The glycation of various biomolecules is the root cause of many pathological conditions associated with diabetic nephropathy and end-stage kidney disease. Glycation imbalances metabolism and increases renal cell injury. Numerous therapeutic measures have narrowed down the adverse effects of endogenous glycation, but efficient and potent measures are miles away. Recent advances in the identification and characterization of noncoding RNAs, especially the long noncoding RNAs (lncRNAs), have opened a mammon of new biology to explore the mitigations for glycation-associated diabetic nephropathy. Furthermore, tissue-specific distribution and condition-specific expression make lncRNA a promising key for second-generation therapeutic interventions. Though the techniques to identify and exemplify noncoding RNAs are rapidly evolving, the lncRNA study encounters multiple methodological constraints.
- diabetes
- diabetic nephropathy
- glycation
- long noncoding RNA
- biomarkers
- therapeutics
1. Introduction
| Diabetic Complications | lncRNA Involved | Mode of Action | References |
|---|---|---|---|
| Diabetic neuropathy | lncRNA NEAT1 | Regulate disease progression by targeting two miRNAs, miR-183-5p and miR-433-3p. | [9] |
| lncRNA TP73-AS1 | Sponges decreases miR-142 and upregulates HMGB1 expression as well as promotes cell proliferation. Its silencing decreases neuropathic pain. | [10] | |
| Diabetic retinopathy | lncRNA HOTTIP | Induces p38/MAPK signaling and promotes retinal cell inflammatory response and diabetic retinopathy progression. | [11] |
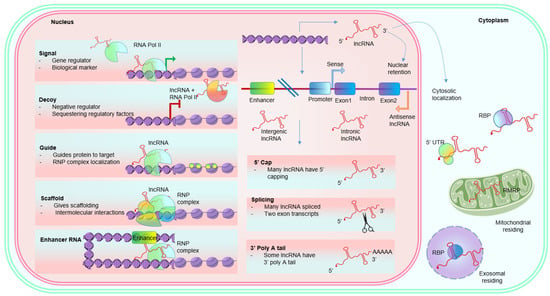
1.2. Functional Properties of LncRNAs
1.2.1. Role in Transcription Regulation
1.2.2. Role in Post-Transcriptional Regulation
1.2.3. Role in Epigenetic Regulation
2. Understanding the Role of LncRNAs in AGEs-Related DN
Studies have shown that lncRNAs play a significant role in AGEs-mediated metabolic malfunctions [38]. So far, the involvement of lncRNAs-mediated AGEs-RAGE signaling in cancer, the immune system, and neurodegenerative diseases has been studied in detail [39]. However, few studies have reported the role of lncRNAs in glycation-associated diabetic complications [15][40][41][42][43] (Table 2).| LncRNA Involved | Diabetic Complication | Mode of Action | References | ||||
|---|---|---|---|---|---|---|---|
| lncRNA Arid2-IR |
Diabetic retinopathy | Regulates oxidative stress, inflammatory responses, and endothelial cell dysfunction via interacting with Smad3 | [40] | ||||
| lncRNA MIAT | Diabetic retinopathy | AGE-induced HRPCs MIAT and CASP1 expressions increase, followed by the release of IL-1β, IL-18, and suppression of cell viability. | [42] | ||||
| lncRNA MEG3 | Diabetic vascular diseases | Upregulates in AGE-induced cells and suppresses cell viability and proliferation by modulating the MEG3/miR-193/p21 pathway | [41] | ||||
| lncRNA BANCR | Regulates cell apoptosis | [12] | |||||
| lncRNA URIDS | Diabetic Wound Healing | Upregulates upon AGE induction. Regulates collagen production and deposition by targeting Plod1. It delays the wound healing process. | [ | lncRNA MEG3 | Regulates VEGF and TGF-β1 expressions | [13] | |
| Inflammatory diabetes complications | lncRNA DRAIR | Regulates macrophages/monocyte pro/anti-inflammatory phenotypes in T2DM. Down-regulated by diabetogenic factors | [14] | ||||
| Diabetic wound healing | lncRNA URIDS | Impairs collagen production and crosslinking by interacting with Plod1 and delays wound healing | [15] | ||||
| lncRNA MALAT1 | Increases wound healing and upregulate fibroblast activation in diabetic mice by activating the HIF-1α signaling pathway. | [16] | |||||
| 15 | ] | ||||||
