Insects and ectoparasites are causes for major concern throughout the world due to their economic and welfare impacts on livestock agriculture. Current control measures involve chemicals such as acaricides which pose challenges like chemical resistance and longer withholding periods. To enable more sustainable agriculture practices, it is important to develop technologies that combine targeted effectiveness with minimal environmental footprint. RNA interference (RNAi) is a eukaryotic process in which transcript expression is reduced in a sequence-specific manner. This makes it a perfect tool for developing efficient and effective biological control against pests and pathogens. Double-stranded RNA (dsRNA) is the key trigger molecule for inducing RNAi; this concept is widely studied for development of RNA-based biopesticides as an alternative to chemical controls in crop protection for targeting pests and pathogens with accuracy and specificity.
- RNA interference
- dsRNA delivery
- biological control
1. Introduction
2. RNA-Interference (RNAi)
RNAi is an endogenous, post-transcriptional gene regulation mechanism which has been identified in almost all eukaryotes; from plants, fungi, algae, protozoans, invertebrates, to vertebrates [18,19][9][10]. It was first reported by Napoli and Jorgensen in 1990 when they observed endogenous gene co-suppression while studying pigmented petunias [20,21][11][12]. In animals, RNAi was first documented in the nematode Caenorhabditis elegans [22,23][13][14]. While the intracellular components of RNAi are similar across species, it was initially known as quelling in fungi, and post transcriptional gene silencing in plants [24][15]. It is a highly conserved mechanism, which is highly sequence specific and selective in its activity [25][16].2.1. RNAi Mechanism
While a comprehensive review on RNAi mechanism discussing all the different facets of this technology is outside the scope of this revisewarch, the basic mechanism of RNAi when initiated via delivered exogenously applied dsRNA are well described and are illustrated in Figure 1.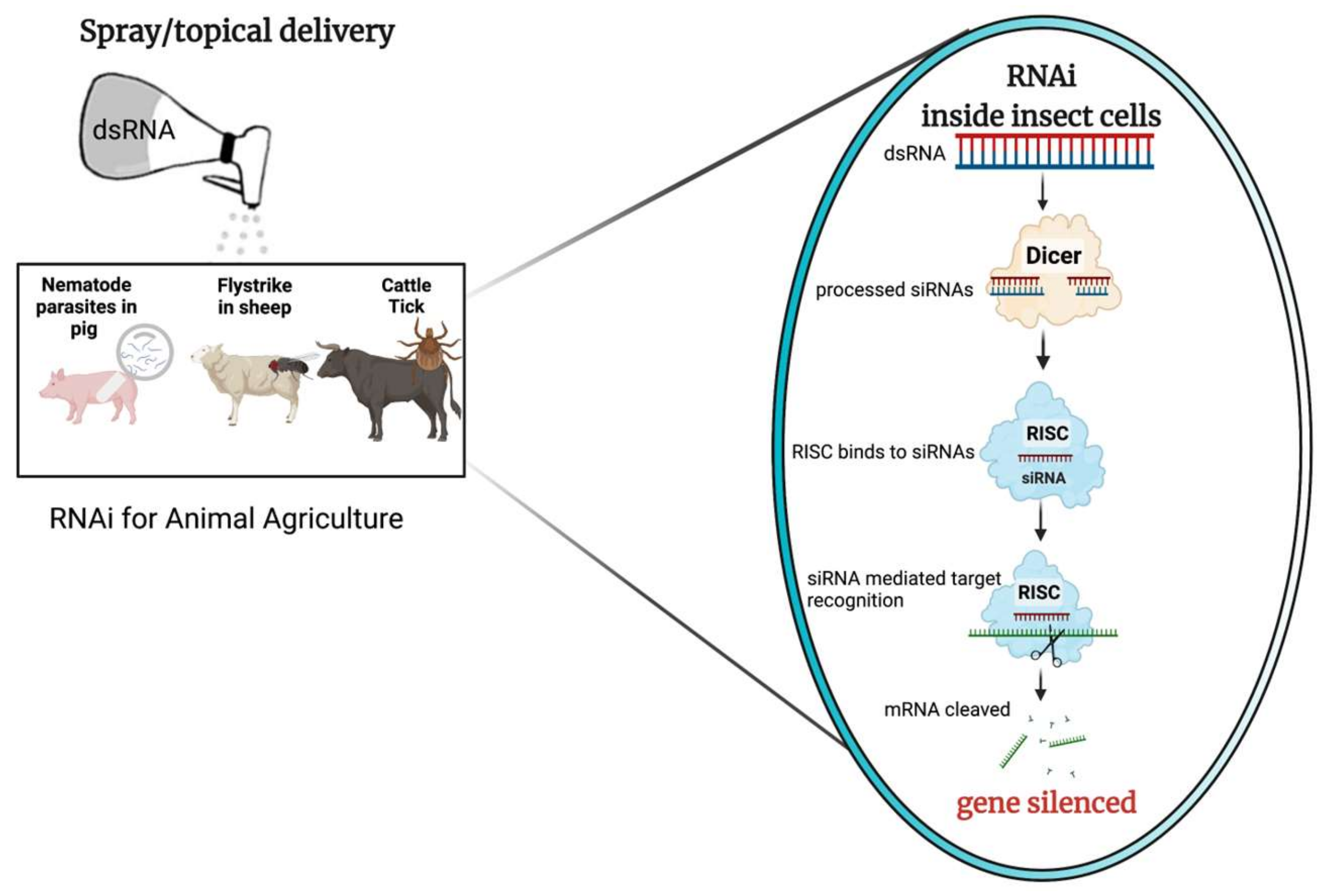
2.2. RNAi Targets
Target gene selection depends on the class of RNA effector molecule to be used. While both siRNA and piRNA are generated from long, complementary dsRNAs, miRNA is endogenous in nature, usually processed from stem-loops, and siRNA are exogenous and are directly generated from cleaved dsRNAs [30][21]. The first step in developing an RNAi product is to identify potential target genes and design dsRNA targeting them. In 2010, a research study suggested that there are five factors which play a key role in influencing the efficiency of RNAi as biocontrol, namely, the concentration, nucleotide sequence, length of the dsRNA, effective period of the dsRNA induced silencing, and life stage of the target [31][22]. The perfect RNAi target gene should be transcribed into an mRNA with a high turn overrate, that is translated into a protein with a short half-life, and is transcribed at all stages of life cycle, with the loss of function leading to mortality or severe impairment in the pest or pathogen of interest. In addition, the target transcript should be poorly conserved across species to