Cow’s milk allergy (CMA) is the most prevalent food allergy (FA) in infancy and early childhood and can be present with various clinical phenotypes. The significant increase in FA rates recorded has been associated with environmental and lifestyle changes that limit microbial exposure in early life and induce changes in gut microbiome composition. Gut microbiome is a diverse community of microbes that colonize the gastrointestinal tract (GIT) and perform beneficial functions for the host. This complex ecosystem interacts with the immune system and has a pivotal role in the development of oral tolerance to food antigens. Emerging evidence indicates that alterations of the gut microbiome (dysbiosis) in early life cause immune dysregulation and render the host susceptible to immune-mediated diseases later in life.
- food allergy
- cow’s milk allergy
- gut microbiome
- dysbiosis
- probiotics
- prebiotics
1. Introduction
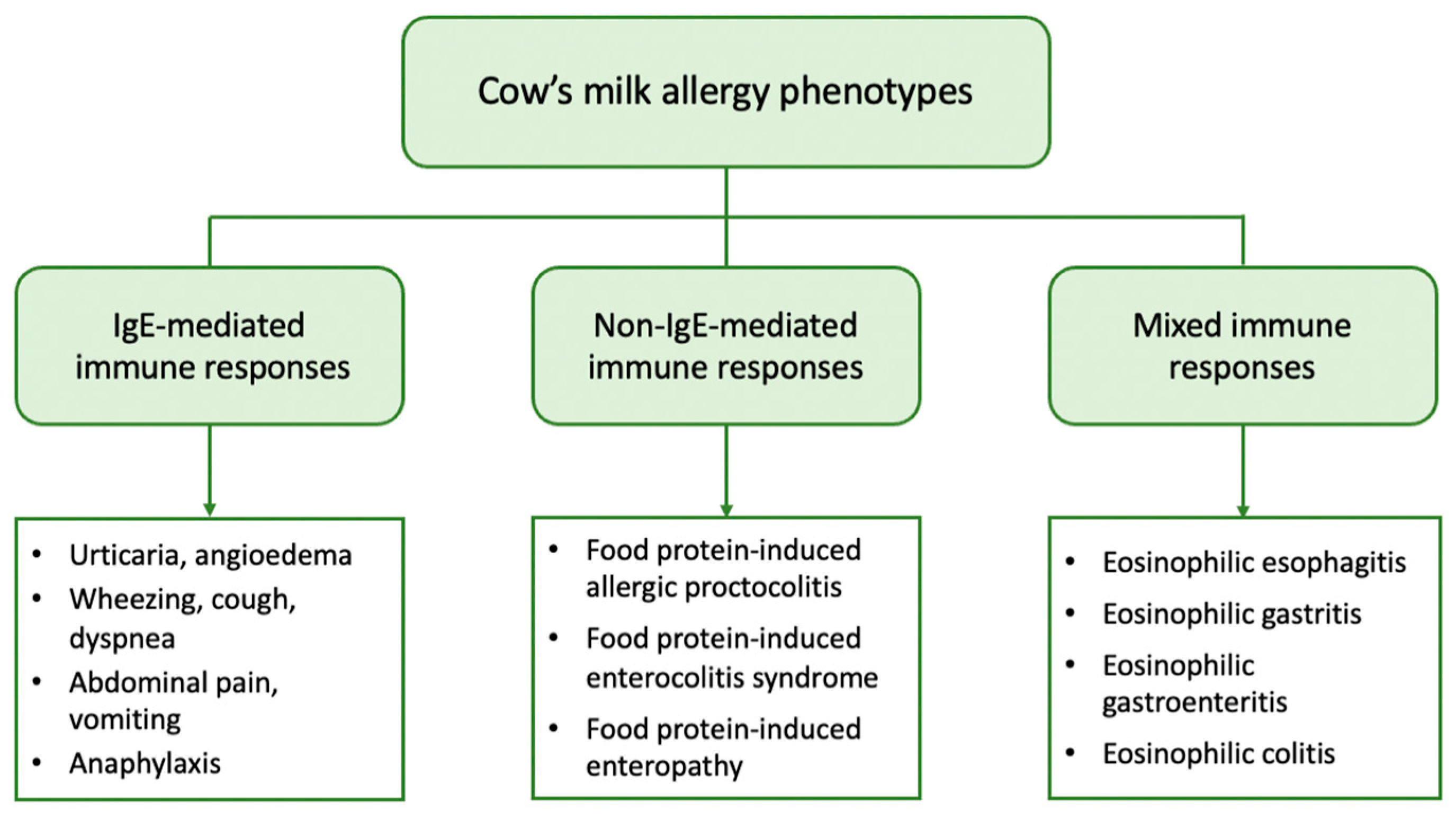
2. Clinical Phenotypes of CMA
From a clinical point of view, patients with CMA may present with a variety of symptoms linked to the immune mechanism underlying the allergic reaction. Based on the mechanism involved, CMA can be further classified into (a) IgE-mediated, which is caused by antibodies against milk proteins belonging to IgE, (b) non-IgE (or cell-mediated), where the cellular immune system, and especially T-cells, are responsible for the allergic reaction, and (c) mixed type when both IgE and immune cells are involved [18][17]. IgE-mediated CMA is the most common form and accounts for approximately 60% of all CM-induced allergic reactions [19][18]. It is caused by the production of specific IgE against CM proteins that bind to high-affinity IgE receptors (FcƐRI) on basophils and mast cells. Upon exposure, CM proteins are recognized by two or more specific IgEs that bind to FcƐRI with subsequent cross-linking of the receptor and activation of mast cells to release mediators such as histamine, tryptase, and platelet-activating factor [20][19]. These mediators cause vasodilation and elicit acute symptoms in the skin, gastrointestinal, respiratory, and cardiovascular systems. IgE-mediated reactions typically occur immediately after CM ingestion, or within 1 to 2 h, and may present as acute urticaria, angioedema, cough, wheezing, dyspnea, abdominal pain, vomiting, and hypotension. In severe cases, anaphylaxis, a systemic reaction that can be fatal, can also occur [18][17]. The diagnosis of IgE-mediated CMA is based on the combination of a compatible medical history and evidence of CM sensitization, i.e., the presence of specific IgE in skin mast cells (skin prick tests, SPTs) and/or in serum [21][20]. The non-IgE-mediated types of CMA have been far less studied. They encompass a wide range of disorders including food protein-induced allergic proctocolitis (FPIAP), food protein-induced enterocolitis syndrome (FPIES), and food protein-induced enteropathy (FPE). These disorders affect different segments of the gastrointestinal tract (GIT) and are distinguished by the delayed onset of symptoms, which can range from 2 h to several days after CM ingestion. They are caused by different immune cell-mediated mechanisms but have in common that they all lead to inflammation of the GIT. In these types of CMA, there is no evidence of CM sensitization, as no circulating specific IgE is detected [22,23][21][22]. However, a localized IgE response to the gut has been described [24][23]. FPIAP is one of the most common allergic diseases in infancy. It is characterized by the presence of mucus and blood in the stools of an otherwise healthy infant. Unlike other types of CMA, this clinical phenotype most commonly affects breastfed infants. It usually occurs in infants up to 3 months of age, and most of them develop tolerance within the first year of life. Diagnosis is based on clinical manifestations [23][22]. FPIES occurs less frequently. In two large prospective cohort studies from Israel and Spain, the cumulative incidence of CM-FPIES was 0.34% and 0.35%, respectively, in children during the first two to three years of life [25,26][24][25]. It mainly affects infants younger than nine months and is differentiated into acute and chronic FPIES. Infants with acute FPIES appear with multiple vomitings within a few hours of CM intake, which may be accompanied by pallor, lethargy, and diarrhea. In contrast, infants with chronic FPIES exhibit more chronic symptoms, such as vomiting, chronic diarrhea, and inadequate growth. Diagnosis in both cases is based on clinical criteria [27][26]. FPE is considered relatively uncommon, although its prevalence has not been thoroughly investigated. This disorder affects the small intestine and presents with malabsorption symptoms, such as chronic diarrhea, bloating, flatulence, anemia, and failure to thrive [28][27]. Mixed types of FA comprise a group of diseases generally referred to as eosinophilic gastrointestinal disorders (EGID). They are triggered by complex immune mechanisms with only partial involvement of IgE, leading to pathological eosinophilic infiltration of different parts of the GIT. Symptoms have a delayed onset and depend on the organs affected, as well as the extent of eosinophilic infiltration [29][28]. Eosinophilic esophagitis (EoE) is the most common disease, with CM and wheat being the main culprits. EoE in children presents with feeding difficulties, gagging, vomiting, and food refusal due to gastroesophageal dysfunction, and the diagnosis is based on endoscopic findings [22][21]. Atopic dermatitis (AD) is the most common chronic skin disease in childhood with an estimated prevalence of up to 20% [30][29]. It is a complex disease that is characterized by impaired skin barrier function due to various genetic, environmental, and immunologic factors. AD is usually the initial manifestation of the “atopic march” and predisposes to other allergic disorders such as FA, asthma, and allergic rhinitis [31][30]. Approximately one-third of children with moderate to severe AD develop FA, with CM being one of the main allergens [32][31]. However, the role of CM in the development of AD remains controversial, with many authors supporting the idea that CMA is a consequence of AD rather than AD being an expression or clinical phenotype of CMA [33][32]. In contrast, there are limited reports in the literature of isolated delayed-type eczematous reactions that typically occur within 6 to 48 h after food ingestion with eczematous flare-ups at AD-predisposed sites, suggesting a non-IgE-mediated pattern [32][31]. Lactose intolerance (LI) is a common gastrointestinal disorder that is caused by lactase deficiency and the inability to digest and absorb dietary lactose [34][33]. Lactose is the major carbohydrate in human and mammalian milk and is absorbed in the small intestine after first being hydrolyzed into D-glucose and D-galactose by the enzyme lactase [35][34]. Lactase expression is genetically programmed to decline gradually after weaning in about 70% of the world’s population [36][35]. As a result, lactase levels progressively decrease over time and its deficiency leads to lactose malabsorption and subsequent microbial fermentation of unabsorbed lactose in the colon [34][33]. Fermentation products, especially gasses, affect gastrointestinal function and may lead to the development of clinical symptoms, such as abdominal pain, flatulence, and diarrhea after consuming milk and dairy products. Despite similarities in clinical presentation with CMA, LI does not affect the immune system and is therefore not considered a FA. Moreover, it rarely manifests clinically in children younger than five years of age [37][36] and can be diagnosed by a variety of diagnostic methods, such as the hydrogen breath test [38][37].3. Gut Microbiome and the Factors Influencing Its Development in Early Life
