14, R57. [Google Scholar] [CrossRef]
- Chen, Z.J. Genomic and epigenetic insights into themolecular bases of heterosis. Nat. Rev. Genet. 2013, 14, 471–482. [Google Scholar] [CrossRef]
- Chodavarapu, R.K.; Feng, S.; Ding, B.; Simon, S.A.; Lopez, D.; Jia, Y.; Wang, G.-L.; Meyers, B.C.; Jacobsen, S.E.; Pellegrini, M. Transcriptome and methylome interactions in rice hybrids. Proc. Natl. Acad. Sci. USA 2012, 109, 12040–12045. [Google Scholar] [CrossRef]
- Shi, X.; Ng, D.W.-K.; Zhang, C.; Comai, L.; Ye, W.; Chen, Z.J. Cis- and trans-regulatory divergence between progenitor species determines gene-expression novelty in Arabidopsis allopolyploids. Nat. Commun. 2012, 3, 950. [Google Scholar] [CrossRef]
- Nakamura, S.; Hosaka, K. DNA methylation in diploid inbred lines of potatoes and its possible role in the regulation of heterosis. Theor. Appl. Genet. 2009, 120, 205–214. [Google Scholar] [CrossRef] [PubMed]
- Greaves, I.K.; Groszmann, M.; Ying, H.; Taylor, J.; Peacock, W.J.; Dennis, E.S. Trans chromosomal methylation in Arabidopsis hybrids. Proc. Natl. Acad. Sci. USA 2012, 109, 3570–3575. [Google Scholar] [CrossRef] [PubMed]
- Greaves, I.K.; Eichten, S.R.; Groszmann, M.; Wang, A.; Ying, H.; Peacock, W.J.; Dennis, E.S. Twenty-four–nucleotide siRNAs produce heritable trans-chromosomal methylation in F1 Arabidopsis hybrids. Proc. Natl. Acad. Sci. USA 2016, 113, E6895–E6902. [Google Scholar] [CrossRef] [PubMed]
- Berger, S.L. The complex language of chromatin regulation during transcription. Nature 2007, 447, 407–412. [Google Scholar] [CrossRef] [PubMed]
- Roudier, F.; Teixeira, F.K.; Colot, V. Chromatin indexing in Arabidopsis: An epigenomic tale of tails and more. Trends Genet. 2009, 25, 511–517. [Google Scholar] [CrossRef]
- Jahnke, S.; Sarholz, B.; Thiemann, A.; Kühr, V.; Gutierrez-Marcos, J.F.; Geiger, H.H.; Piepho, H.-P.; Scholten, S. Heterosis in early seed development: A comparative study of F1 embryo and endosperm tissues 6 days after fertilization. Theor. Appl. Genet. 2009, 120, 389–400. [Google Scholar] [CrossRef]
- Chen, X. Small RNAs and their roles in plant development. Annu. Rev. Cell Dev. Biol. 2009, 25, 21–44. [Google Scholar] [CrossRef]
- Xie, F.; He, Z.; Esguerra, M.Q.; Qiu, F.; Ramanathan, V. Determination of heterotic groups for tropical Indica hybrid rice germplasm. Theor. Appl. Genet. 2013, 127, 407–417. [Google Scholar] [CrossRef]
- Ng, D.W.-K.; Lu, J.; Chen, Z.J. Big roles for small RNAs in polyploidy, hybrid vigor, and hybrid incompatibility. Curr. Opin. Plant. Biol. 2012, 15, 154–161. [Google Scholar] [CrossRef]
- Greaves, I.K.; Gonzalez-Bayon, R.; Wang, L.; Zhu, A.; Liu, P.-C.; Groszmann, M.; Peacock, W.J.; Dennis, E.S. Epigenetic Changes in Hybrids. Plant. Physiol. 2015, 168, 1197–1205. [Google Scholar] [CrossRef]
- Groszmann, M.; Greaves, I.K.; Fujimoto, R.; Peacock, W.J.; Dennis, E.S. The role of epigenetics in hybrid vigour. Trends Genet. 2013, 29, 684–690. [Google Scholar] [CrossRef] [PubMed]
- Barber, W.T.; Zhang, W.; Win, H.; Varala, K.; Dorweiler, J.E.; Hudson, M.E.; Moose, S.P. Repeat associated small RNAs vary among parents and following hybridization in maize. Proc. Natl. Acad. Sci. USA 2012, 109, 10444–10449. [Google Scholar] [CrossRef] [PubMed]
- Busov, V.; Brunner, A.M.; Strauss, S.H. Genes for control of plant stature and form. New Phytol. 2008, 177, 589–607. [Google Scholar] [CrossRef] [PubMed]
- Krizek, B.A. Making bigger plants: Key regulators of final organ size. Curr. Opin. Plant. Biol. 2009, 12, 17–22. [Google Scholar] [CrossRef] [PubMed]
- Hu, Y.; Xie, Q.; Chua, N.-H. The Arabidopsis Auxin-Inducible Gene ARGOS Controls Lateral Organ Size. Plant. Cell 2003, 15, 1951–1961. [Google Scholar] [CrossRef] [PubMed]
- Guo, M.; Rupe, M.A.; Dieter, J.A.; Zou, J.; Spielbauer, D.; Duncan, K.E.; Howard, R.J.; Hou, Z.; Simmons, C.R. Cell Number Regulator1 Affects Plant and Organ Size in Maize: Implications for Crop Yield Enhancement and Heterosis. Plant. Cell 2010, 22, 1057–1073. [Google Scholar] [CrossRef]
