Your browser does not fully support modern features. Please upgrade for a smoother experience.
Please note this is a comparison between Version 1 by Usama AlDallal and Version 4 by Dean Liu.
Nanoparticle-based biosensor prototypes illustrate desirable diagnostics qualities, especially during the current pandemic. Such qualities include their ease of use, inexpensive equipment and visual results [77]. Gold nanoparticles (AuNPs) have fostered widespread attention in dthe past decade due to their outstanding optical characteristics, photostability and high extinction coefficient [77].
- nanoparticles
- coronaviruses
- diagnostic techniques
- COVID-19
1. Usage of Nanoparticles in the Detection of Middle East Respiratory Syndrome Coronavirus (MERS-CoV)
MERS-CoV was first reported in Saudi Arabia in September 2012, with further investigations suggesting that the first cases occurred in Jordan in April 2012 [1][36]. Like COVID-19, patients developed severe respiratory illness accompanied by fever, cough and shortness of breath. The mortality rate was approximated 30–40% [1][36].
Over the last few years, major strides have been made in MERS-CoV’s rapid and accurate detection using nanoparticles, especially AuNPs. AuNPs have been implemented both as the sole mechanism of detection or merged with established methods to improve detection rates. Through a competitive electromechanical immunosensor, Layqah and Eissa [2][79] used recombinant spike protein S1 (from spiked nasal samples) as a biomarker for MERS-CoV. The competition that occurred on the device was between the free viruses in the sample and the immobilized MERS-CoV S1 protein in the presence of known concentrations of antibodies [2][79]. The immunosensor was based on an array of carbon electrodes modified with an AuNP coating, with the voltametric response being the signal for detection. AuNPs were employed to increase the surface area, the transfer rate of the electrode’s electrons and ultimately, the peak current of the cathode [2][79]. The immunosensor was able to complete the analysis in 20 min while maintaining a LOD of 1.0 pg/mL and a high level of selectivity for MERS-CoV. Compared to the conventional methods used during that period, the immunosensor’s LOD was lower than the reported ELISA’s LOD of 1 ng/mL; the authors attributed the superior sensitivity to the AuNPs-modified electrodes [2][79].
Furthermore, AuNPs have been incorporated in colorimetric hybridization assays. The disulfide bond-based colorimetric assay developed by Kim et al. [3][80] for MERS-CoV is an example of one such assay. Using an extended form of “double-stranded DNA (dsDNA) self-assembly shielded” AuNPs, they could detect the presence of MERS-CoV viral molecules. The mechanism of detection involved color changes and the shift of a localized surface plasmon resonance (LSPR) in the visible range [3][80] (shown in Figure 13). The thiolated single-stranded DNA probes would identify specific regions of the MERS-CoV genome, genes upstream of the E protein gene and Open Reading Frame 1a to form a self-assembled complex. The complex would protect the nanoparticles from salt-induced clumping. Thereby, in the absence of the MERS-CoV targets, the AuNPs would aggregate together, resulting in a color change. The assay was reported to provide a result within 10 min; the naked eye would easily see the result. Further, with a potential detection limit of 1 pmol/µL, it can detect the coronavirus with little-to-no amplification [3][80]. Such assay would be of great use in high-demand and resource-limited sites.