maintain maximum specificity of mRNA for minimal environmental effects on non-target species [27][18]. This is a challenging criterion to meet, as those genes that are essential to cell viability tend to be more conserved between species. It has reported that the efficiency of dsRNA uptake is length dependent, requiring an optimal length and dosage to induce RNAi successfully [32][23]. Studies have shown that dsRNA as short as 30 and 60-bp can induce 30% and 70% gene knockdown in the red flour beetle (Tribolium castaneum), and the optimum length 200–520 bps for most insects [31,33][22][24]. This ideal dosage changes according to the organism’s susceptibility to RNAi and their potential for systemic RNAi, formulation for delivery, gene expression abundance, life and development stage for gene expression [27][18]. Unfortunately, genes that often would lead to mortality if targeted by RNAi are highly conserved across nature, due to their evident importance for homeostasis [27][18]. Various selection methods can be used to develop efficient RNAi targets to minimize side effects and enhance expected silencing. Since RNAi relies on high gene specificity to the selected target species based on sequence divergence, dsRNA needs to be screened and designed to be specific to a target gene or to target genes on a broad spectrum, closer to related species [34][25]. To increase knockdown or have higher than the expected RNAi result, multiple targets can be selected for a single transcript to ensure variation within the target species is accounted for. A common application that can be used to evaluate off-target genes is the Basic Local Alignment Search Tool (BLAST), and it has been used to find contiguous matching sections of 17 nt or more in the genomes of interesting organisms [34][25]. BLAST is a common approach for finding regions of identity, and for identifying the functionality of siRNA. Thereafter, a wide range of specialised programs can be used to design the sequences. These include siRNA-Finder (si-Fi), siDirect, dsCheck, and RNAi Designer by ThermoFisher [35][26]. Amongst insect researchers it is widely accepted that several dsRNA should be screened as a combination as each gene is highly unique to the target gene and the insect species [33][24]. Whyard et al. demonstrated species specific insecticides could be achieved by targeting the variable 3′-UTR regions of the y-Tubulin transcript, allowing for a specific species knockdown between four closely related Drosophila species [36][27]. The results of this study clearly demonstrate that with sufficient sequence information and thorough interrogation of this available data, the development of highly specific RNAi based insecticides is highly feasible. While the application of RNAi animal agriculture is still in its infancy, there are more dosage trials in plant agriculture, such as efficiency of dosage needed per acre regarding insect pests. Despite the fact that sufficient field experimentation is lacking, it is estimated that near 2–10 g of dsRNA is needed per acre of plants [30][21]. As this depends on the organisms response efficiency and target gene, this number is open to high levels of variability [30][21].2.3. dsRNA Uptake Mechanism