The GIT is colonized by a wide variety of microorganisms, mainly bacteria, which perform substantial metabolic, immunological, and gut protective functions. The structure of these complex microbial communities varies along the GIT and shows greater diversity in the colon where the bacterial density reaches 1012 colony-forming units (CFU)/mL [39][38]. Despite the large diversity, only four major phyla, Firmicutes, Bacteroidetes, Actinobacteria, and Proteobacteria, are represented in the human gut [40][39]. However, the exact taxonomic composition depends on many host-related factors, including genetic variation, age, diet, and geographic location, and thus varies significantly among healthy individuals [41][40]. The gut microbiome is acquired at birth and undergoes dynamic changes during the first years of life. In newborns it is initially dominated by Proteobacteria (e.g., Escherichia) and Actinobacteria (e.g., Bifidobacterium). Its composition is constantly changing and increasing in diversity throughout infancy and is largely mature by the age of two to three years. It then tends to acquire a more stable composition dominated by Firmicutes and Bacteroidetes and is characterized by distinct functions [42][41]. The ratio of Enterobacteriaceae to Bacteroidaceae, referred to as the E/B ratio, reflects maturation in the adult-type microbiome and decreases with age [43][42]. However, recent evidence suggests that the colonization of the gut may begin in utero, contrary to the widespread theory that the fetal environment is sterile [44,45,46][43][44][45]. Hence, the first 1000 days of life, or the time period from conception to two years of age, is considered the critical window for shaping the microbiome and regulating the immune system [47][46]. Mode of delivery, gestational age, breastfeeding, early exposure to antibiotics, and the timing and type of complementary feeding are the dominant factors (Figure 2) that influence the composition and function of the gut microbiome in this period and, therefore, determine the development of oral tolerance to different antigens [48,49,50,51][47][48][49][50].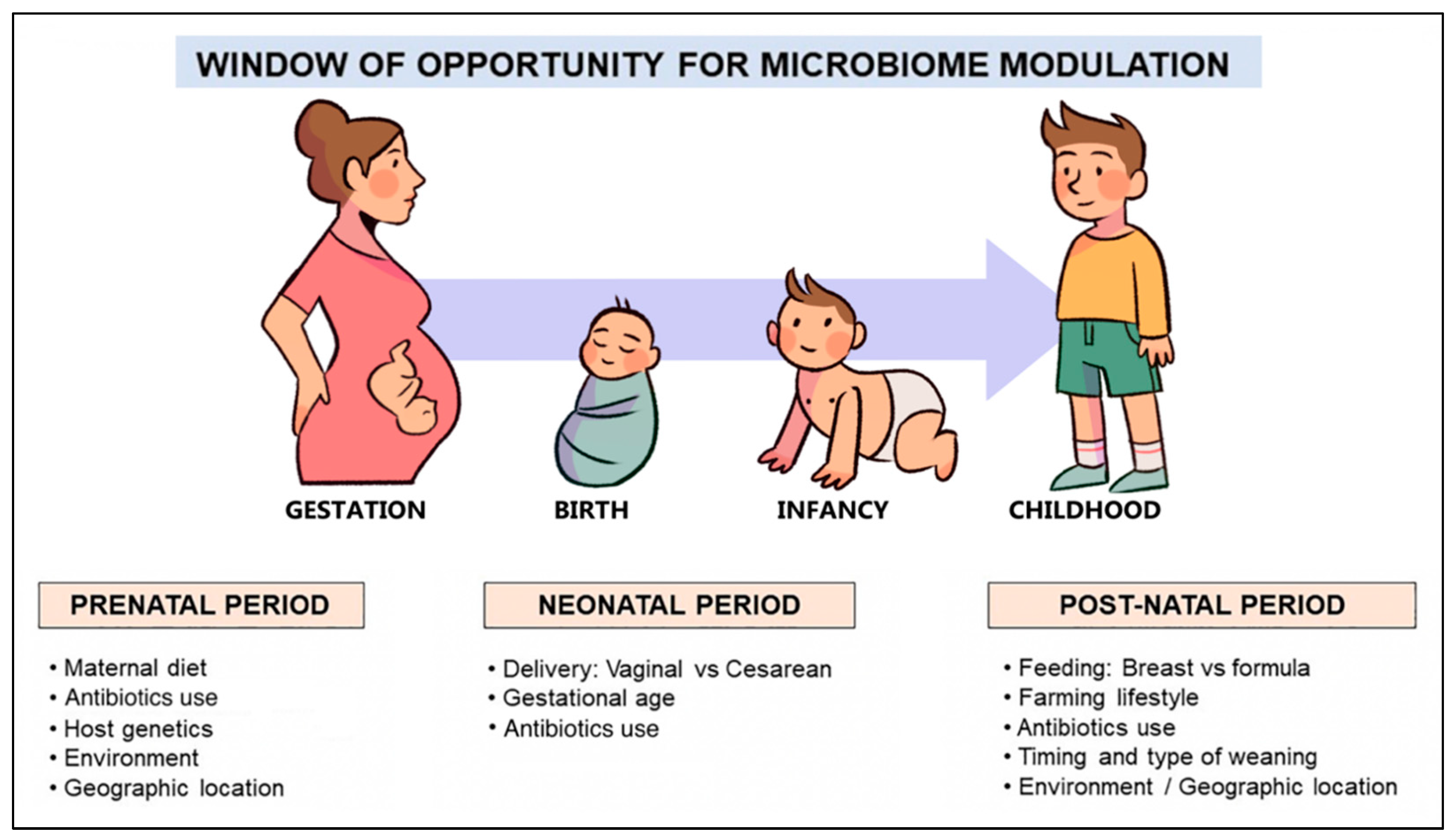
4. The Role of the Gut Microbiome in the Development of FA
Recent data indicate that the composition and metabolic activities of the gut microbiome are inextricably linked to the development of oral tolerance, which is defined as an active process of local and systemic immune unresponsiveness to food antigens [69][68]. Studies in animal models revealed that germ-free mice were more likely to be sensitized to CM proteins than those whose gut was colonized with microbes, suggesting a protective effect exerted by the gut microbiome [70,71][69][70]. Remarkably, the transfer of fecal samples from healthy infants to germ-free mice reduced the risk of CM sensitization and the occurrence of anaphylaxis when exposed to beta-lactalbumin [72][71]. These findings suggest that the commensal microbial community drives the host towards the development of oral tolerance. However, the exact mechanism by which this is achieved remains to be further elucidated. According to accepted theory, the process of oral tolerance occurs in the gut-associated lymphoid tissue (GALT) and involves the recognition of dietary antigens by dendritic cells and the induction of regulatory T cells (Tregs), as well as regulatory B cells (Bregs). Food antigens that cross the epithelial barrier are presented to naïve T cells of the mesenteric lymph nodes by specialized populations of dendritic cells in the presence of retinoic acid and TGF-β. This leads to the robust induction of antigen-specific FOXP3+ Tregs, which suppress the activation and differentiation of naïve T cells to Th types and have various anti-inflammatory roles. Suppression of Th2 responses