- Krieger, U.; Lippman, Z.B.; Zamir, D. The flowering gene SINGLE FLOWER TRUSS drives heterosis for yield in tomato. Nat. Genet. 2010, 42, 459–463. [Google Scholar] [CrossRef]
- Li, A.; Zhou, Y.; Jin, C.; Song, W.; Chen, C.; Wang, C. LaAP2L1, a Heterosis-Associated AP2/EREBP Transcription Factor of Larix, Increases Organ Size and Final Biomass by Affecting Cell Proliferation in Arabidopsis. Plant. Cell Physiol. 2013, 54, 1822–1836. [Google Scholar] [CrossRef]
- Li, D.; Huang, Z.; Song, S.; Xin, Y.; Mao, D.; Lv, Q.; Zhou, M.; Tian, D.; Tang, M.; Wu, Q.; et al. Integrated analysis of phenome, genome, and transcriptome of hybrid rice uncovered multiple heterosis-related loci for yield increase. Proc. Natl. Acad. Sci. USA 2016, 113, E6026–E6035. [Google Scholar] [CrossRef]
- Huang, Y.; Zhang, L.; Zhang, J.; Yuan, D.; Xu, C.; Li, X.; Zhou, D.-X.; Wang, S.; Zhang, Q. Heterosis and polymorphisms of gene expression in an elite rice hybrid as revealed by a microarray analysis of 9198 unique ESTs. Plant. Mol. Biol. 2006, 62, 579–591. [Google Scholar] [CrossRef]
- Shapira, R.; Levy, T.; Shaked, S.; Fridman, E.; David, L. Extensive heterosis in growth of yeast hybrids is explained by a combination of genetic models. Heredity 2014, 113, 316–326. [Google Scholar] [CrossRef] [PubMed]
- Steinmetz, L.M.; Sinha, H.; Richards, D.R.; Spiegelman, J.I.; Oefner, P.J.; McCusker, J.H.; Davis, R.W. Dissecting the architecture of a quantitative trait locus in yeast. Nature 2002, 416, 326–330. [Google Scholar] [CrossRef] [PubMed]
- Fiévet, J.B.; Nidelet, T.; Dillmann, C.; De Vienne, D. Heterosis Is a Systemic Property Emerging From Non-linear Genotype-Phenotype Relationships: Evidence From in Vitro Genetics and Computer Simulations. Front. Genet. 2018, 9, 1–26. [Google Scholar] [CrossRef]
- Cornish-Bowden, A.; Cárdenas, M.L. Contrasting theories of life: Historical context, current theories. In search of an ideal theory. Biosystems 2020, 188, 104063. [Google Scholar] [CrossRef] [PubMed]
- Hauben, M.; Haesendonckx, B.; Standaert, E.; Van Der Kelen, K.; Azmi, A.; Akpo, H.; Van Breusegem, F.; Guisez, Y.; Bots, M.; Lambert, B.; et al. Energy use efficiency is characterized by an epigenetic component that can be directed through artificial selection to increase yield. Proc. Natl. Acad. Sci. USA 2009, 106, 20109–20114. [Google Scholar] [CrossRef]
- Ni, Z.; Kim, E.-D.; Ha, M.; Lackey, E.; Liu, J.; Zhang, Y.; Sun, Q.; Chen, Z.J. Altered circadian rhythms regulate growth vigour in hybrids and allopolyploids. Nature 2008, 457, 327–331. [Google Scholar] [CrossRef]
- Meyer, R.C.; Witucka-Wall, H.; Becher, M.; Blacha, A.; Boudichevskaia, A.; Dörmann, P.; Fiehn, O.; Friedel, S.; Von Korff, M.; Lisec, J.; et al. Heterosis manifestation during early Arabidopsis seedling development is characterized by intermediate gene expression and enhanced metabolic activity in the hybrids. Plant J. 2012, 71, 669–683. [Google Scholar] [CrossRef]
- Mulualem, T.; Abate, M. Heterotic Response in Major Cereals and Vegetable Crops. Int. J. Plant. Breed. Genet. 2016, 10, 69–78. [Google Scholar] [CrossRef]
- Shull, G.H. The Composition of a Field of Maize. J. Hered. 1908, 4, 296–301. [Google Scholar] [CrossRef]
- Ramya, A.R.; Ahamed, M.L.; Satyavathi, C.T.; Rathore, A.; Katiyar, P.; Raj, A.G.B.; Kumar, S.; Gupta, R.; Mahendrakar, M.; Yadav, R.; et al. Towards Defining Heterotic Gene Pools in Pearl Millet [Pennisetum glaucum (L.) R. Br.]. Front. Plant Sci. 2018, 8, 8. [Google Scholar] [CrossRef]
- Bollam, S.; Pujarula, V.; Srivastava, R.K.; Gupta, R.K. Genomic Approaches to Enhance Stress Tolerance for Productivity Improvements in Pearl Millet. In Biotechnologies of Crop Improvement, Volume 3; Springer: Berlin/Heidelberg, Germany, 2018; Volume 3, pp. 239–264. [Google Scholar]