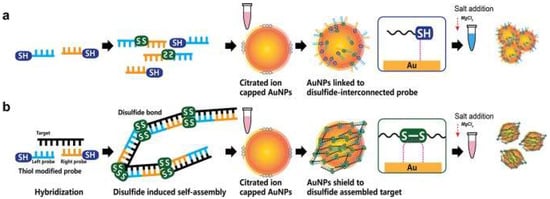
To accelerate detection and avoid the requirement of specialized facilities, flow detection strips utilizing NPs have been developed. By using a combination of vertical flow (VF) detection and reverse transcription loop-mediated isothermal amplification (RT-LAMP-VF), the N gene of MERS-CoV was visually detected [4][81]. MERS-CoV RNA was amplified and the resultant amplicons were marked with biotin and fluorescein isothiocyanate (FITC), allowing them to bind streptavidin-AuNPs conjugates and form complexes. This complex would then be targeted by anti-FITC antibodies, producing a visible colored line [4][81]. Considering preparation time and the time required to present a result, the test was reported only to take 35 min. The RT-LAMP-VF assay was able to detect 1 × 101 copies/µL of MERS-CoV RNA, which was more sensitive than the traditional RT-LAMP (1 × 102 copies/µL) but less sensitive than RT-PCR (1 × 100 copies/µL) [4][81]. The assay also displayed a high degree of specificity when evaluated with other coronaviruses, including SARS-related (SARSr)-CoV, HKU4, HKU1, OC43 and 229E [4][81]. The assay’s rapidity, good sensitivity/selectivity and, most importantly, lack of any need for expensive equipment make it a promising option that can be adapted in various laboratory settings.
Like AuNPs, silver nanoparticles (AgNPs) have been explored in the detection of MERS-CoV. Teengam et al. [5][82] developed a multiplex paper-based colorimetric sensor to detect infection-associated DNA using AgNPs as the colorimetric agent. The analytical device was tested to screen MERS-CoV and other microbes such as Mycobacterium tuberculosis and human papillomavirus. Using pyrrolidinyl peptide nucleic acid probes as opposed to DNA or RNA probes, the team measured the color change associated with AgNPs dispersion, giving a detection limit of 1.53 nM for MERS-CoV [5][82]. Moreover, the probe displayed high selectivity for the desired oligonucleotides, even when introduced to single-base mismatch, two-base mismatch and non-complementary DNA targets [5][82]. The multiplex was also described as capable of providing accurate detection for simultaneous testing of multiple DNA targets within a single device, simplifying analysis when compared to conventional techniques [5][82].
The use of NPs in detecting MERS-CoV is not limited to metal NPs such as AuNPs and AgNPs. Indeed, NPs like bacteria-based nanobioparticles have shown promise in the field of diagnosis. Through the biosynthesis of NPs (MERS nucleoprotein) inside Staphylococcus Aureus cells, Qioa et al. [6][83] were able to detect viral antibodies against MERS-CoV. MERS nucleoproteins were fabricated using a cell wall binding domain (CBD) from a bacteriophage lysin PlyV12 [6][83]. An agglutination test was performed to detect IgG using the Staphylococcus Aureus nanobioparticles. The agglutination was credited to the reaction between the IgG antibodies of MERS nucleoprotein, protein A on the surface of a staphylococcal particle and the MERS nucleoprotein-CBD adhered to another particle [6][83]. The specificity of this approach was tested by introducing the sensitized nanobioparticle to 38 clinical sera. No agglutination was observed in all 38 clinical sera and the nanobioparticle was deemed to have appropriate specificity. As the approach can provide a result within 20 min, this novel incorporation of NPs can provide an avenue for developing simple agglutination tests without unique instrumentation or expertise [6][83].
2. Usage of Nanoparticles in the Detection of Severe Acute Respiratory Syndrome-Associated Coronavirus (SARS-CoV)
Identified in 2003 in China, SARS-CoV was responsible for the 2002–2003 epidemic that infected 8000 people in 26 countries. It led to the death of 774 people [1][36]. A patient with SARS typically presents with fever, dry cough and diarrhea within the first or second week of infection. Often, the disease would progress and a patient’s condition would worsen rapidly, resulting in respiratory distress and intensive care management [1][36].