The use of RNAi to protect plants and animals by suppressing essential gene function in pest species has been well documented in model invertebrates such as Caenorhabditis elegans, and Drosophila melanogaster [37][28]. While most of the reported experiments involved injecting dsRNA into the target organism, this is not a viable delivery approach for in-field applications in animal production, thus creating the need for autonomous dsRNA uptake [31,37][22][28]. Studies have reported two dsRNA uptake mechanisms, the transmembrane channel-mediated uptake and an alternate endocytosis-mediated uptake [37][28]. Transmembrane channel-mediated uptake mechanism has been best studied in C. elegans, more specifically with worms carrying mutant sid genes that lack systemic RNAi capacity [31][22]. This pathway involves two proteins, SID-1 and SID-2, with changes being made to the protein molecule affecting other cells and their activity. SID-1 is a hypothesized transmembrane protein which passively transports dsRNA into the cells [31,38][22][29]. The role played by SID-2 has been hypothesized to help facilitate RNAi by coordinating with SID-1 either by modifying the molecule to activate it or binding to dsRNA for delivery or by inducing endocytosis [31][22]. It has been noted that SID-1 is considered to be a ‘cell surface conduit’ for dsRNA uptake [32][23]. Meanwhile, endocytosis-mediated mechanism was first studied in D. melanogaster, since it had no sid gene orthologues and the mechanism was later confirmed in C. elegans [31][22]. This is a cell-autonomous pathway wherein dsRNA is transported intracellularly via vesicles [31,38][22][29]. Drosophila S2 cells were able to take up dsRNA and initiate an RNAi response by receptor-mediated endocytosis uptake which is an active process involving vacuolar H+ (V-H) ATPase [31,32][22][23]. This pathway is active, specific and is conserved in other animal cells [32][23]. Some organisms can take up exogenous dsRNA and trigger an RNAi response in the entire animal systemically, like C. elegans, while others cannot transmit this RNAi response in another cell [32][23]. In Drosophila, it has been confirmed that transposon inversion can activate RNAi, and the homologous gene silencing caused by it is similar to co-inhibition in plants [39][30]. In a C. elegans assay, it was found that the quelling defective protein, QDE-1, a necessary protein for RNAi process, is homologous to RNA-dependent RNA polymerase (RDRP), supporting RNAi process potential involved in RNA replication and regulation [40][31]. Similarly, dsRNA injection into phloem of the plant can spread throughout the whole plant to initiate systemic RNAi [41][32]. Interestingly, RNAi can also be induced by immersing nematodes in dsRNA-containing liquids or by feeding nematodes with E. coli bacteria engineered to express dsRNA [42][33]. The pathway for entry of dsRNA into the target cell determines RNAi efficiency and provides a starting point for the development of dsRNA delivery systems [32][23].3. Application of RNAi for Animal Health