| lncRNA E330013P06 | Inflammatory diabetes complications | Increases inflammatory response upon AGE induction; by triggering pro-inflammatory gene. It also enhances foam cell formation. | [43] | Diabetic cardiomyopathy | lncRNA HOTAIR | Downregulates DCM effects by activating SIRT1 expressions and sponging miR-34a. | [17] |
| lncRNA Kcnq1ot1 | Promotes pyroptosis by regulating expressions of miR-214-3p and caspase-1 | [18] | |||||
| lncRNA H19 | Regulates high glucose-induced apoptosis by targeting VDAC1. It also improves left ventricular function when overexpressed. | [19] | |||||
| lncRNA Crende | Negatively regulates cardiac fibroblast differentiation. Also, its expression is induced by Smad3 in cardiac fibroblasts. It inhibits myofibroblastic gene transcription. | [20] | |||||
| lncRNA TUG1 | Its knockdown lessened DCM-induced hypertrophy and diastolic function. Also, its silencing upregulates the expression of some miRNAs | [21] | |||||
| Diabetic nephropathy | lncRNA Blnc1 | Attenuates renal fibrosis and inflammation and affects oxidative stress by NF-κB and NRF2/HO-1 pathways | [22] | ||||
| lncRNA TCF7 | It acts as a sponge against miR-200c and triggers endoplasmic reticulum stress in patients with DN. | [23] | |||||
| lncRNA Gas5 | Alleviates cell proliferation and fibrosis sponging miR-221 and upregulates SIRT1 | [24] |
1.1. Structural Properties of LncRNAs
2.1. LncRNA-Mediated RAGE Gene Expression and Signaling
The lncRNAs such as H9, MIAT, and MEG3 significantly regulate the pathogenesis of various diabetic complications. Recently, the role of lncRNA Arid2-IR in AGEs-induced retinal endothelial cells was also observed. By binding to Smad3, LncRNA Arid2-IR regulated levels of oxidative stress, inflammation, and apoptosis [40]. Similarly, lncRNA HOTAIR regulated RAGE expression and inflammation in acute myocardium infarction-induced rat models [44]. Likewise, the delayed process of diabetic wound healing in AGEs-induced fibroblast was associated with lncRNA URIDS [15]. Overexpression and knockdown studies also confirmed the role of another lncRNA MVIH in the AGEs-RAGE signaling pathway responsible for tumor induction and progression in cancer [45]. In AGEs-induced endothelial cells, lncRNA MEG3 regulated cell viability, proliferation, and apoptosis through the lncRNA MEG3/MiR-93/p21 mediated pathway during the onset of diabetic vascular diseases [41]. In a cell-based study on hypoxia/reoxygenation-injured cardiomyocytes, lncRNA SNHG12 down-regulated RAGE, NF-κB expression, and pro-inflammatory responses [46]. In a different context, the lncRNA TP73-AS1-mediated RAGE-HMGB1 signaling pathway upregulated NF-κB expression and pro-inflammatory cytokine levels [12][47]. The above-discussed lncRNAs can be a promising biomarker to detect complications in AGEs-RAGE signaling-mediated metabolism.2.2. LncRNAs That Regulate AGER Gene Expression and Signaling
Notably, the expression pattern of lncRNA AGER-1 correlates with AGER expression levels (r = 0.360, p = 2.15 × 10−18) possibly by binding and sponging miRNA-185. Animal-based studies confirmed these results with suppressed cell proliferation rate, migration, and colony-forming efficiency in nude mice [48]. In a different context, lncRNA AGER-1 induced cell cycle arrest and promoted apoptosis, thereby inhibiting cell proliferation and migration efficiency of tumor cells in several types of cancer [49]. Yet another study indicated the sponging effect of lncRNA AGER-1 for miR-182, an inhibitor of AGER1 [50]. Recently, a study showed that both AGER and its positive regulator lncRNA AGER-1 have the significant diagnostic potential for lung adenocarcinoma. Both AGER and lncRNA AGER-1 regulate apoptosis, cell migration, and antitumor responses [51]. In summary, lncRNA AGER-1 can be a promising agent against AGEs signaling in metabolic disorders.References
- Palazzo, A.F.; Lee, E.S. Non-Coding RNA: What Is Functional and What Is Junk? Front. Genet. 2015, 5, 1–11.