With the wide range of applicability and the unique optical qualities AuNPs offer, their use in SARS-CoV detection has been previously studied. Park et al. [7][84] effectively utilized AuNPs’ optical properties to detect SARS-CoV. Using a gold-binding polypeptide (GBP) as a fusion agent, the SARS-CoV E protein was immobilized on AuNPs. These AuNPs would then be modified with a green fluorescent protein and adhered to a bare gold surface to generate a nanopattern. The AuNP-E protein complex would demonstrate changes in both absorbance and color when exposed to its complementary antibody. These changes were recorded and subsequently used to determine the presence of SARS-CoV. The authors elaborate on how such a mechanism can create effective and high-throughput bioassays when combined with surface plasma resonance and microfluidic channels [7][84]. AuNPs’ characteristics were also exploited by Huang et al. [8][85], where they were utilized in a localized surface plasmon coupled fluorescence (LSPCF) fiber-optic biosensor. The biosensor’s mechanism of detection involved monitoring the fluorescence of a fluorophore-labeled anti-SARS-CoV N protein [8][85]. The device exhibited a detection sensitivity of 1 pg/mL for N protein in human serum. A linear response was demonstrated between the fluorescence signal and concentration of N protein from 0.1 pg/mL to 1 ng/mL. When compared with the conventional ELISA, the detection limit of the LSPCF fiber-optic biosensor was shown to be increased by 104-fold. The device’s very high sensitivity, quantitative ability, ease of use and disposability make it a desirable option for the early diagnosis of SARS infection [8][85].
NP-based hybridization assays for the detection of SARS-CoV have also been researched. An electrochemical hybridization assay based on AuNP has been described using a gene-based biosensor [9][86]. The gene-sensor (illustrated in Figure 24) consists of immobilized oligonucleotide probes on disposable AuNP-modified screen-printed carbon electrodes; the probes hybridize the biotinylated target DNA of SARS-CoV [9][86]. The catalyzation of the reduction of silver (Ag) ions and subsequent deposition of metallic Ag on the electrode’s surface would follow. The amount of metallic Ag can be quantified, which is directly proportional to viral DNA load, and a result can be reported within 20 min. The sensor’s selectivity was assessed by measuring analytical signals when hybridization was conducted using complementary strands 1, 2 and 3-base mismatch. Under 25% formamide, the sensor was able to discriminate between the complementary strands and the mismatched strands. Furthermore, a linear sensor response was reported between 2.5 to 50 pmol/L, suggesting that 2.5 pmol/L was the sensor’s detection limit. The sensor’s sensitivity was also described to be greater than that achieved by surfaces without NPs [9][86].
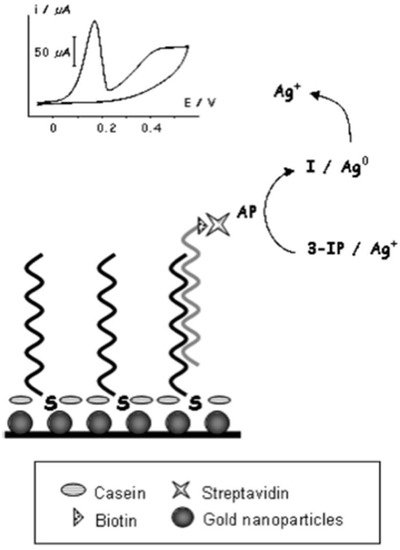
Figure 24. Basic schematic representation of the genosensor device developed by Martinez-Paredes et al. [9].
Carbon nanotube (CNT) field-effect transistors and biosensors hold great potential in diagnostics owing to their mechanical durability and high thermal conductivity. In addition, single-walled CNTs have the smallest diameter compared to other one-dimensional nanomaterials, making them a remarkable option in terms of sensors, as every atom in the CNTs is in contact with its environment [10][87]. To study the relationship between CNT density and sensitivity, Ishikawa et al. [10][87] tested their density-optimized CNT biosensors to detect the N protein of SARS-CoV under physiological conditions. The device successfully detected the biomarker protein with a detection limit of 5 nM [10][87].