Over the past ten years, there has been a significant increase in published papers regarding application of RNAi to control in parasites affecting livestock, the data of which has been figuratively described in Figure 2A. This section highlights those RNAi studies which have been focused on the control of pests that affect pigs, sheep, and cattle pest.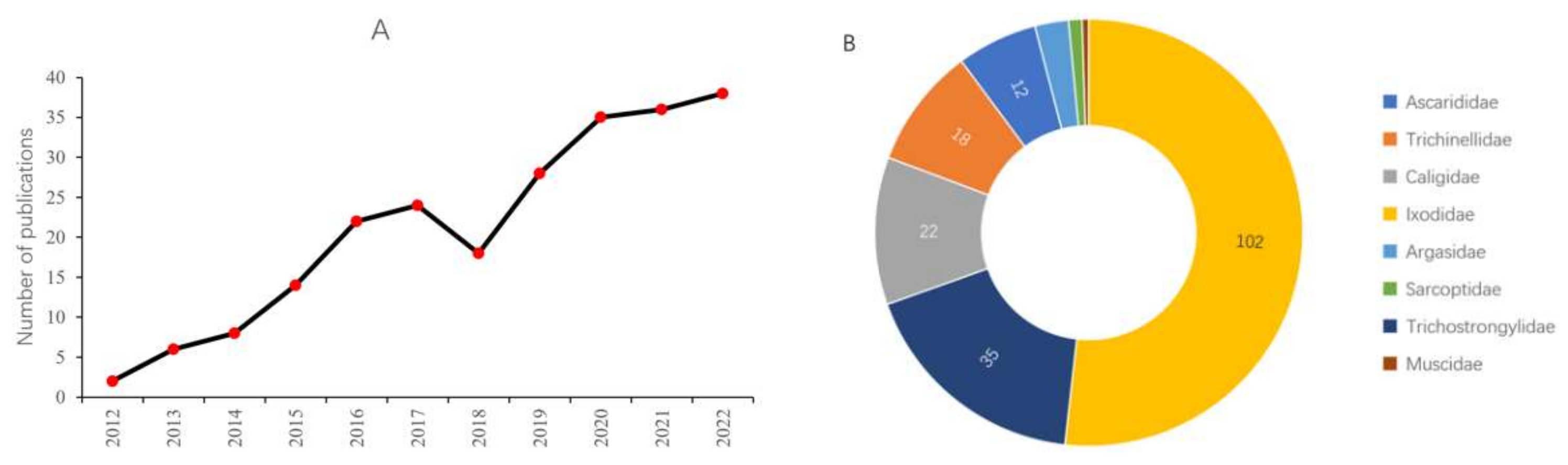
| RNAi Target | Parasites | Delivery | Livestock | Efficiency | Ref | |||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| As-eft-1, As-eft-2, As-gmpr, As-tnc-1, As-rab-3, As-hb-1, As-unc-29, As-unc-38 |
Ascaris suum | Soaking | Pig | Different targets in adult | A. suum | were tested for gene knockdown. All targets were significantly silenced in different region of adult | A. suum | . Furthermore, all genes indicate they are susceptible to RNAi in adults | A. suum | , especially neuronal genes. | [15] | [39] |
| As-enol-1 | Ascaris suum | Soaking | Pig | Enolase gene expression was significantly silenced, and the mortality rate increased by 20.11% ( | p | < 0.01) after soaking expressed sequence tag (EST) representing | As-enol-1 | dsRNA derived from | A. suum | for 72 h. | [65] | [34] |
| EST-06G09 | Ascaris suum | Soaking | Pig | The gene of EST 06G09 was silenced after 72 h soaking. The survival rate achieved highest peak at 48 h (20.37%, | p | < 0.01). The gene 06G09 has function in larval development. | [70] | [40] | ||||
| TsSPIs | Trichinella spiralis | Soaking | Pig | TsSPI mRNA and protein expression levels in larvae decreased by 68.7% and 68.4%, respectively ( | p | < 0.05). RNAi-mediated suppression of TsSPI expression in | T. spiralis | drastically decreased larval infectivity and survival inside the host. | [71] | [41] | ||
| LsalMS | Lepeophtheirus salmonis | Soaking | Salmon Lice | Myosuppressin (LsalMS) knockdown resulted in reduction in the amount of muscle, with skeletal and visceral muscles both showing anomalies in the lice. Additionally, LsalMS down-regulation also had an impact on feeding, spermatophore deposition, and moulting. | [72] | [42] | ||||||
| Ubiquitin-63E | Rhipicephalus microplus; | Injection | Cattle | All ticks treated with dsRNA did not have offspring. Ubiquitin-63E knockdown was confirmed by microarray and qRT-PCR. | [73] | [43] | ||||||
| HBP | Amblyomma mericanum | Injection | Cattle | dsRNA specific for histamine binding protein (HBP) transcript was injected into the haemolymph. Less expression of HBP mRNA was shown by molecular evidence, resulting in lower histamine binding ability. | [74] | [44] | ||||||
| Subolesin | Ornithodoros spp. | Injection | Cattle | By injecting dsRNA to silence the subolesin gene (Silencing 70–78%), oviposition was prevented (88.6% and 95.4%). | [75] | [45] | ||||||
| Fhteg1, Fhteg5, Fhteg8, | Fasciola hepatica | Soaking | Cattle | 92.9% transcript reduction of Fhteg1, 74.1% transcript reduction of Fhteg5. But no observable effect on phenotype. | [76] | [46] | ||||||
| SsGST-mu1 | Sarcoptes scabiei | Soaking | Sheep | Compared to controls, knockdown caused a 35% decrease in transcription of the target gene. | [77] | [47] | ||||||
| Hyaluronidase | Haemonchus contortus | Oral feeding | Sheep | A significant drop in worm burden and significant decrease in hyaluronidase activity in siRNA treated group. | [78] | [48] |
References
- FAO. The State of Food and Agriculture 2001; Food & Agriculture Organization: Rome, Italy, 2001.