also involves Bregs that contribute to oral tolerance by producing specific IgG4 [73][72]. Experimental research has shown that the interplay between the gut microbiome and the immune system is crucial for the production of Tregs which are necessary for the establishment and maintenance of oral tolerance. The colonization of the intestine of mice with specific bacterial strains, such as the Bifidobacterium and Clostridium species, was shown to have a protective effect on food allergen sensitization by induction of mucosal Tregs [74,75][73][74]. There are also studies denoting that commensal bacteria, particularly the Clostridia class, are likely to suppress FA through bacterial fermentation products, such as SCFAs, that promote the regulatory activity of dendritic cells and result in the induction of Tregs [76,77,78][75][76][77]. In addition to the induction of antigen specific Tregs, there is evidence that commensal microbes accelerate the development of oral tolerance through their protective effect on the intestinal epithelial barrier [79][78]. Indeed, some Clostridium species have been found to stimulate innate lymphoid cells to produce IL-22, which enhances epithelial barrier integrity and reduces intestinal permeability to protein antigens [80][79]. Another factor that has emerged as a crucial determinant for the protection and homeostatic regulation of the intestinal mucosal epithelium is the production of secretory IgA (sIgA), which is the major immunoglobulin on mucosal surfaces. Indeed, reduced fecal IgA levels and altered gut microbiome composition have been associated with food sensitization and an increased risk of anaphylaxis in mice [81][80]. The main function of sIgA, referred to as immune exclusion, is to limit the access of pathogens and food allergens to the mucosal barrier, and thus prevents their spread to the systemic compartment [82][81]. In addition, it regulates the balance of gut microbes by favoring the maintenance of beneficial members and the removal of opportunistic pathogens, which in turn prevents the absorption of pathogens and toxins [83,84][82][83]. There is evidence that mucosal secretion of sIgA is partially controlled by the gut microbiome and even that specific microbial species can induce its production to a different extent [84][83]. As a result, the composition of gut microbes strongly influences sIgA production and, consequently, the development of oral tolerance. These data demonstrate a causal role for the gut microbiome in protecting against allergic responses to food antigens. Consequently, alterations in its composition, collectively referred to as dysbiosis, lead to immune dysfunction and predispose the host to the development of FA.References
- Sicherer, S.H.; Sampson, H.A. Food allergy: Epidemiology, pathogenesis, diagnosis, and treatment. J. Allergy Clin. Immunol. 2014, 133, 291–307.
- Savage, J.; Johns, C.B. Food allergy: Epidemiology and natural history. Immunol. Allergy Clin. N. Am. 2015, 35, 45–59.
- Lifschitz, C.; Szajewska, H. Cow’s milk allergy: Evidence-based diagnosis and management for the practitioner. Eur. J. Pediatr. 2015, 174, 141–150.
- Nwaru, B.I.; Hickstein, L.; Panesar, S.S.; Roberts, G.; Muraro, A.; Sheikh, A. Prevalence of common food allergies in Europe: A systematic review and meta-analysis. Allergy 2014, 69, 992–1007.
- Schoemaker, A.A.; Sprikkelman, A.B.; Grimshaw, K.E.; Roberts, G.; Grabenhenrich, L.; Rosenfeld, L.; Siegert, S.; Dubakiene, R.; Rudzeviciene, O.; Reche, M.; et al. Incidence and natural history of challenge-proven cow’s milk allergy in European children--EuroPrevall birth cohort. Allergy 2015, 70, 963–972.
- Rona, R.J.; Keil, T.; Summers, C.; Gislason, D.; Zuidmeer, L.; Sodergren, E.; Sigurdardottir, S.T.; Lindner, T.; Goldhahn, K.; Dahlstrom, J.; et al. The prevalence of food allergy: A meta-analysis. J. Allergy Clin. Immunol. 2007, 120, 638–646.