NPs, such as superparamagnetic NPs, can be used to capture targeted viral genetic material from a mixture of numerous genetic molecules for amplification. To this end, Gong et al. [11][88] designed functionalized silica-coated superparamagnetic NPs to isolate the desired cDNA of SARS-CoV from a sample that contains the target cDNA and non-target cDNA. The isolated cDNA was then amplified through a general symmetry PCR. Next, the cDNA was once again separated from the PCR products using another batch of the same NPs [11][88]. Lastly, through fluorescence, the functionalized NPs were to be used as signaling probes with a sandwich hybridization method. The approach was able to detect the cDNA with a detection limit of 2.0 × 103 within 6 h [11][88].
3. Usage of Nanoparticles in the Detection of Severe Acute Respiratory Syndrome-Associated Coronavirus-2 (SARS-CoV-2)
As the cause behind the COVID-19 pandemic of 2020, the rapid and accurate detection of SARS-CoV-2 has been a priority for researchers globally. While RT-PCR and CT have been the most used diagnostic methods thus far, NP-based diagnostic techniques are being actively developed and optimized to provide utility in the clinical setting.
Similar to how AuNPs found application in detecting MERS-CoV and SARS-CoV, it was only inevitable that they would also be used to detect SARS-CoV-2. Many of the current diagnostic tests are unreliable when tested against a viral load at its early stages of infection or a viral gene that has mutated during its spread. To this end, Moitra et al. [12][77] have attempted to overcome such challenges by designing an AuNP-based colorimetric assay that can directly target the SARS-CoV-2 N gene at multiple gene positions simultaneously. The AuNPs used in this biosensor are labeled with thiol-modified antisense oligonucleotides (ASOs) that specifically target the N gene of SARS-CoV-2 [12][77]. In the presence of the N gene, these AuNPs would aggregate, displaying a change in their surface plasmon resonance (SPR). This agglomeration is further enhanced by the addition of RNaseH, resulting in the precipitation of the AuNPs, which are visible to the naked eye [12][77]. The test’s limit of detection was reported to be 0.18 ng/µL. Additionally, the selectivity was also assessed by introducing the Au-ASO nanoparticles to RNA isolated from cell lysate of MERS-CoV infected Vero cells, where there was no significant change in the absorbance, making the test rather selective.
Similarly, Kumar et al. [13][89] developed a colorimetric nanoparticle-based assay to detect the presence of the RdRp gene of SARS-CoV-2. Through the usage of AuNPs, a pink-to-blue color change can be appreciated upon the formation of the oligo probe-target complex within assays containing SARS-CoV-2 pharyngeal RNA samples. The assay would maintain its pink color when introduced to samples from COVID-19-negative patients or human papillomavirus-infected women. When tested against RT-PCR, the developed instrument had a sensitivity of 85.29% and a specificity of 94.12%. Lastly, the detection limit was 0.5 ng of SARS-CoV-2 RNA and results can be obtained within 30 min [13][89]. As such, these studies exemplify how NPs can be utilized to develop a visual, reliable and reproducible COVID-19 diagnostic test without the need for sophisticated equipment.
AuNPs have also shown promise in providing extremely rapid detection (≤2 min) of SARS-CoV-2 RNA, bypassing the need for PCR amplification. Li et al. [14][90] designed a graphene field-effect transistor sensor studded with AuNPs, on which complementary phospho-rodiamidate morpholino oligos probes were seeded to target the RdRp gene. Through the usage of a semiconductor characterization system and a probe station, electrical changes on the sensor chip can be monitored. When tested on 30 clinical samples (20 confirmed COVID-19 patients and 10 healthy controls), a statistically significant difference in the average levels of the sensor’s signal was reported. The limit of detection was determined to be 2.29 fM for throat swabs and 3.99 fM for serum. Furthermore, a Kappa test was performed to scrutinize the reliability of the developed method and the gold-standard RT-PCR, with a result of 0.92, suggesting near-perfect agreeability. Such a tool could be adopted in the emergency setting or close-contact investigations as a screening tool, given its ability to provide rapid and reliable results [14][90].