- Manning, K.; Pelling, R.; Higham, T.; Schwenniger, J.-L.; Fuller, D. 4500-Year old domesticated pearl millet (Pennisetum glaucum) from the Tilemsi Valley, Mali: New insights into an alternative cereal domestication pathway. J. Archaeol. Sci. 2011, 38, 312–322. [Google Scholar] [CrossRef]
- Hash, T.; Raj, A.B.; Lindup, S.; Sharma, A.; Beniwal, C.; Folkertsma, R.; Mahalakshmi, V.; Zerbini, E.; Blümmel, M. Opportunities for marker-assisted selection (MAS) to improve the feed quality of crop residues in pearl millet and sorghum. Field Crop. Res. 2003, 84, 79–88. [Google Scholar] [CrossRef]
- Basavaraj, S.H.; Singh, V.K.; Singh, A.; Singh, A.; Singh, A.; Anand, D.; Yadav, S.; Ellur, R.K.; Singh, D.; Krisnan, S.G.; et al. Marker-assisted improvement of bacterial blight resistance in parental lines of Pusa RH10, a superfine grain aromatic rice hybrid. Mol. Breed. 2010, 26, 293–305. [Google Scholar] [CrossRef]
- Nambiar, V.S.; Dhaduk, J.J.; Sareen, N.; Shahu, T. and Desai, R. Potential functional implications of pearl millet (Pennisetum glaucum) in health and disease. J. Appl. Pharm. Sci. 2011, 1, 62–67. [Google Scholar]
- Vadez, V.; Hash, T.; Bidinger, F.R.; Kholova, J. Phenotyping pearl millet for adaptation to drought. In Drought Phenotyping in Crops: From Theory to Practice; Monneveux, P., Ribaut, J.M., Eds.; Frontiers E-books; 2014; p. 158. Available online: https://books.google.com.ph/books?hl=zh-TW&lr=&id=zRApAwAAQBAJ&oi=fnd&pg=PP1&dq=Drought+phenotyping+in+crops:+From+theory+to+practice&ots=bCvnXEKxEL&sig=e8dxZUCtlfbRzGX_vXhOOT2T25A&redir_esc=y#v=onepage&q=Drought%20phenotyping%20in%20crops%3A%20From%20theory%20to%20practice&f=false (accessed on 15 June 2020).
- Varshney, R.K.; Shi, C.; Thudi, M.; Mariac, C.; Wallace, J.G.; Qi, P.; Zhang, H.; Zhao, Y.; Wang, X.; Rathore, A.; et al. Pearl millet genome sequence provides a resource to improve agronomic traits in arid environments. Nat. Biotechnol. 2017, 35, 969–976. [Google Scholar] [CrossRef] [PubMed]
- Gupta, S.K.; Nepolean, T.; Sankar, S.M.; Rathore, A.; Das, R.R.; Rai, K.N.; Hash, C.T. Patterns of Molecular Diversity in Current and Previously Developed Hybrid Parents of Pearl Millet [Pennisetum glaucum (L.) R. Br.]. Am. J. Plant Sci. 2015, 6, 1697–1712. [Google Scholar] [CrossRef]
- Souci, S.W.; Fachmann, W.; Kraut, H. Food Composition and Nutrition Tables (No. Ed. 6); Medpharm GmbH Scientific Publishers: Stuttgart, Germany, 2000. [Google Scholar]
- Tako, E.; Reed, S.M.; Budiman, J.; Hart, J.; Glahn, R. Higher iron pearl millet (Pennisetum glaucum L.) provides more absorbable iron that is limited by increased polyphenolic content. Nutr. J. 2015, 14, 11. [Google Scholar] [CrossRef]
- Finkelstein, J.L.; Mehta, S.; Udipi, S.A.; Ghugre, P.S.; Luna, S.V.; Murray-Kolb, L.E.; Przybyszewski, E.M.; Haas, J.D. A randomized trial of iron-biofortified pearl millet in school children in India. J. Nutr. 2015, 145, 1576–1581. [Google Scholar] [CrossRef]
- Lardy, G.P.; Adams, D.C.; Klopfenstein, T.J.; Patterson, H.H. Building beef cow nutritional programs with the 1996 NRC beef cattle requirements model. J. Anim. Sci. 2004, 82, 83–92. [Google Scholar]
- Vadez, V.; Hash, T.; Bidinger, F.R.; Kholova, J. II.1.5 Phenotyping pearl millet for adaptation to drought. Front. Physiol. 2012, 3, 1–12. [Google Scholar] [CrossRef]
- Smith, R.L.; Jensen, L.S.; Hoveland, C.S.; Hanna, W.W. Use of Pearl Millet, Sorghum, and Triticale Grain in Broiler Diets. J. Prod. Agric. 1989, 2, 78–82. [Google Scholar] [CrossRef]
- Burton, G.W. Cytoplasmic Male-Sterility in Pearl Millet (Pennisetum glaucum) (L.) R. Br.1. Agron. J. 1907, 50, 230. [Google Scholar] [CrossRef]
- Serba, D.D.; Perumal, R.; Tesso, T.; Min, D. Status of Global Pearl Millet Breeding Programs and the Way Forward. Crop. Sci. 2017, 57, 2891–2905. [Google Scholar] [CrossRef]
- Yadav, O.P.; Rai, K.N. Genetic Improvement of Pearl Millet in India. Agric. Res. 2013, 2, 275–292. [Google Scholar] [CrossRef]
- Burton, G.W. Pearl millet Tift 23A released. Crops Soils 1965, 19, 17–19. [Google Scholar]