- Katz, Y.; Rajuan, N.; Goldberg, M.R.; Eisenberg, E.; Heyman, E.; Cohen, A.; Leshno, M. Early exposure to cow’s milk protein is protective against IgE-mediated cow’s milk protein allergy. J. Allergy Clin. Immunol. 2010, 126, 77–82.
- García-Ara, M.C.; Boyano-Martínez, M.T.; Díaz-Pena, J.M.; Martín-Muñoz, M.F.; Martín-Esteban, M. Cow’s milk-specific immunoglobulin E levels as predictors of clinical reactivity in the follow-up of the cow’s milk allergy infants. Clin. Exp. Allergy 2004, 34, 866–870.
- Fiocchi, A.; Terracciano, L.; Bouygue, G.R.; Veglia, F.; Sarratud, T.; Martelli, A.; Restani, P. Incremental prognostic factors associated with cow’s milk allergy outcomes in infant and child referrals: The Milan Cow’s Milk Allergy Cohort study. Ann. Allergy Asthma Immunol. 2008, 101, 166–173.
- Skripak, J.M.; Matsui, E.C.; Mudd, K.; Wood, R.A. The natural history of IgE-mediated cow’s milk allergy. J. Allergy Clin. Immunol. 2007, 120, 1172–1177.
- Tsabouri, S.; Douros, K.; Priftis, K.N. Cow’s milk allergenicity. Endocr. Metab. Immune Disord. Drug Targets 2014, 14, 16–26.
- Yeung, J.P.; Kloda, L.A.; McDevitt, J.; Ben-Shoshan, M.; Alizadehfar, R. Oral immunotherapy for milk allergy. Cochrane Database Syst. Rev. 2012, 11, CD009542.
- Vandenplas, Y. Prevention and Management of Cow’s Milk Allergy in Non-Exclusively Breastfed Infants. Nutrients 2017, 9, 731.
- McBurney, M.I.; Davis, C.; Fraser, C.M.; Schneeman, B.O.; Huttenhower, C.; Verbeke, K.; Walter, J.; Latulippe, M.E. Establishing What Constitutes a Healthy Human Gut Microbiome: State of the Science, Regulatory Considerations, and Future Directions. J. Nutr. 2019, 149, 1882–1895.
- Moran, N.A.; Ochman, H.; Hammer, T.J. Evolutionary and ecological consequences of gut microbial communities. Annu. Rev. Ecol. Evol. Syst. 2019, 50, 451–475.
- Young, V.B. The role of the microbiome in human health and disease: An introduction for clinicians. BMJ Br. Med. J. 2017, 356, j831.
- Fiocchi, A.; Schünemann, H.J.; Brozek, J.; Restani, P.; Beyer, K.; Troncone, R.; Martelli, A.; Terracciano, L.; Bahna, S.L.; Rancé, F.; et al. Diagnosis and Rationale for Action Against Cow’s Milk Allergy (DRACMA): A summary report. J. Allergy Clin. Immunol. 2010, 126, 1119–1128.
- Sampson, H.A. Food allergy. Part 1: Immunopathogenesis and clinical disorders. J. Allergy Clin. Immunol. 1999, 103, 717–728.
- Vickery, B.P.; Chin, S.; Burks, A.W. Pathophysiology of food allergy. Pediatr. Clin. N. Am. 2011, 58, 363–376.
- Luyt, D.; Ball, H.; Makwana, N.; Green, M.R.; Bravin, K.; Nasser, S.M.; Clark, A.T. BSACI guideline for the diagnosis and management of cow’s milk allergy. Clin. Exp. Allergy 2014, 44, 642–672.
- Cianferoni, A. Non-IgE Mediated Food Allergy. Curr. Pediatr. Rev. 2020, 16, 95–105.
- Nowak-Węgrzyn, A.; Katz, Y.; Mehr, S.S.; Koletzko, S. Non-IgE-mediated gastrointestinal food allergy. J. Allergy Clin. Immunol. 2015, 135, 1114–1124.
- Lin, X.P.; Magnusson, J.; Ahlstedt, S.; Dahlman-Höglund, A.; Hanson, L.L.; Magnusson, O.; Bengtsson, U.; Telemo, E. Local allergic reaction in food-hypersensitive adults despite a lack of systemic food-specific IgE. J. Allergy Clin. Immunol. 2002, 109, 879–887.
- Katz, Y.; Goldberg, M.R.; Rajuan, N.; Cohen, A.; Leshno, M. The prevalence and natural course of food protein-induced enterocolitis syndrome to cow’s milk: A large-scale, prospective population-based study. J. Allergy Clin. Immunol. 2011, 127, 647–653.
- Alonso, S.B.; Ezquiaga, J.G.; Berzal, P.T.; Tardón, S.D.; San José, M.M.; López, P.A.; Bermejo, T.B.; Teruel, S.Q.; Echeverría Zudaire, L. Food protein-induced enterocolitis syndrome: Increased prevalence of this great unknown-results of the PREVALE study. J. Allergy Clin. Immunol. 2019, 143, 430–433.
- Leonard, S.A.; Pecora, V.; Fiocchi, A.G.; Nowak-Wegrzyn, A. Food protein-induced enterocolitis syndrome: A review of the new guidelines. World Allergy Organ. J. 2018, 11, 4.