By combining the plasmonic photothermal (PPT) effect and localized surface plasmon resonance (LSPR) sensing transduction, Qiu et al. [15][91] were able to create a dual-functional plasmonic biosensor based around gold nano-islands (AuNIs) to detect SARS-CoV-2 genetic material. These AuNIs were functionalized by forming bonds with the complementary receptors of the ORF1ab, RdPd, or E gene sequences, allowing for their sensitive detection through nucleic acid hybridization [15][91]. To maximize the device’s sensitivity, heat was generated by exposing the AuNIs to their plasmonic resonance frequency. Sensing stability and reliability were further improved by utilizing two different angles of incidence, permitting the use of two varying wavelengths to excite the plasmonic resonances of PPT and LSPR [15][91]. Consequently, the biosensor’s detection limit was reported to be 0.22 pM while simultaneously differentiating the desired targets from a multigene mixture [15][91]. This technique could enable clinicians to generate a reliable result in real-time without labeling the desired gene sequences.
Along with the detection of viral DNA/RNA, NPs have been developed to detect anti-SARS-CoV-2 IgG in human serum. In the lateral flow immunoassay (LFIA) created by Chen et al. [16][92], lanthanide-doped polystyrene nanoparticles (LNPs) were used to detect anti-SARS-CoV-2 IgG; mouse anti-human IgG marked with LNPs functioned as a fluorescent signaler (portrayed in Figure 35). The assay was tested with 100-µL aliquots of human serum samples that were diluted 1000-fold; the test’s duration was reported to only take 10 min [16][92]. Moreover, the diagnostic accuracy of the assay was evaluated by testing it against samples that were previously tested with RT-PCT, seven of which were positive and 12 were negative but clinically suspicious [16][92]. The assay was able to accurately replicate the results of the RT-PCT, giving a positive result for all seven samples. In fact, the assay also reported a positive result in one of the clinically suspicious samples. Unlike the other suspected samples, this specific case presented with relevant symptoms in addition to fever, justifying the high clinical suspicion [16][92]. Chen et al. showcased how an assay based on NPs can be used to ascertain the results provided by RT-PCT, provide a rapid diagnosis and also be applicable in monitoring a patient’s response to treatment.
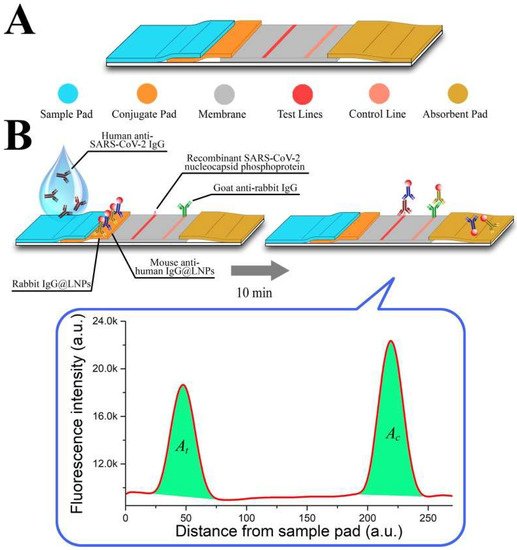
Figure 35.