- Savary, S.; Willocquet, L.; Pethybridge, S.J.; Esker, P.D.; McRoberts, N.; Nelson, A.D. The global burden of pathogens and pests on major food crops. Nat. Ecol. Evol. 2019, 3, 430–439.
- Narladkar, B. Projected economic losses due to vector and vector-borne parasitic diseases in livestock of India and its significance in implementing the concept of integrated practices for vector management. Vet. World 2018, 11, 151.
- Worrall, E.; Hamid, A.; Mody, K.; Mitter, N.; Pappu, H. Nanotechnology for Plant Disease Management. Agronomy 2018, 8, 285.
- Chagnon, M.; Kreutzweiser, D.; Mitchell, E.A.D.; Morrissey, C.A.; Noome, D.A.; Van der Sluijs, J.P. Risks of large-scale use of systemic insecticides to ecosystem functioning and services. Environ. Sci. Pollut. Res. Int. 2014, 22, 119–134.
- Bai, G.; Shaner, G. Management and resistance in wheat and barley to Fusarium head blight. Annu. Rev. Phytopathol. 2004, 42, 135–161.
- Kim, Y.H.; Issa, M.S.; Cooper, A.M.; Zhu, K.Y. RNA interference: Applications and advances in insect toxicology and insect pest management. Pestic. Biochem. Physiol. 2015, 120, 109–117.
- Jain, R.G.; Robinson, K.E.; Fletcher, S.J.; Mitter, N. RNAi-based functional genomics in Hemiptera. Insects 2020, 11, 557.
- Schuster, S.; Miesen, P.; van Rij, R.P. Antiviral RNAi in insects and mammals: Parallels and differences. Viruses 2019, 11, 448.
- Chen, J.; Peng, Y.; Zhang, H.; Wang, K.; Zhao, C.; Zhu, G.; Reddy Palli, S.; Han, Z. Off-target effects of RNAi correlate with the mismatch rate between dsRNA and non-target mRNA. RNA Biol. 2021, 18, 1747–1759.
- Napoli, C.; Lemieux, C.; Jorgensen, R. Introduction of a chimeric chalcone synthase gene into petunia results in reversible co-suppression of homologous genes in trans. Plant Cell 1990, 2, 279–289.
- Sen, G.L.; Blau, H.M. A brief history of RNAi: The silence of the genes. FASEB J. 2006, 20, 1293–1299.
- Guo, S.; Kemphues, K.J. par-1, a gene required for establishing polarity in C. elegans embryos, encodes a putative Ser/Thr kinase that is asymmetrically distributed. Cell 1995, 81, 611–620.
- Fire, A.; Xu, S.; Montgomery, M.K.; Kostas, S.A.; Driver, S.E.; Mello, C.C. Potent and specific genetic interference by double-stranded RNA in Caenorhabditis elegans. Nature 1998, 391, 806–811.
- Wytinck, N.; Manchur, C.L.; Li, V.H.; Whyard, S.; Belmonte, M.F. dsRNA uptake in plant pests and pathogens: Insights into RNAi-based insect and fungal control technology. Plants 2020, 9, 1780.
- Bramlett, M.; Plaetinck, G.; Maienfisch, P. RNA-based biocontrols—A new paradigm in crop protection. Engineering 2020, 6, 522–527.
- Darlington, M.; Reinders, J.D.; Sethi, A.; Lu, A.L.; Ramaseshadri, P.; Fischer, J.R.; Boeckman, C.J.; Petrick, J.S.; Roper, J.M.; Narva, K.E. RNAi for Western Corn Rootworm Management: Lessons Learned, Challenges, and Future Directions. Insects 2022, 13, 57.