- Labrosse, R.; Graham, F.; Caubet, J.C. Non-IgE-Mediated Gastrointestinal Food Allergies in Children: An Update. Nutrients 2020, 12, 2086.
- Gonsalves, N. Eosinophilic Gastrointestinal Disorders. Clin. Rev. Allergy Immunol. 2019, 57, 272–285.
- Nutten, S. Atopic dermatitis: Global epidemiology and risk factors. Ann. Nutr. Metab. 2015, 66 (Suppl. 1), 8–16.
- Zhu, T.H.; Zhu, T.R.; Tran, K.A.; Sivamani, R.K.; Shi, V.Y. Epithelial barrier dysfunctions in atopic dermatitis: A skin-gut-lung model linking microbiome alteration and immune dysregulation. Br. J. Dermatol. 2018, 179, 570–581.
- Bergmann, M.M.; Caubet, J.C.; Boguniewicz, M.; Eigenmann, P.A. Evaluation of food allergy in patients with atopic dermatitis. J. Allergy Clin. Immunol. Pract. 2013, 1, 22–28.
- Marzuillo, P.; Guarino, S.; Perrone, L. Atopic eczema could be a cause and not an effect of cow’s milk protein allergy. J. Pediatr. Gastroenterol. Nutr. 2014, 58, e23.
- Misselwitz, B.; Butter, M.; Verbeke, K.; Fox, M.R. Update on lactose malabsorption and intolerance: Pathogenesis, diagnosis and clinical management. Gut 2019, 68, 2080–2091.
- Fassio, F.; Facioni, M.S.; Guagnini, F. Lactose Maldigestion, Malabsorption, and Intolerance: A Comprehensive Review with a Focus on Current Management and Future Perspectives. Nutrients 2018, 10, 1599.
- Storhaug, C.L.; Fosse, S.K.; Fadnes, L.T. Country, regional, and global estimates for lactose malabsorption in adults: A systematic review and meta-analysis. Lancet Gastroenterol. Hepatol. 2017, 2, 738–746.
- Heine, R.G.; AlRefaee, F.; Bachina, P.; De Leon, J.C.; Geng, L.; Gong, S.; Madrazo, J.A.; Ngamphaiboon, J.; Ong, C.; Rogacion, J.M. Lactose intolerance and gastrointestinal cow’s milk allergy in infants and children—Common misconceptions revisited. World Allergy Organ. J. 2017, 10, 41.
- Catanzaro, R.; Sciuto, M.; Marotta, F. Lactose intolerance: An update on its pathogenesis, diagnosis, and treatment. Nutr. Res. 2021, 89, 23–34.
- Eckburg, P.B.; Bik, E.M.; Bernstein, C.N.; Purdom, E.; Dethlefsen, L.; Sargent, M.; Gill, S.R.; Nelson, K.E.; Relman, D.A. Diversity of the human intestinal microbial flora. Science 2005, 308, 1635–1638.
- Arumugam, M.; Raes, J.; Pelletier, E.; Le Paslier, D.; Yamada, T.; Mende, D.R.; Fernandes, G.R.; Tap, J.; Bruls, T.; Batto, J.M.; et al. Enterotypes of the human gut microbiome. Nature 2011, 473, 174–180.
- Adak, A.; Khan, M.R. An insight into gut microbiota and its functionalities. Cell. Mol. Life Sci. 2019, 76, 473–493.
- Bäckhed, F.; Roswall, J.; Peng, Y.; Feng, Q.; Jia, H.; Kovatcheva-Datchary, P.; Li, Y.; Xia, Y.; Xie, H.; Zhong, H.; et al. Dynamics and Stabilization of the Human Gut Microbiome during the First Year of Life. Cell Host Microbe 2015, 17, 690–703.
- Yatsunenko, T.; Rey, F.E.; Manary, M.J.; Trehan, I.; Dominguez-Bello, M.G.; Contreras, M.; Magris, M.; Hidalgo, G.; Baldassano, R.N.; Anokhin, A.P.; et al. Human gut microbiome viewed across age and geography. Nature 2012, 486, 222–227.
- Gosalbes, M.J.; Llop, S.; Vallès, Y.; Moya, A.; Ballester, F.; Francino, M.P. Meconium microbiota types dominated by lactic acid or enteric bacteria are differentially associated with maternal eczema and respiratory problems in infants. Clin. Exp. Allergy 2013, 43, 198–211.
- Moles, L.; Gómez, M.; Heilig, H.; Bustos, G.; Fuentes, S.; de Vos, W.; Fernández, L.; Rodríguez, J.M.; Jiménez, E. Bacterial diversity in meconium of preterm neonates and evolution of their fecal microbiota during the first month of life. PLoS ONE 2013, 8, e66986.
- Aagaard, K.; Ma, J.; Antony, K.M.; Ganu, R.; Petrosino, J.; Versalovic, J. The placenta harbors a unique microbiome. Sci. Transl. Med. 2014, 6, 237ra65.
- Robertson, R.C.; Manges, A.R.; Finlay, B.B.; Prendergast, A.J. The Human Microbiome and Child Growth—First 1000 Days and Beyond. Trends Microbiol. 2019, 27, 131–147.
- Francino, M.P. Early development of the gut microbiota and immune health. Pathogens 2014, 3, 769–790.