- Dunham, I.; Kundaje, A.; Aldred, S.F.; Collins, P.J.; Davis, C.A.; Doyle, F.; Epstein, C.B.; Frietze, S.; Harrow, J.; Kaul, R.; et al. An Integrated Encyclopedia of DNA Elements in the Human Genome. Nature 2012, 489, 57–74.
- Carninci, P.; Kasukawa, T.; Katayama, S.; Gough, J.; Frith, M.C.; Maeda, N.; Oyama, R.; Ravasi, T.; Lenhard, B.; Wells, C.; et al. Molecular Biology: The Transcriptional Landscape of the Mammalian Genome. Science 2005, 309, 1559–1563.
- Hon, C.C.; Ramilowski, J.A.; Harshbarger, J.; Bertin, N.; Rackham, O.J.L.; Gough, J.; Denisenko, E.; Schmeier, S.; Poulsen, T.M.; Severin, J.; et al. An Atlas of Human Long Non-Coding RNAs with Accurate 5′ Ends. Nature 2017, 543, 199–204.
- Hangauer, M.J.; Vaughn, I.W.; McManus, M.T. Pervasive Transcription of the Human Genome Produces Thousands of Previously Unidentified Long Intergenic Noncoding RNAs. PLoS Genet. 2013, 9, e1003569.
- Bushati, N.; Cohen, S.M. MicroRNA Functions. Annu. Rev. Cell Dev. Biol. 2007, 23, 175–205.
- Esteller, M. Non-Coding RNAs in Human Disease. Nat. Rev. Genet. 2011, 12, 861–874.
- Chen, M. PlncRNADB: A Repository of Plant LncRNAs and LncRNA-RBP Protein Interactions. Curr. Bioinform. 2019, 14, 621–627.
- Asadi, G.; Rezaei Varmaziar, F.; Karimi, M.; Rajabinejad, M.; Ranjbar, S.; Gorgin Karaji, A.; Salari, F.; Afshar Hezarkhani, L.; Rezaiemanesh, A. Determination of the Transcriptional Level of Long Non-Coding RNA NEAT-1, Downstream Target MicroRNAs, and Genes Targeted by MicroRNAs in Diabetic Neuropathy Patients. Immunol. Lett. 2021, 232, 20–26.
- Zhang, R.; Jin, H.; Lou, F. The Long Non-Coding RNA TP73-AS1 Interacted with MiR-142 to Modulate Brain Glioma Growth through HMGB1/RAGE Pathway. J. Cell. Biochem. 2018, 119, 3007–3016.
- Sun, Y.; Liu, Y. LncRNA HOTTIP Improves Diabetic Retinopathy by Regulating the P38-MAPK Pathway. Eur. Rev. Med. Pharmacol. Sci. 2018, 22, 2941–2948.
- Zhang, X.; Zou, X.; Li, Y.; Wang, Y. Downregulation of lncRNA BANCR participates in the development of retinopathy among diabetic patients. Exp. Ther. Med. 2019, 17, 4132–4138.
- Zhang, D.; Qin, H.; Leng, Y.; Li, X.; Zhang, L.; Bai, D.; Meng, Y.; Wang, J. LncRNA MEG3 Overexpression Inhibits the Development of Diabetic Retinopathy by Regulating TGF-β1 and VEGF. Exp. Ther. Med. 2018, 16, 2337–2342.