- Vogel, E.; Santos, D.; Mingels, L.; Verdonckt, T.-W.; Broeck, J.V. RNA interference in insects: Protecting beneficials and controlling pests. Front. Physiol. 2019, 9, 1912.
- Nunes, C.C.; Dean, R.A. Host-induced gene silencing: A tool for understanding fungal host interaction and for developing novel disease control strategies. Mol. Plant Pathol. 2012, 13, 519–529.
- You, L.; Zhang, F.; Huang, S.; Merchant, A.; Zhou, X.; Li, Z. Over-expression of rna interference (rnai) core machinery improves susceptibility to rnai in silkworm larvae. Insect Mol. Biol. 2020, 29, 353–362.
- Zotti, M.; Dos Santos, E.A.; Cagliari, D.; Christiaens, O.; Taning, C.N.T.; Smagghe, G. RNA interference technology in crop protection against arthropod pests, pathogens and nematodes. Pest Manag. Sci. 2018, 74, 1239–1250.
- Huvenne, H.; Smagghe, G. Mechanisms of dsRNA uptake in insects and potential of RNAi for pest control: A review. J. Insect Physiol. 2010, 56, 227–235.
- Saleh, M.-C.; van Rij, R.P.; Hekele, A.; Gillis, A.; Foley, E.; O’Farrell, P.H.; Andino, R. The endocytic pathway mediates cell entry of dsRNA to induce RNAi silencing. Nat. Cell Biol. 2006, 8, 793–802.
- Joga, M.R.; Zotti, M.J.; Smagghe, G.; Christiaens, O. RNAi Efficiency, Systemic Properties, and Novel Delivery Methods for Pest Insect Control: What We Know So Far. Front. Physiol. 2016, 7, 553.
- Fletcher, S.J.; Reeves, P.T.; Hoang, B.T.; Mitter, N. A Perspective on RNAi-Based Biopesticides. Front. Plant Sci. 2020, 11, 51.
- Luck, S.; Kreszies, T.; Strickert, M.; Schweizer, P.; Kuhlmann, M.; Douchkov, D. siRNA-Finder (si-Fi) Software for RNAi-Target Design and Off-Target Prediction. Front. Plant Sci. 2019, 10, 1023.
- Whyard, S.; Singh, A.D.; Wong, S. Ingested double-stranded RNAs can act as species-specific insecticides. Insect Biochem. Mol. Biol. 2009, 39, 824–832.
- Katoch, R.; Sethi, A.; Thakur, N.; Murdock, L.L. RNAi for insect control: Current perspective and future challenges. Appl. Biochem. Biotechnol. 2013, 171, 847–873.
- Feinberg, E.H.; Hunter, C.P. Transport of dsRNA into cells by the transmembrane protein SID-1. Science 2003, 301, 1545–1547.
- Okamura, K.; Chung, W.-J.; Ruby, J.G.; Guo, H.; Bartel, D.P.; Lai, E.C. The Drosophila hairpin RNA pathway generates endogenous short interfering RNAs. Nature 2008, 453, 803–806.
- Mourrain, P.; Béclin, C.; Elmayan, T.; Feuerbach, F.; Godon, C.; Morel, J.-B.; Jouette, D.; Lacombe, A.-M.; Nikic, S.; Picault, N. Arabidopsis SGS2 and SGS3 genes are required for posttranscriptional gene silencing and natural virus resistance. Cell 2000, 101, 533–542.
- Price, D.R.; Gatehouse, J.A. RNAi-mediated crop protection against insects. Trends Biotechnol. 2008, 26, 393–400.
- Alper, S.; McBride, S.J.; Lackford, B.; Freedman, J.H.; Schwartz, D.A. Specificity and complexity of the Caenorhabditis elegans innate immune response. Mol. Cell. Biol. 2007, 27, 5544–5553.
- Chen, N.; Xu, M.-J.; Nisbet, A.J.; Huang, C.-Q.; Lin, R.-Q.; Yuan, Z.-G.; Song, H.-Q.; Zhu, X.-Q. Ascaris suum: RNAi mediated silencing of enolase gene expression in infective larvae. Exp. Parasitol. 2011, 127, 142–146.
- Jex, A.R.; Liu, S.; Li, B.; Young, N.D.; Hall, R.S.; Li, Y.; Yang, L.; Zeng, N.; Xu, X.; Xiong, Z. Ascaris suum draft genome. Nature 2011, 479, 529–533.