(
A
) An illustration of the developed lateral flow strip; (
Immunoassays revolving around AuNPs were also developed for the detection of SARS-CoV-2. Wen et al. [17][71] described the creation of a point-of-care LFIA used to detect the IgG antibodies against SARS-CoV-2. The strip was compromised of an immobilized SARS-CoV-2 nucleocapsid protein on the strip’s surface and anti-human IgG coupled with colloidal AuNPs. Colloidal AuNPs were mainly chosen for this sensor due to their stability, ability to provide interpretable results with the naked eye and excellent biocompatibility [17][71]. The detection mechanism is based on antigen-antibody interactions, where the presence of IgG antibodies of SARS-CoV-2 would form a complex with the anti-human-antibody-AuNP compound and subsequently migrate to the bound nucleocapsid protein (test line). Thereby, a positive result would be indicated based on whether there is a red color on the test line. In order to gauge the practicability of the assay in a clinical setting, Wen et al. applied the assay to detect SARS-CoV-2 in 55 different patient samples. The reported sensitivity was 69.1%, where 17 out of the 55 samples were not identified as positive. Nonetheless, when tested against samples from patients with severe fever with thrombocytopenia syndrome (SFTS) and avian influenza A (H7N9), there was no cross-reactivity signifying that the assay was of high specificity [17][71]. Each sample was repeated in triplets at intervals ranging from 1 to 3 weeks; the results remained unchanged, suggesting that the LFIA strip had exceptional stability. Due to its speed and cost-effectiveness, the authors propose that the LFIA strip can provide a preliminary result for physicians to initiate treatment in clinically suspicious patients. However, given its limitations in terms of sensitivity, physicians may need to use alternative testing methods to make a confident diagnosis. Similarly, Huang et al. [18][93] synthesized a colloidal gold nanoparticle-based LFIA that allows for the on-site detection of IgM antibodies against SARS-CoV-2. Detection is interpreted through an indirect immunochromatography technique. The strips were comprised of a SARS-CoV-2 nucleoprotein-coated membrane and an anti-human IgM-AuNP complex functioning as the reporting agent. The result can easily be seen as a red-colored line appearing on the test line and control lines. By testing it against serum samples of COVID-19 patients and normal individuals, the assay achieved a 100% accuracy [18][93]. It also demonstrated a degree of selectivity, showing no cross-reactions or alterations when tested against severe fever with thrombocytopenia syndrome virus and dengue virus [18][93]. Considering that the test requires 15 min and 10–20µL of serum to perform, it presents itself as a very attractive and feasible method to detect SARS-CoV-2.
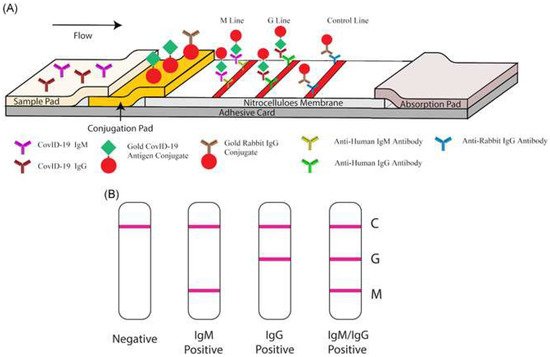
Interestingly, in an attempt to identify patients at varying infection stages, Li et al. [19][94] developed a point-of-care lateral flow immunoassay to detect both IgM and IgG antibodies against SARS-CoV-2 within 15 min, where the antigen is conjugated on AuNPs. Similar to the previous devices, results are communicated through detection lines on the strip. However, instead of two, three lines are used: a control (C) line, an anti-SARS-CoV-2 IgM line and an anti-SARS-CoV-2-IgG line (showcased in Figure 46). The device’s sensitivity and specificity were 88.66% and 90.63%, respectively, calculated using blood samples from 397 PCR confirmed COVID-19 patients and 128 COVID-19 negative patients. Moreover, the results were deemed reliable across sample types, whether they may be fingerstick blood or plasma of venous blood [19][94].