- Francino, M.P. Birth Mode-Related Differences in Gut Microbiota Colonization and Immune System Development. Ann. Nutr. Metab. 2018, 73 (Suppl. 3), 12–16.
- Peroni, D.G.; Nuzzi, G.; Trambusti, I.; Di Cicco, M.E.; Comberiati, P. Microbiome Composition and Its Impact on the Development of Allergic Diseases. Front. Immunol. 2020, 11, 700.
- Tsabouri, S.; Priftis, K.N.; Chaliasos, N.; Siamopoulou, A. Modulation of gut microbiota downregulates the development of food allergy in infancy. Allergol. Immunopathol. 2014, 42, 69–77.
- Dominguez-Bello, M.G.; Costello, E.K.; Contreras, M.; Magris, M.; Hidalgo, G.; Fierer, N.; Knight, R. Delivery mode shapes the acquisition and structure of the initial microbiota across multiple body habitats in newborns. Proc. Natl. Acad. Sci. USA 2010, 107, 11971–11975.
- Shao, Y.; Forster, S.C.; Tsaliki, E.; Vervier, K.; Strang, A.; Simpson, N.; Kumar, N.; Stares, M.D.; Rodger, A.; Brocklehurst, P.; et al. Stunted microbiota and opportunistic pathogen colonization in caesarean-section birth. Nature 2019, 574, 117–121.
- Kristensen, K.; Henriksen, L. Cesarean section and disease associated with immune function. J. Allergy Clin. Immunol. 2016, 137, 587–590.
- Kolokotroni, O.; Middleton, N.; Gavatha, M.; Lamnisos, D.; Priftis, K.N.; Yiallouros, P.K. Asthma and atopy in children born by caesarean section: Effect modification by family history of allergies—A population based cross-sectional study. BMC Pediatr. 2012, 12, 179.
- Karlström, A.; Lindgren, H.; Hildingsson, I. Maternal and infant outcome after caesarean section without recorded medical indication: Findings from a Swedish case-control study. BJOG Int. J. Obstet. Gynaecol. 2013, 120, 479–486.
- Magne, F.; Abély, M.; Boyer, F.; Morville, P.; Pochart, P.; Suau, A. Low species diversity and high interindividual variability in faeces of preterm infants as revealed by sequences of 16S rRNA genes and PCR-temporal temperature gradient gel electrophoresis profiles. FEMS Microbiol. Ecol. 2006, 57, 128–138.
- Arboleya, S.; Solís, G.; Fernández, N.; de los Reyes-Gavilán, C.G.; Gueimonde, M. Facultative to strict anaerobes ratio in the preterm infant microbiota: A target for intervention? Gut Microbes 2012, 3, 583–588.
- Jacquot, A.; Neveu, D.; Aujoulat, F.; Mercier, G.; Marchandin, H.; Jumas-Bilak, E.; Picaud, J.C. Dynamics and clinical evolution of bacterial gut microflora in extremely premature patients. J. Pediatr. 2011, 158, 390–396.
- Mai, V.; Torrazza, R.M.; Ukhanova, M.; Wang, X.; Sun, Y.; Li, N.; Shuster, J.; Sharma, R.; Hudak, M.L.; Neu, J. Distortions in development of intestinal microbiota associated with late onset sepsis in preterm infants. PLoS ONE 2013, 8, e52876.
- Cuna, A.; Morowitz, M.J.; Ahmed, I.; Umar, S.; Sampath, V. Dynamics of the preterm gut microbiome in health and disease. Am. J. Physiol. Gastrointest. Liver Physiol. 2021, 320, G411–G419.
- Lyons, K.E.; Ryan, C.A.; Dempsey, E.M.; Ross, R.P.; Stanton, C. Breast Milk, a Source of Beneficial Microbes and Associated Benefits for Infant Health. Nutrients 2020, 12, 1039.
- Levin, A.M.; Sitarik, A.R.; Havstad, S.L.; Fujimura, K.E.; Wegienka, G.; Cassidy-Bushrow, A.E.; Kim, H.; Zoratti, E.M.; Lukacs, N.W.; Boushey, H.A.; et al. Joint effects of pregnancy, sociocultural, and environmental factors on early life gut microbiome structure and diversity. Sci. Rep. 2016, 6, 31775.
- Francino, M.P. Antibiotics and the Human Gut Microbiome: Dysbioses and Accumulation of Resistances. Front. Microbiol. 2015, 6, 1543.
- Tanaka, S.; Kobayashi, T.; Songjinda, P.; Tateyama, A.; Tsubouchi, M.; Kiyohara, C.; Shirakawa, T.; Sonomoto, K.; Nakayama, J. Influence of antibiotic exposure in the early postnatal period on the development of intestinal microbiota. FEMS Immunol. Med. Microbiol. 2009, 56, 80–87.
- Fouhy, F.; Guinane, C.M.; Hussey, S.; Wall, R.; Ryan, C.A.; Dempsey, E.M.; Murphy, B.; Ross, R.P.; Fitzgerald, G.F.; Stanton, C.; et al. High-throughput sequencing reveals the incomplete, short-term recovery of infant gut microbiota following parenteral antibiotic treatment with ampicillin and gentamicin. Antimicrob. Agents Chemother. 2012, 56, 5811–5820.
- Laforest-Lapointe, I.; Arrieta, M.C. Patterns of Early-Life Gut Microbial Colonization during Human Immune Development: An Ecological Perspective. Front. Immunol. 2017, 8, 788.