- Peter, R.; Van den Bossche, P.; Penzhorn, B.L.; Sharp, B. Tick, fly, and mosquito control—Lessons from the past, solutions for the future. Vet. Parasitol. 2005, 132, 205–215.
- de la Fuente, J.; Estrada-Pena, A.; Venzal, J.M.; Kocan, K.M.; Sonenshine, D.E. Overview: Ticks as vectors of pathogens that cause disease in humans and animals. Front. Biosci. 2008, 13, 6938–6946.
- Fang, E.; Liu, X.; Li, M.; Zhang, Z.; Song, L.; Zhu, B.; Wu, X.; Liu, J.; Zhao, D.; Li, Y. Advances in COVID-19 mRNA vaccine development. Signal Transduct. Target. Ther. 2022, 7, 94.
- McCoy, C.J.; Warnock, N.D.; Atkinson, L.E.; Atcheson, E.; Martin, R.J.; Robertson, A.P.; Maule, A.G.; Marks, N.J.; Mousley, A. RNA interference in adult Ascaris suum–an opportunity for the development of a functional genomics platform that supports organism-, tissue-and cell-based biology in a nematode parasite. Int. J. Parasitol. 2015, 45, 673–678.
- Xu, M.; Chen, N.; Song, H.; Lin, R.; Huang, C.; Yuan, Z.; Zhu, X. RNAi-mediated silencing of a novel Ascaris suum gene expression in infective larvae. Parasitol. Res. 2010, 107, 1499–1503.
- Yi, N.; Yu, P.; Wu, L.; Liu, Z.; Guan, J.; Liu, C.; Liu, M.; Lu, Y. RNAi-mediated silencing of Trichinella spiralis serpin-type serine protease inhibitors results in a reduction in larval infectivity. Vet. Res. 2020, 51, 139.
- Komisarczuk, A.Z.; Kongshaug, H.; Li, M.; Nilsen, F. RNAi mediated myosuppressin deficiency affects muscle development and survival in the salmon louse (Lepeophtheirus salmonis). Sci. Rep. 2019, 9, 6944.
- Lew-Tabor, A.; Kurscheid, S.; Barrero, R.; Gondro, C.; Moolhuijzen, P.; Valle, M.R.; Morgan, J.; Covacin, C.; Bellgard, M. Gene expression evidence for off-target effects caused by RNA interference-mediated gene silencing of Ubiquitin-63E in the cattle tick Rhipicephalus microplus. Int. J. Parasitol. 2011, 41, 1001–1014.
- Aljamali, M.; Bior, A.; Sauer, J.; Essenberg, R. RNA interference in ticks: A study using histamine binding protein dsRNA in the female tick Amblyomma americanum. Insect Mol. Biol. 2003, 12, 299–305.
- Manzano-Román, R.; Díaz-Martín, V.; Oleaga, A.; Siles-Lucas, M.; Pérez-Sánchez, R. Subolesin/akirin orthologs from Ornithodoros spp. soft ticks: Cloning, RNAi gene silencing and protective effect of the recombinant proteins. Vet. Parasitol. 2012, 185, 248–259.
- McCusker, P.; Toet, H.; Rathinasamy, V.; Young, N.; Beddoe, T.; Anderson, G.; Dempster, R.; McVeigh, P.; McCammick, E.; Wells, D. Molecular characterisation and vaccine efficacy of two novel developmentally regulated surface tegument proteins of Fasciola hepatica. Vet. Parasitol. 2020, 286, 109244.
- Fernando, D.D.; Marr, E.J.; Zakrzewski, M.; Reynolds, S.L.; Burgess, S.T.; Fischer, K. Gene silencing by RNA interference in Sarcoptes scabiei: A molecular tool to identify novel therapeutic targets. Parasites Vectors 2017, 10, 289.
- Yang, X.; Khan, S.; Zhao, X.; Zhang, J.; Nisar, A.; Feng, X. Suppression of hyaluronidase reduces invasion and establishment of Haemonchus contortus larvae in sheep. Vet. Res. 2020, 51, 106.
- Cagliari, D.; Dias, N.P.; Galdeano, D.M.; Dos Santos, E.Á.; Smagghe, G.; Zotti, M.J. Management of pest insects and plant diseases by non-transformative RNAi. Front. Plant Sci. 2019, 10, 1319.