Other forms of nanotechnology, including graphene sheets, magnetic nanoparticles (MNPs) and nanotubular formations of conducting polymers (CPs), have been used to develop novel approaches for the detection of SARS-CoV-2. Seo et al. have developed an antibody-based biosensor functioning as a field-effect transistor (FET) to detect SARS-CoV-2 in nasopharyngeal samples [20][95]. The graphene sheets were coated with an antibody targeting the SARS-CoV-2 spike protein. With this approach, the device detected the protein in phosphate-buffered saline and clinical transport medium at concentrations of 1 fg/mL and 100 fg/mL, respectively [20][95]. When used in unprocessed clinical samples, the sensor detected the virus with a detection limit of 242 copies/mL, which could potentially be lowered by reducing noise signals. Compared to the detection limits of currently used molecular diagnostic techniques (50–100 copies/mL), the graphene-based biosensor can provide a comparable diagnostic outcome within significantly shorter periods of time [20][95]. Another form of applicable nanotechnology are MNPs, one of the most promising methods for sensitive biomolecule detection [21][96]. As an example, Zhong et al. formulated an approach to detect SARS-CoV-2 through the utilization of functionalized MNPs and the measuring of their magnetic response in an AC magnetic field [21][96]. As a proof of concept, the team fabricated SARS-CoV-2 replicas; each replicate is composed of 100 biotinylated spike proteins and a streptavidin-coated polystyrene bead. Using magnetic particle spectroscopy (MPS), it was seen that the binding of mimic SARS-CoV-2 particles and functionalized MNPs changed the MPS signal as it increased the effective Brownian relaxation time (illustrated in Figure 57). With a calculated detection limit of 0.084 nM (5.9 fM), the authors believe that the approach could be a promising candidate for sensitive, easy-to-perform and cheap point-of-care for rapid SARS-CoV-2 detection [21][96]. Lastly, nanoarchitectures of CPs, such as poly-pyrrole have been investigated for COVID-19 diagnostic applications due to their stability, ease of synthesis and electroactivity in phosphate buffer (pH 7.4) [22][97]. By incorporating AuNPs within poly-pyrrole nanotubular morphologies, Hryniewicz et al. added further stability, biocompatibility and selectivity to the CPs formulations, making it a promising biosensing tool for COVID-19 antibodies [22][97]. The authors tested two morphologies (globular and nanotubular) of poly-pyrrole implanted with AuNPs against SARS-CoV-2 Nucleocapsid (N) protein. The detection limits for the globular and nanotubular formulations were 7.442 and 0.4 ng/mL, respectively. As such, when adopting a nano-scale form, the nanotubular morphology showcased 8-fold higher sensitivity than its globular counterpart [22][97]. Furthermore, the nanotubular biosensor was tested against serum from positive and negative COVID-19 patients, where it successfully distinguished between all tested samples within 1 h. Given the small volume of serum needed to obtain a result, finger-prick blood tests for COVID-19 diagnosis and immunosurveillance could be a promising translation of this technology in a clinical setting [22][97].
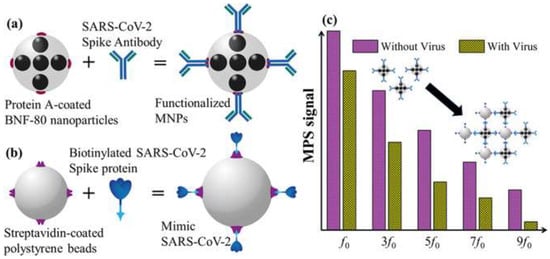
Figure 57. (a) Illustration of functionalized magnetic NPs; (b) illustration of SARS-CoV-2 mimic; (c) magnetic particle spectroscopy signal with and without SARS-CoV-2 mimic. Reprinted (adapted) with permission from Zhong et al., Toward Rapid and Sensitive Detection of SARS-CoV-2 with Functionalized Magnetic Nanoparticles. ACS Sens. 2021, 6, 976–984, doi:10.1021/acssensors.0c02160. Copyright 2021 American Chemical Society.