- Reddy, M.A.; Amaram, V.; Das, S.; Tanwar, V.S.; Ganguly, R.; Wang, M.; Lanting, L.; Zhang, L.; Abdollahi, M.; Chen, Z.; et al. LncRNA DRAIR Is Downregulated in Diabetic Monocytes and Modulates the Inflammatory Phenotype via Epigenetic Mechanisms. JCI Insight 2021, 6, 1–22.
- Hu, M.; Wu, Y.; Yang, C.; Wang, X.; Wang, W.; Zhou, L.; Zeng, T.; Zhou, J.; Wang, C.; Lao, G.; et al. Novel Long Noncoding RNA Lnc-URIDS Delays Diabetic Wound Healing by Targeting Plod1. Diabetes 2020, 69, 2144–2156.
- Liu, X.; Duan, L.; Chen, Y.; Jin, X.; Zhu, N.; Zhou, X.; Wei, H. LncRNA MALAT1 Accelerates Wound Healing of Diabetic Mice Transfused with Modified Autologous Blood via the HIF-1 a Signaling Pathway. Mol. Ther. Nucleic Acid 2019, 17, 504–515.
- Gao, L.; Wang, X.; Guo, S.; Xiao, L.; Liang, C.; Wang, Z.; Li, Y.; Liu, Y.; Yao, R.; Liu, Y.; et al. LncRNA HOTAIR Functions as a Competing Endogenous RNA to Upregulate SIRT1 by Sponging MiR-34a in Diabetic Cardiomyopathy. J. Cell. Physiol. 2019, 234, 4944–4958.
- Yang, F.; Qin, Y.; Wang, Y.; Li, A.; Lv, J.; Sun, X.; Che, H.; Han, T.; Meng, S.; Bai, Y.; et al. LncRNA KCNQ1OT1 mediates pyroptosis in diabetic cardiomyopathy. Cell. Physiol. Biochem. 2018, 5, 1230–1244.
- Li, X.; Wang, H.; Yao, B.; Xu, W.; Chen, J.; Zhou, X. LncRNA H19/MiR-675 Axis Regulates Cardiomyocyte Apoptosis by Targeting VDAC1 in Diabetic Cardiomyopathy. Sci. Rep. 2016, 6, 1–9.
- Zheng, D.; Zhang, Y.; Hu, Y.; Guan, J.; Xu, L.; Xiao, W.; Zhong, Q.; Ren, C.; Lu, J.; Liang, J.; et al. Long Noncoding RNA Crnde Attenuates Cardiac Fibrosis via Smad3-Crnde Negative Feedback in Diabetic Cardiomyopathy. FEBS J. 2019, 286, 1645–1655.
- Zhao, L.; Li, W.; Zhao, H. Inhibition of Long Non-Coding RNA TUG1 Protects against Diabetic Cardiomyopathy Induced Diastolic Dysfunction by Regulating MiR-499-5p. Am. J. Transl. Res. 2020, 12, 718–730.
- Feng, X.; Zhao, J.; Ding, J.; Shen, X.; Zhou, J.; Xu, Z. LncRNA Blnc1 Expression and Its Effect on Renal Fibrosis in Diabetic Nephropathy. Am. J. Transl. Res. 2019, 11, 5664–5672.
- Liu, H.; Sun, H. LncRNA TCF7 Triggered Endoplasmic Reticulum Stress through a Sponge Action with MiR-200c in Patients with Diabetic Nephropathy. Eur. Rev. Med. Pharmacol. Sci. 2019, 23, 5912–5922.
- Ge, X.; Xu, B.; Xu, W.; Xia, L.; Xu, Z.; Shen, L.; Peng, W.; Huang, S. Long Noncoding RNA GAS5 Inhibits Cell Proliferation and Fibrosis in Diabetic Nephropathy by Sponging MiR-221 and Modulating SIRT1 Expression. Aging 2019, 11, 8745–8759.
- Derrien, T.; Johnson, R.; Bussotti, G.; Tanzer, A.; Djebali, S.; Tilgner, H.; Guernec, G.; Martin, D.; Merkel, A.; Knowles, D.G.; et al. The GENCODE v7 Catalog of Human Long Noncoding RNAs: Analysis of Their Gene Structure, Evolution, and Expression. Genome Res. 2012, 22, 1775–1789.