- Burton, G.W.; Athwal, D.S. Two Additional Sources of Cytoplasmic Male-Sterility in Pearl Millet and Their Relationship to Tift 23A 1. Crop. Sci. 1967, 7, 209–211. [Google Scholar] [CrossRef]
- Singh, S.P.; Satyavathi, C.T.; Sankar, S.M. Diversification of male sterility sources with special reference to biotic stresses. In Proceedings of the New paradigms in heterosis breeding: Conventional and molecular approaches, G.B Path university of agriculture and technology, Pantnagar, Uttarakhand, India, 10 September 2014. [Google Scholar]
- Andrews, D.J.; Kumar, K.A. Use of the West African pearl millet landrace Iniadi in cultivar development. Plant Gen. Res. News. 1996, 105, 15–22. [Google Scholar]
- Chen, L.; Liu, Y.-G. Male Sterility and Fertility Restoration in Crops. Annu. Rev. Plant. Biol. 2014, 65, 579–606. [Google Scholar] [CrossRef]
- Chang, Z.; Chen, Z.; Wang, N.; Xie, G.; Lu, J.; Yan, W.; Zhou, J.; Tang, X.; Deng, X.W. Construction of a male sterility system for hybrid rice breeding and seed production using a nuclear male sterility gene. Proc. Natl. Acad. Sci. USA 2016, 113, 14145–14150. [Google Scholar] [CrossRef]
- Rao, M.K.; Devi, K.U. Variation in expression of genic male sterility in pearl millet. J. Hered. 1983, 74, 34–38. [Google Scholar] [CrossRef]
- Yadav, D.; Gupta, S.K.; Kulkarni, V.N.; Rai, K.N.; Behl, R.K. Inheritance of A1system of cytoplasmic-nuclear male sterility in pearl millet [Pennisetum glaucum(L). R. Br.]. Cereal. Res. Commun. 2010, 38, 285–293. [Google Scholar] [CrossRef]
- Gupta, S.; Rai, K.N.; Govindaraj, M.; Rao, A. Genetics of fertility restoration of the A4 cytoplasmic-nuclear male sterility system in pearl millet. Czech. J. Genet. Plant. Breed. 2012, 48, 87–92. [Google Scholar] [CrossRef]
- Li, X.Q.; Jean, M.; Landry, B.S.; Brown, G.G. Restorer genes for different forms of Brassicacytoplasmic Male Sterility Map to a Single Nuclear Locus that Modifies Transcripts of Several Mitochondrial Genes. Proc. Natl. Acad. Sci. USA 1998, 95, 10032–10037. [Google Scholar] [CrossRef] [PubMed]
- Islam, A.; Mian, M.A.; Rasul, G.; Bashar, K.; Johora, F.-T. Development of Component Lines (CMS, Maintainer and Restorer lines) and their Maintenance Using Diversed Cytosources of Rice. Rice Res. Open Access 2015, 3, 1–5. [Google Scholar] [CrossRef]
- Delorme, V.; Keen, C.L.; Rai, K.N.; Leaver, C.J. Cytoplasmic-Nuclear Male Sterility in Pearl Millet: Comparative RFLP and Transcript Analyses of Isonuclear Male-Sterile Lines. Theor. Appl. Genet. 1997, 95, 961–968. [Google Scholar] [CrossRef]
- Pucher, A.; Hash, C.T.; Wallace, J.G.; Han, S.; Leiser, W.L.; Haussmann, B.I.G. Mapping a male-fertility restoration locus for the A4 cytoplasmic-genic male-sterility system in pearl millet using a genotyping-by-sequencing-based linkage map. BMC Plant. Biol. 2018, 18, 65. [Google Scholar] [CrossRef]
- Burton, G.W. Fertile Sterility Maintainer Mutants in Cytoplasmic Male Sterile Pearl Millet 1. Crop. Sci. 1977, 17, 635–637. [Google Scholar] [CrossRef]
- Burton, G.W. Pearl Millet. In Hybridization of Crop Plants; Wiley: Tifton, GA, USA, 1980; Volume 1, pp. 457–469. [Google Scholar]
- Hanna, W.W.; Wells, H.D.; Burton, G.W.; Monson, W.G. Registration of ÔTifleaf 2Õ pearl millet. Crop Sci. 1988, 28, 1023. [Google Scholar] [CrossRef]
- Hanna, W.W.; Hill, G.M.; Gates, R.N.; Wilson, J.; Burton, G.W. Registration of ‘Tifleaf 3’ Pearl Millet. Crop. Sci. 1997, 37, 1388. [Google Scholar] [CrossRef]
- Gulia, S.K.; Wilson, J.P.; Carter, J.; Singh, B.P. Progress in grain pearl millet research and market development. Issues New Crops New Uses 2007, 1, 196–203. [Google Scholar]
- Hanna, W.; Wilson, J.; Timper, P. Registration of Pearl Millet Parental Lines Tift 99D2A1/B1. Crop sci. 2005, 45, 2671. [Google Scholar] [CrossRef]
- Hanna, W.; Wilson, J.; Timper, P. Registration of Pearl Millet Parental Line Tift 454. Crop. Sci. 2005, 45, 2670–2671. [Google Scholar] [CrossRef]