- Penders, J.; Thijs, C.; Vink, C.; Stelma, F.F.; Snijders, B.; Kummeling, I.; van den Brandt, P.A.; Stobberingh, E.E. Factors influencing the composition of the intestinal microbiota in early infancy. Pediatrics 2006, 118, 511–521.
- Iweala, O.I.; Nagler, C.R. Immune privilege in the gut: The establishment and maintenance of non-responsiveness to dietary antigens and commensal flora. Immunol. Rev. 2006, 213, 82–100.
- Rodriguez, B.; Prioult, G.; Bibiloni, R.; Nicolis, I.; Mercenier, A.; Butel, M.J.; Waligora-Dupriet, A.J. Germ-free status and altered caecal subdominant microbiota are associated with a high susceptibility to cow’s milk allergy in mice. FEMS Microbiol. Ecol. 2011, 76, 133–144.
- Morin, S.; Bernard, H.; Przybylski-Nicaise, L.; Corthier, G.; Rabot, S.; Wal, J.M.; Hazebrouck, S. Allergenic and immunogenic potential of cow’s milk β-lactoglobulin and caseins evidenced without adjuvant in germ-free mice. Mol. Nutr. Food Res. 2011, 55, 1700–1707.
- Feehley, T.; Plunkett, C.H.; Bao, R.; Choi Hong, S.M.; Culleen, E.; Belda-Ferre, P.; Campbell, E.; Aitoro, R.; Nocerino, R.; Paparo, L.; et al. Healthy infants harbor intestinal bacteria that protect against food allergy. Nat. Med. 2019, 25, 448–453.
- Satitsuksanoa, P.; Jansen, K.; Głobińska, A.; van de Veen, W.; Akdis, M. Regulatory Immune Mechanisms in Tolerance to Food Allergy. Front. Immunol. 2018, 9, 2939.
- Lyons, A.; O’Mahony, D.; O’Brien, F.; MacSharry, J.; Sheil, B.; Ceddia, M.; Russell, W.M.; Forsythe, P.; Bienenstock, J.; Kiely, B.; et al. Bacterial strain-specific induction of Foxp3+ T regulatory cells is protective in murine allergy models. Clin. Exp. Allergy 2010, 40, 811–819.
- Atarashi, K.; Tanoue, T.; Shima, T.; Imaoka, A.; Kuwahara, T.; Momose, Y.; Cheng, G.; Yamasaki, S.; Saito, T.; Ohba, Y.; et al. Induction of colonic regulatory T cells by indigenous Clostridium species. Science 2011, 331, 337–341.
- Furusawa, Y.; Obata, Y.; Fukuda, S.; Endo, T.A.; Nakato, G.; Takahashi, D.; Nakanishi, Y.; Uetake, C.; Kato, K.; Kato, T.; et al. Commensal microbe-derived butyrate induces the differentiation of colonic regulatory T cells. Nature 2013, 504, 446–450.
- Smith, P.M.; Howitt, M.R.; Panikov, N.; Michaud, M.; Gallini, C.A.; Bohlooly, Y.M.; Glickman, J.N.; Garrett, W.S. The microbial metabolites, short-chain fatty acids, regulate colonic Treg cell homeostasis. Science 2013, 341, 569–573.
- Arpaia, N.; Campbell, C.; Fan, X.; Dikiy, S.; van der Veeken, J.; deRoos, P.; Liu, H.; Cross, J.R.; Pfeffer, K.; Coffer, P.J.; et al. Metabolites produced by commensal bacteria promote peripheral regulatory T-cell generation. Nature 2013, 504, 451–455.
- Cao, S.; Feehley, T.J.; Nagler, C.R. The role of commensal bacteria in the regulation of sensitization to food allergens. FEBS Lett. 2014, 588, 4258–4266.
- Stefka, A.T.; Feehley, T.; Tripathi, P.; Qiu, J.; McCoy, K.; Mazmanian, S.K.; Tjota, M.Y.; Seo, G.Y.; Cao, S.; Theriault, B.R.; et al. Commensal bacteria protect against food allergen sensitization. Proc. Natl. Acad. Sci. USA 2014, 111, 13145–13150.
- Smeekens, J.M.; Johnson-Weaver, B.T.; Hinton, A.L.; Azcarate-Peril, M.A.; Moran, T.P.; Immormino, R.M.; Kesselring, J.R.; Steinbach, E.C.; Orgel, K.A.; Staats, H.F.; et al. Fecal IgA, Antigen Absorption, and Gut Microbiome Composition Are Associated With Food Antigen Sensitization in Genetically Susceptible Mice. Front. Immunol. 2020, 11, 599637.
- Corthésy, B. Multi-faceted functions of secretory IgA at mucosal surfaces. Front. Immunol. 2013, 4, 185.
- Zhang, B.; Liu, E.; Gertie, J.A.; Joseph, J.; Xu, L.; Pinker, E.Y.; Waizman, D.A.; Catanzaro, J.; Hamza, K.H.; Lahl, K.; et al. Divergent T follicular helper cell requirement for IgA and IgE production to peanut during allergic sensitization. Sci. Immunol. 2020, 5, eaay2754.
- León, E.D.; Francino, M.P. Roles of Secretory Immunoglobulin A in Host-Microbiota Interactions in the Gut Ecosystem. Front. Microbiol. 2022, 13, 880484.