- Bridges, M.C.; Daulagala, A.C.; Kourtidis, A. LNCcation: LncRNA Localization and Function. J. Cell Biol. 2021, 220, 1–17.
- Klapproth, C.; Sen, R.; Stadler, P.F.; Findeiß, S.; Fallmann, J. Common Features in LncRNA Annotation and Classification: A Survey. Non-Coding RNA 2021, 7, 77.
- Parasramka, M.A.; Maji, S.; Matsuda, A.; Yan, I.K.; Patel, T. Pharmacology & Therapeutics Long Non-Coding RNAs as Novel Targets for Therapy in Hepatocellular Carcinoma. Pharmacol. Ther. 2016, 161, 67–78.
- Su, P.P.; Liu, D.W.; Zhou, S.J.; Chen, H.; Wu, X.M.; Liu, Z.S. Down—Regulation of Risa Improves Podocyte Injury by Enhancing Autophagy in Diabetic Nephropathy. Mil. Med. Res. 2022, 9, 1–13.
- Gil, N.; Ulitsky, I. Regulation of Gene Expression by Cis-acting Long non-coding RNAs. Nat. Rev. Genet. 2020, 21, 102–117.
- Wutz, A. Gene Silencing in X-Chromosome Inactivation: Advances in Understanding Facultative Heterochromatin Formation. Nat. Rev. Genet. 2011, 12, 542–553.
- Grossi, E.; Raimondi, I.; Goñi, E.; González, J.; Marchese, F.P.; Chapaprieta, V.; Martín-Subero, J.I.; Guo, S.; Huarte, M.A. LncRNA-SWI/SNF Complex Crosstalk Controls Transcriptional Activation at Specific Promoter Regions. Nat. Commun. 2020, 11, 1–16.
- He, R.; Luo, D.; Mo, Y. Emerging Roles of LncRNAs in the Post-Transcriptional Regulation in Cancer. Genes Dis. 2019, 6, 6–15.
- Lee, S.; Kopp, F.; Chang, T.; Sataluri, A.; Chen, B.; Sivakumar, S.; Yu, H.; Xie, Y.; Mendell, J.T.; Comprehensive, C.S. Noncoding RNA NORAD Regulates Genomic Stability by Sequestering PUMILIO proteins. Cell 2017, 164, 69–80.
- Morlando, M.; Fatica, A. Alteration of Epigenetic Regulation by Ong Noncoding RNAs in Cancer. Int. J. Mol. Sci. 2018, 19, 570.
- Isoda, T.; Moore, A.J.; He, Z.; Chandra, V.; Aida, M.; Denholtz, M.; Piet van Hamburg, J.; Fisch, K.M.; Chang, A.N.; Fahl, S.P.; et al. Non-Coding Transcription Instructs Chromatin Folding and Compartmentalization to Dictate Enhancer-Promoter Communication and T cell fate. Cell 2017, 171, 103–119.
- Dueva, R.; Akopyan, K.; Pederiva, C.; Trevisan, D.; Dhanjal, S.; Lindqvist, A.; Farnebo, M.; Dueva, R.; Akopyan, K.; Pederiva, C.; et al. Neutralization of the Positive Charges on Histone tails by RNA Promotes an Open Chromatin Structure. Cell Chem. Biol. 2019, 26, 1436–1449.
- Sanchez, C.A.; Kawamura, Y.; Yamamoto, Y.; Takeshita, F.; Ochiya, T. Emerging Roles of Long Non-Coding RNA in Cancer. Cancer Sci. 2018, 109, 2093–2100.
- Chrysanthou, M.; Estruch, I.M.; Rietjens, I.M.C.M.; Wichers, H.J.; Hoppenbrouwers, T. In Vitro Methodologies to Study the Role of Advanced Glycation End Products (AGEs) in Neurodegeneration. Nutrients 2022, 14, 363.