- Burton, G.W.; Athwal, D.S. Registration of Pearl Millet Inbreds Tift 239DB2 and Tift 239DA2 1 (Reg. No. PL 5, PL 6). Corp Sci. 1969, 9, 398. [Google Scholar]
- Athwal, D.S. Hybrid bajra-1 marks a new era. Ind. Farm. 1965, 15, 6–7. [Google Scholar]
- Aken’Ova, M.E.; Chheda, H.R. A New Source of Cytoplasmic—Genic Male Sterility in Pearl Millet 1. Crop. Sci. 1981, 21, 984–985. [Google Scholar] [CrossRef]
- Appadurai, R.; Raveendran, T.S.; Nagarajan, C. A new male-sterility system in pearl millet. Indian J. Genet. Plant Breed. 1982, 52, 832–834. [Google Scholar]
- Rai, K.N. A new cytoplasmic-nuclear male sterility system in pearl millet. Plant Breed. 1995, 114, 445–447. [Google Scholar] [CrossRef]
- Marchais, L.; Pernes, J. Genetic divergence between wild and cultivated pearl millets (Pennisetum typhoides): 1. Male sterility. Zeitschrift fur Pflanzenzüchtung 1985, 95, 103–111. [Google Scholar]
- Hanna, W.W. Characteristics and Stability of a New Cytoplasmic-Nuclear Male-Sterile Source in Pearl Millet. Crop. Sci. 1989, 29, 1457–1459. [Google Scholar] [CrossRef]
- Govindaraj, M.; Rai, K.N.; Cherian, B.; Pfeiffer, W.H.; Kanatti, A.; Shivade, H. Breeding Biofortified Pearl Millet Varieties and Hybrids to Enhance Millet Markets for Human Nutrition. Agriculture 2019, 9, 106. [Google Scholar] [CrossRef]
- Khairwal, I.S.; Yadav, O.P. Pearl millet (Pennisetum glatlcum) improvement in India-retrospect and prospects. Indian Agric. Sci. 2005, 75, 183–191. [Google Scholar]
- Yadav, O.P.; Khairwal, I.S. Progress towards developing dual-purpose cultivars of pearl millet (Pennisetum glaucum) in India. Indian J. Agric. Sci. 2007, 77, 645–648. [Google Scholar]
- Gill, K.S.; Phul, P.S.; Jindla, L.N. New bajra hybrids resistant to the downy mildew green ear disease. Seed Forms 1975, 1, 3–4. [Google Scholar]
- Pokhriyal, S.C.; Unnikrishnan, K.V.; Singh, B.; Dass, R.; Patil, R.R. Combining ability of downy mildew resistant lines in pearl millet. Indian J. Genet. Plant Breed 1976, 36, 403–419. [Google Scholar]
- Kumar, P.; Sharma, V.K.; Prasad, B.D. Characterization of maintainer and restorer lines for wild abortive cytoplasmic male sterility in indica rice (’Oryza sativa’L.) using pollen fertility and microsatellite (SSR) markers. Aust. J. Crop. Sci. 2015, 9, 384. [Google Scholar]
- Amiribehzadi, A.; Satyavathi, C.T. Fertility restoration studies in different cytoplasms of pearl millet [Pennisetum glaucum (L.) R. BR.]. Ann. Agric. Sci. 2012, 3, 1–12. [Google Scholar]
- Shivhare, R.; Lata, C. Exploration of Genetic and Genomic Resources for Abiotic and Biotic Stress Tolerance in Pearl Millet. Front. Plant Sci. 2017, 7, 1125. [Google Scholar] [CrossRef]
- Lachagari, V.B.R.; Gupta, R.; Lekkala, S.P.; Mahadevan, L.; Kuriakose, B.; Chakravartty, N.; Katta, A.V.S.K.M.; Santhosh, S.; Reddy, A.R.; Thomas, G. Whole Genome Sequencing and Comparative Genomic Analysis Reveal Allelic Variations Unique to a Purple Colored Rice Landrace (Oryza sativa ssp. indica cv. Purpleputtu). Front. Plant Sci. 2019, 10, 513. [Google Scholar] [CrossRef]
- Alaux, M.; International Wheat Genome Sequencing Consortium; Rogers, J.; Letellier, T.; Flores, R.; Alfama, F.; Pommier, C.; Mohellibi, N.; Durand, S.; Kimmel, E.; et al. Linking the International Wheat Genome Sequencing Consortium bread wheat reference genome sequence to wheat genetic and phenomic data. Genome Biol. 2018, 19, 111. [Google Scholar] [CrossRef]
- Varshney, R.K.; Song, C.; Saxena, R.K.; Azam, S.; Yu, S.; Sharpe, A.G.; Cannon, S.; Baek, J.; Rosen, B.D.; Tar’An, B.; et al. Draft genome sequence of chickpea (Cicer arietinum) provides a resource for trait improvement. Nat. Biotechnol. 2013, 31, 240–246. [Google Scholar] [CrossRef]
- Kreplak, J.; Madoui, M.-A.; Cápal, P.; Novak, P.; Labadie, K.; Aubert, G.; Bayer, P.E.; Gali, K.K.; Syme, R.; Main, R.; et al. A reference genome for pea provides insight into legume genome evolution. Nat. Genet. 2019, 51, 1411–1422. [Google Scholar] [CrossRef] [PubMed]