With the rising rates of SARS-CoV-2 mutants, nanoparticle-based methods may provide novel detection methods or improve upon current methods to maintain high sensitivity and selectivity rates. Through an immunosensor utilizing MNPs conjugated to a mixture of antibodies, Durmus at al. were able to obtain higher detection rates of SARS-CoV-2 variants when compared to commercially available anti-SARS-CoV-2 S1 (anti-S1) and anti-S2 monoclonal antibodies [23][98]. The device’s LOD ranged from 0.53–0.75 ng/mL as opposed to the anti-S1 LOD of 0.93 ng/mL and the anti-S2 LOD of 0.99 ng/mL. The performance of the three immunosensor platforms were tested using 50 RT-PCR confirmed clinical samples (nasopharyngeal swabs; 40 positive and 10 negative). The positive results were further sub-classified according to the virus’s variant: original, alpha, beta and delta. The MNP immunosensor had a sensitivity and selectivity of 100% while the anti-S1 and anti-S2 platforms had sensitivity/selectivity values of 90%/100% and 87.5%/100%, respectively. To this end, the developed device showcased better ability to separate positive and negative samples with higher rates of variant detection. Nonetheless, it should be stated that the mixture of antibodies is derived from human samples, a method that is relatively unsustainable and has limit potential of mass production [23][98]. An alternative method of variant detection could be aptamer-based biosensing, an approach successfully demonstrated by Ellipilli et al., where DNA aptamer targeting a variant’s S protein is conjugated to AuNPs. [24][99] The aptamer-AuNP complex is then used as a detection probe, subsequently exhibiting a red color change when binding pre-anchored spike-antibodies on a nitrocellulose membrane. The aforementioned experiment served as a proof of concept, where the authors hope that such a method could be adopted to rapidly (<5 min) detect variant SARS-CoV-2 strains by cloning respective antibodies and customizing aptamers for mutated spike proteins [24][99]. Further, nanoparticles have been shown to markedly expediate the process of quantitative PCR, allowing for point-of-care application. Using gold nanorods, Blumenfeld et al. demonstrated plasmonic thermocycling through irradiation of the nanoparticles to facilitate rapid heating of a sample [25][100]. The incorporation of gold nanorods led to quenching of the overall fluorescence signal. However, the advantages gained are arguably worth the loss in fluorescence: the sample-to-result time was reported to be <25 min while still maintaining an LOD of 2.2–4.4 copies/μL and sensitivity/specificity values of 100%/100% from either saliva or nasal samples [25][100].
In addition to the detection of genetic material, antigens and antibodies against SARS-CoV-2, nanoparticles have also been applied to detect reactive oxygen species (ROS) and volatile organic compounds (VOCs) attributable to COVID-19. Using an electrochemical sensor decorated with Multi-Wall Carbon Nanotubes (MWCNTs) on its electrodes, Miripour et al. [26][101] were able to detect ROS in sputum samples. In its three-electrode conformation (working, reference and counter electrodes), changes in electrode potential were monitored using cyclic voltammetry. When validated with results from CT scans and RT-PCR, the device’s sensitivity/specificity results were 92%/94% and 97%/91% when compared to healthy individuals, respectively. Indeed, 97% of true positive patients were identified within 30 s. However, it needs to be noted that other respiratory conditions, such as asthma, acute pneumonia, etc., can also increase ROS [26][101]. Therefore, the device should not be used as a definitive diagnostic tool but rather as a rapid, assistive screening technique. In a paper revolving around a similar concept, Shan et al. [27][102] describe their multiplexed nanomaterial-based sensor array used to detect COVID-19 patients using their breath, where each array consists of 8 AuNPs linked to organic ligands. As a COVID-19 patient breathes into the device, the VOCs would react with the AuNPs, resulting in several detectable electrical signals. Through a case-control study design involving 49 confirmed COVID-19 patients, 58 healthy individuals and 33 non-COVID lung infection controls, the device exhibited an accuracy of 94% (training data set: 70% of the total sample) and 76% (test data set: 30% of the sample) in distinguishing COVID-19 patients from controls. Further, it had an accuracy of 90% (training data set) and 95% (test data set) in distinguishing between COVID-19 patients and other lung infection patients [27][102].