- Xiao, H.; Yang, H.; Zeng, Y. Long Non-Coding RNA Arid2-IR Affects Advanced Glycation End Products-Induced Human Retinal Endothelial Cell Injury by Binding to Smad3. Int. Ophthalmol. 2020, 40, 1123–1133.
- Ju, C.; Sheng, Z.; Wang, Q.; Li, Y.; Wang, X.; Li, S.; Qi, Q.; Yuan, Z. Advanced Glycation End Products of Bovine Serum Albumin Affect the Cell Growth of Human Umbilical Vein Endothelial Cells via Modulation of MEG3/MiR-93/P21 Pathway. Acta Biochim. Biophys. Sin. 2018, 51, 41–50.
- Yu, X.; Ma, X.; Lin, W.; Xu, Q.; Zhou, H.; Kuang, H.Y. Long Noncoding RNA MIAT Regulates Primary Human Retinal Pericyte Pyroptosis by Modulating MiR-342–3p Targeting of CASP1 in Diabetic Retinopathy. Exp. Eye Res. 2021, 202, 108300.
- Reddy, M.A.; Chen, Z.; Park, J.T.; Wang, M.; Lanting, L.; Zhang, Q.; Bhatt, K.; Leung, A.; Wu, X.; Putta, S.; et al. Regulation of Inflammatory Phenotype in Macrophages by a Diabetes-Induced Long Noncoding RNA. Diabetes 2014, 63, 4249–4261.
- Lu, W.; Zhu, L.; Ruan, Z.B.; Wang, M.X.; Ren, Y.; Li, W. HOTAIR Promotes Inflammatory Response after Acute Myocardium Infarction by Upregulating RAGE. Eur. Rev. Med. Pharmacol. Sci. 2018, 22, 7423–7430.
- Hu, S.; Zheng, Q.; Xiong, J.; Wu, H.; Wang, W.; Zhou, W. Long Non-Coding RNA MVIH Promotes Cell Proliferation, Migration, Invasion through Regulating Multiple Cancer-Related Pathways, and Correlates with Worse Prognosis in Pancreatic Ductal Adenocarcinomas. Am. J. Transl. Res. 2020, 12, 2118–2135.
- Lu, P.; Xiao, S.; Chen, S.; Fu, Y.; Zhang, P.; Yao, Y.; Chen, F. LncRNA SNHG12 Downregulates RAGE to Attenuate Hypoxia-Reoxygenation-Induced Apoptosis in H9c2 Cells. Biosci. Biotechnol. Biochem. 2021, 85, 866–873.
- Li, S.; Huang, Y.; Huang, Y.; Fu, Y.; Tang, D.; Kang, R.; Zhou, R.; Fan, X.G. The Long Non-Coding RNA TP73-AS1 Modulates HCC Cell Proliferation through MiR-200a-Dependent HMGB1/RAGE Regulation. J. Exp. Clin. Cancer Res. 2017, 36, 1–12.
- Pan, Z.; Liu, L.; Nie, W.; Miggin, S.; Qiu, F.; Cao, Y.; Chen, J.; Yang, B.; Zhou, Y.; Lu, J.; et al. Long Non-Coding RNA AGER-1 Functionally Upregulates the Innate Immunity Gene AGER and Approximates Its Anti-Tumor Effect in Lung Cancer. Mol. Carcinog. 2018, 57, 305–318.
- Lin, M.; Li, Y.; Xian, J.; Chen, J.; Feng, Y.; Mao, C.; Pan, Y.; Li, Z.; Zeng, Y.; Yang, L.; et al. Long Non-Coding RNA AGER-1 Inhibits Colorectal Cancer Progression through Sponging MiR-182. Int. J. Biol. Markers 2020, 35, 10–18.
- Pidíková, P.; Herichová, I. MiRNA Clusters with Up-Regulated Expression in Colorectal Cancer. Cancers 2021, 13, 2979.
- Abdelwahab, O.; Awad, N.; Elserafy, M.; Badr, E. A Feature Selection-Based Framework to Identify Biomarkers for Cancer Diagnosis: A Focus on Lung Adenocarcinoma. PLoS ONE 2022, 17, e0269126.