- Varshney, R.K.; Chen, W.; Li, Y.; Bharti, A.K.; Saxena, R.K.; Schlueter, J.; A Donoghue, M.T.; Azam, S.; Fan, G.; Whaley, A.M.; et al. Draft genome sequence of pigeonpea (Cajanus cajan), an orphan legume crop of resource-poor farmers. Nat. Biotechnol. 2011, 30, 83–89. [Google Scholar] [CrossRef] [PubMed]
- Cooper, E.A.; Brenton, Z.W.; Flinn, B.S.; Grimwood, J.; Shu, S.; Flowers, D.; Luo, F.; Wang, Y.; Xia, P.; Barry, K.; et al. A new reference genome for Sorghum bicolor reveals high levels of sequence similarity between sweet and grain genotypes: Implications for the genetics of sugar metabolism. BMC Genom. 2019, 20, 420. [Google Scholar] [CrossRef] [PubMed]
- Meyer, S.; Pospisil, H.; Scholten, S. Heterosis associated gene expression in maize embryos 6 days after fertilization exhibits additive, dominant and overdominant pattern. Plant. Mol. Biol. 2006, 63, 381–391. [Google Scholar] [CrossRef]
- Zhang, X.; Wang, Y.; Yan, Y.; Peng, H.; Long, Y.; Zhang, Y.; Jiang, Z.; Liu, P.; Zou, C.; Peng, H.; et al. Transcriptome sequencing analysis of maize embryonic callus during early redifferentiation. BMC Genom. 2019, 20, 159. [Google Scholar] [CrossRef]
- Wang, C.; Tariq, R.; Ji, Z.; Wei, Z.; Zheng, K.; Mishra, R.; Zhao, K. Transcriptome analysis of a rice cultivar reveals the differentially expressed genes in response to wild and mutant strains of Xanthomonas oryzae pv. oryzae. Sci. Rep. 2019, 9, 3757. [Google Scholar] [CrossRef]
- Yan, L.; Liu, Z.; Xu, H.; Zhang, X.; Zhao, A.; Liang, F.; Xin, M.; Peng, H.; Yao, Y.; Sun, Q.; et al. Transcriptome analysis reveals potential mechanisms for different grain size between natural and resynthesized allohexaploid wheats with near-identical AABB genomes. BMC Plant. Biol. 2018, 18, 28. [Google Scholar] [CrossRef]
- Jaikishan, I.; Rajendrakumar, P.; Hariprasanna, K.; Bhat, B.V. Gene Expression Analysis in Sorghum Hybrids and Their Parental Lines at Critical Developmental Stages in Relation to Grain Yield Heterosis by Exploiting Heterosis-Related Genes from Major Cereals. Plant Mol. Biol. Rep. 2018, 36, 418–428. [Google Scholar] [CrossRef]
- Zhang, H.; Hall, N.; Goertzen, L.R.; Chen, C.Y.; Peatman, E.; Patel, J.; McElroy, J.S. Transcriptome Analysis Reveals Unique Relationships Among Eleusine Species and Heritage of Eleusine coracana. G3 Genes|Genomes|Genet. 2019, 9, 2029–2036. [Google Scholar] [CrossRef]
- Hiremath, P.J.; Farmer, A.; Cannon, S.B.; Woodward, J.; Kudapa, H.; Tuteja, R.; Kumar, A.; Bhanuprakash, A.; Mulaosmanovic, B.; Gujaria, N.; et al. Large-scale transcriptome analysis in chickpea (Cicer arietinum L.), an orphan legume crop of the semi-arid tropics of Asia and Africa. Plant Biotechnol. J. 2011, 9, 922–931. [Google Scholar] [CrossRef]
- Dudhate, A.; Shinde, H.; Tsugama, D.; Liu, S.; Takano, T. Transcriptomic analysis reveals the differentially expressed genes and pathways involved in drought tolerance in pearl millet [Pennisetum glaucum (L.) R. Br]. PLoS ONE 2018, 13, e0195908. [Google Scholar] [CrossRef] [PubMed]
- Singh, R.B.; Singh, B.; Singh, R.K. Development of potential dbEST-derived microsatellite markers for genetic evaluation of sugarcane and related cereal grasses. Ind. Crop. Prod. 2019, 128, 38–47. [Google Scholar] [CrossRef]
- Diack, O.; Kane, N.A.; Berthouly-Salazar, C.; Gueye, M.C.; Diop, B.M.; Fofana, A.; Sy, O.; Tall, H.; Zekraoui, L.; Piquet, M.; et al. New Genetic Insights into Pearl Millet Diversity As Revealed by Characterization of Early- and Late-Flowering Landraces from Senegal. Front. Plant. Sci. 2017, 8, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Anuradha, N.; Satyavathi, C.T.; Bharadwaj, C.; Nepolean, T.; Sankar, S.M.; Singh, S.P.; Meena, M.C.; Singhal, T.; Srivastava, R.K. Deciphering Genomic Regions for High Grain Iron and Zinc Content Using Association Mapping in Pearl Millet. Front. Plant Sci. 2017, 8, 170. [Google Scholar] [CrossRef]
- Taunk, J.; Rani, A.; Yadav, N.R.; Yadav, D.V.; Yadav, R.C.; Raj, K.; Kumar, R.; Yadav, H.P. Molecular breeding of ameliorating commercial pearl millet hybrid for downy mildew resistance. J. Genet. 2018, 97, 1241–1251. [Google Scholar] [CrossRef]
- Serba, D.D.; Muleta, K.T.; Amand, P.S.; Bernardo, A.; Bai, G.; Perumal, R.; Bashir, E. Genetic Diversity, Population Structure, and Linkage Disequilibrium of Pearl Millet. Plant Genome 2019, 12, 180091. [Google Scholar] [CrossRef]
- Srivastava, R.K.; Singh, R.B.; Pujarula, V.L.; Bollam, S.; Pusuluri, M.; Chellapilla, T.S.; Yadav, R.S.; Gupta, R.K. Genome-Wide Association Studies and Genomic Selection in Pearl Millet: Advances and Prospects. Front. Genet. 2020, 10, 1389. [Google Scholar] [CrossRef]
- Liang, Z.; Gupta, S.K.; Yeh, C.-T.; Zhang, Y.; Ngu, D.W.; Kumar, R.; Patil, H.T.; Mungra, K.; Yadav, D.V.; Rathore, A.; et al. Phenotypic Data from Inbred Parents Can Improve Genomic Prediction in Pearl Millet Hybrids. G3 Genes|Genomes|Genet. 2018, 8, 2513–2522. [Google Scholar] [CrossRef]
- Liu, C.J.; Witcombe, J.R.; Pittaway, T.S.; Nash, M.; Hash, C.T.; Busso, C.S.; Gale, M.D. An RFLP-based genetic map of pearl millet (Pennisetum glaucum). Theor. Appl. Genet. 1994, 89, 481–487. [Google Scholar] [CrossRef]
- Devos, K.M.; Pittaway, T.S.; Busso, C.S.; Gale, M.D.; Witcombe, J.R.; Hash, C.T. Molecular tools for the pearl millet nuclear genome. Int. Sorghum Millets Newslett. 1995, 36, 64–66. [Google Scholar]
- Rajaram, V.; Thirunavukkarasu, N.; Senthilvel, S.; Varshney, R.K.; Vadez, V.; Srivastava, R.K.; Shah, T.; Supriya, A.; Kumar, S.; Kumari, B.R.; et al. Pearl millet [Pennisetum glaucum (L.) R. Br.] consensus linkage map constructed using four RIL mapping populations and newly developed EST-SSRs. BMC Genom. 2013, 14, 159. [Google Scholar] [CrossRef] [PubMed]
- Allouis, S.; Qi, X.; Lindup, S.; Gale, M.D.; Devos, K.M. Construction of a BAC library of pearl millet, Pennisetum glaucum. Theor. Appl. Genet. 2001, 102, 1200–1205. [Google Scholar] [CrossRef]
- Kumar, S.; Hash, C.T.; Singh, G.; Basava, R.K.; Srivastava, R.K. Identification of polymorphic SSR markers in elite genotypes of pearl millet and diversity analysis. Ecol. Genet. Genom. 2020, 14, 100051. [Google Scholar] [CrossRef]
- Supriya, A.; Senthilvel, S.; Nepolean, T.; Eshwar, K.; Rajaram, V.; Shaw, R.; Hash, C.T.; Kilian, A.; Yadav, R.C.; Narasu, M.L. Development of a molecular linkage map of pearl millet integrating DArT and SSR markers. Theor. Appl. Genet. 2011, 123, 239–250. [Google Scholar] [CrossRef] [PubMed]
- Sehgal, D.; Rajaram, V.; Armstead, I.; Vadez, V.; Yadav, Y.P.; Hash, C.T.; Yadav, R. Integration of gene-based markers in a pearl millet genetic map for identification of candidate genes underlying drought tolerance quantitative trait loci. BMC Plant. Biol. 2012, 12, 9. [Google Scholar] [CrossRef]
- Singh, R.B.; Singh, B.; Singh, R.K. Cross-taxon transferability of sugarcane expressed sequence tags derived microsatellite (EST-SSR) markers across the related cereal grasses. J. Plant Biochem. Biotechnol. 2019, 28, 176–188. [Google Scholar] [CrossRef]
- Gurung, D.B.; George, M.L.C.; DeLaCruz, Q.D. Determination of Heterotic Groups in Nepalese Yellow Maize Populations. Nepal J. Sci. Technol. 2009, 10, 1–8. [Google Scholar] [CrossRef]
- Singh, S.; Gupta, S.K. Formation of heterotic pools and understanding relationship between molecular divergence and heterosis in pearl millet [Pennisetum glaucum (L.) R. Br.]. PLoS ONE 2019, 14, e0207463. [Google Scholar] [CrossRef]
- Teklewold, A.; Becker, H.C. Comparison of phenotypic and molecular distances to predict heterosis and F1 performance in Ethiopian mustard (Brassica carinata A. Braun). Theor. Appl. Genet. 2005, 112, 752–759. [Google Scholar] [CrossRef]
- 168. Basavaraj, G; Rao, P.P; Bhagavatula, S; Ahmed, W. Availability and utilization of pearl millet in India. SAT eJournal 2010, 8, 1–6.
- Singh, A.M.; Rana, M.K.; Singh, S.; Kumar, S.; Kumar, D.; Arya, L. Assessment of genetic diversity among pearl millet [Pennisetum glaucum (L) R Br] cultivars using SSR markers. Range Manag. Agrofor. 2013, 34, 77–81. [Google Scholar]
- Singh, S.; Gupta, S.K.; Thudi, M.; Das, R.R.; Vemula, A.; Garg, V.; Varshney, R.K.; Rathore, A.; Pahuja, S.K.; Yadav, D.V. Genetic Diversity Patterns and Heterosis Prediction Based on SSRs and SNPs in Hybrid Parents of Pearl Millet. Crop. Sci. 2018, 58, 2379–2390. [Google Scholar] [CrossRef]
- Bhardwaj, R.; Garg, T.; Malik, E.A.; Vikal, Y.; Sohu, R.S.; Gupta, S.K. Genetic divergence studies in pearl millet (Pennisetum glaucum L. (R.) Br.) inbred lines. Indian J. Genet. Plant. Breed. 2018, 78, 382–385. [Google Scholar] [CrossRef]
- Kapadia, V.N. Estimation of Heterosis for Yield and Its Relevant Traits in Forage Pearl Millet [Pennisetum glaucum LR Br.]. Int. J. Agric. Sci. 2016, 8. [Google Scholar] [CrossRef]
- Satyavathi, C.T.; Tiwari, S.; Bharadwaj, C.; Rao, A.R.; Bhat, J.; Singh, S.P. Genetic diversity analysis in a novel set of restorer lines of pearl millet [Pennisetum glaucum (L.) R. Br] using SSR markers. Vegetos 2013, 26, 72–82. [Google Scholar]
- Sumanth, M.; Sumathi, P.; Vinodhana, N.K.; Sathya, M. Assessment of genetic distance among the inbred lines of pearl millet (Pennisetum glaucum (L.) R. Br.) using SSR markers. Int. J. Biotechnol. Allied Fields 2013, 1, 153–162. [Google Scholar]
- Stich, B.; Haussmann, B.I.; Pasam, R.K.; Bhosale, S.; Hash, C.T.; Melchinger, A.E.; Parzies, H.K. Patterns of molecular and phenotypic diversity in pearl millet [Pennisetum glaucum (L.) R. Br.] from West and Central Africa and their relation to geographical and environmental parameters. BMC Plant. Biol. 2010, 10, 216. [Google Scholar] [CrossRef]
- Spindel, J.; Begum, H.; Akdemir, D.; Virk, P.; Collard, B.; Redona, E.; Atlin, G.; Jannink, J.L.; McCouch, S.R. Genomic selection and association mapping in rice (Oryza sativa): Effect of trait genetic architecture, training population composition, marker number and statistical model on accuracy of rice genomic selection in elite, tropical rice breeding lines. PLoS Genet. 2015, 11, e1004982. [Google Scholar]
- Zhong, S.; Dekkers, J.C.M.; Fernando, R.L.; Jannink, J.-L. Factors Affecting Accuracy From Genomic Selection in Populations Derived From Multiple Inbred Lines: A Barley Case Study. Genetics 2009, 182, 355–364. [Google Scholar] [CrossRef]
- Heffner, E.L.; Sorrells, M.E.; Jannink, J. Genomic Selection for Crop Improvement. Crop. Sci. 2009, 49, 1–12. [Google Scholar] [CrossRef]
- Crossa, J.; Campos, G.D.L.; Pérez-Rodríguez, P.; Gianola, D.; Burgueño, J.; Araus, J.L.; Makumbi, D.; Singh, R.P.; Dreisigacker, S.; Yan, J.; et al. Prediction of Genetic Values of Quantitative Traits in Plant Breeding Using Pedigree and Molecular Markers. Genetics 2010, 186, 713–724. [Google Scholar] [CrossRef] [PubMed]
- Poland, J.; Endelman, J.B.; Dawson, J.; Rutkoski, J.; Wu, S.; Manès, Y.; Dreisigacker, S.; Crossa, J.; Sanchez-Villeda, H.; Sorrells, M.; et al. Genomic Selection in Wheat Breeding using Genotyping-by-Sequencing. Plant. Genome 2012, 5, 103–113. [Google Scholar] [CrossRef]
- Ornella, L.; Singh, S.; Pérez-Rodríguez, P.; Burgueño, J.; Singh, R.; Tapia, E.; Bhavani, S.; Dreisigacker, S.; Braun, H.-J.; Mathews, K.; et al. Genomic Prediction of Genetic Values for Resistance to Wheat Rusts. Plant. Genome 2012, 5, 136–148. [Google Scholar] [CrossRef]
- Muleta, K.T.; Pressoir, G.; Morris, G.P. Optimizing Genomic Selection for a Sorghum Breeding Program in Haiti: A Simulation Study. G3 Genes|Genomes|Genet. 2018, 9, 391–401. [Google Scholar] [CrossRef] [PubMed]
- Meuwissen, T.H.; Hayes, B.J.; E Goddard, M. Prediction of total genetic value using genome-wide dense marker maps. Genetics 2001, 157, 1819–1829. [Google Scholar] [PubMed]
- Lorenz, A.J.; Smith, K.; Jannink, J.-L. Potential and Optimization of Genomic Selection for Fusarium Head Blight Resistance in Six-Row Barley. Crop. Sci. 2012, 52, 1609–1621. [Google Scholar] [CrossRef]
- Heffner, E.L.; Lorenz, A.J.; Jannink, J.; Sorrells, M.E. Plant Breeding with Genomic Selection: Gain per Unit Time and Cost. Crop. Sci. 2010, 50, 1681–1690. [Google Scholar] [CrossRef]
- Crossa, J.; Pérez-Rodríguez, P.; Cuevas, J.; Montesinos-Lopez, O.A.; Jarquin, D.; Campos, G.D.L.; Burgueño, J.; González-Camacho, J.M.; Elizalde, S.P.; Beyene, Y.; et al. Genomic Selection in Plant Breeding: Methods, Models, and Perspectives. Trends Plant Sci. 2017, 22, 961–975. [Google Scholar] [CrossRef]
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
References
- Rakesh K. Srivastava; Srikanth Bollam; Vijayalakshmi Pujarula; Madhu Pusuluri; Ram Singh; Gopi Potupureddi; Rajeev Gupta; Exploitation of Heterosis in Pearl Millet: A Review. MDPI PLANTS 2020, 9, 807, 10.3390/plants9070807.
