In all eukaryotes, autophagy is the main pathway for nutrient recycling, which encapsulates parts of the cytoplasm and organelles in double-membrane vesicles, and then fuses with lysosomes/vacuoles to degrade them. Autophagy is a highly dynamic and relatively complex process influenced by multiple factors. Under normal growth conditions, it is maintained at basal levels. However, when plants are subjected to biotic and abiotic stresses, such as pathogens, drought, waterlogging, nutrient deficiencies, etc., autophagy is activated to help cells to survive under stress conditions. TAt present, the regulation of autophagy is mainly reflected in hormones, second messengers, post-transcriptional regulation, and protein post-translational modification.
- autophagy-related protein
- degradation
- ubiquitin
- proteasome
- autophagy
1. Introduction
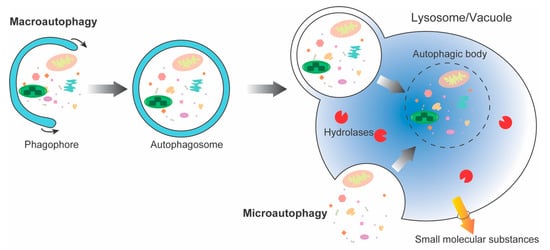
2. ATG Complexes in Plants and Animals
ATG proteins play an indispensable role in the process of autophagy in plant and animal cells. According to their diverse functions in different stages of autophagy, core ATG proteins can be divided into four major complexes (Figure 2).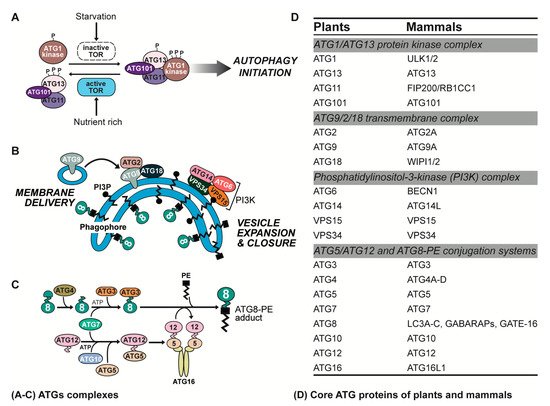
3. Degradation of Yeast Autophagy-Related Proteins
|
Eukaryotes |
ATGs |
E3 Ligases |
Ubiquitin Chain Types |
References |
|---|---|---|---|---|
|
Yeast |
Atg9 |
Met30 |
Unknown |
[23] |
|
Atg32 |
Rsp5 |
Unknown |
[25] |
|
|
Mammalian |
ULK1 |
CUL3-KLHL20 |
K48 |
[26] |
|
BTRC |
K48 |
[27] |
||
|
TRAF3 |
K48 |
[28] |
||
|
NEDD4L |
K27 and K29 |
[29] |
||
|
ATG101 |
HUWE1 |
K48 |
[30] |
|
|
WIPI2 |
HUWE1 |
Unknown |
[31] |
|
|
ATG14L |
ZBTB16-CUL3-Roc1 |
Unknown |
[32] |
|
|
BECN1 |
NEDD4 |
K11 |
[33] |
|
|
RNF216 |
K48 |
[34] |
||
|
CUL3-KLHL20 |
Unknown |
[35] |
||
|
CUL3-KLHL38 |
K48 |
[35] |
||
|
TRAF6 |
K63 |
[36] |
||
|
VPS34 |
UBE3C |
K29/K48 branched |
[37] |
|
|
FBXL20-CUL1-SKP1 |
Unknown |
[38] |
||
|
ATG16L1 |
Gigaxonin |
K48 |
[39] |
|
|
LC3B |
BIRC6 |
Single ubiquitin |
[40] |
|
|
ATG3 |
PTK2 |
Unknown |
[41] |
|
|
Plants |
ATG6 |
SINAT1/2 |
Unknown |
[42] |
|
ATG13 |
SINAT1/2 |
K48 |
[43] |
|
|
SH3P2 |
XopL |
Unknown |
[44] |
4. Degradation of Autophagy-Related Proteins in Metazoa
4.1. Degradation of ULK Complex Components
4.2. Degradation of ATG9 Complex Components
4.3. Degradation of PI3K Complex Components
4.4. Degradation of the ATG12-Conjugation System Components
4.5. Degradation of the LC3-Conjugation System Components
5. Degradation of Plant Autophagy-Related Proteins
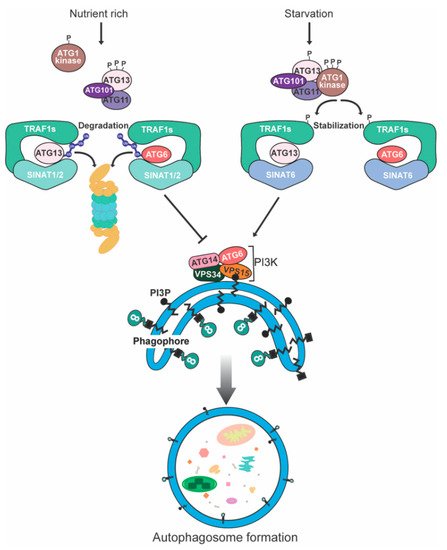
References
- Sarika, C.; Sreedevi, P.; Mishra, S.; Singh, S.N. Multifaceted Housekeeping Functions of Autophagy. J. Indian Inst. Sci. 2017, 97, 79–94.
- Huang, H.; Ayaz, A.; Zheng, M.; Yang, X.; Zaman, W.; Zhao, H.; Lü, S. Arabidopsis KCS5 and KCS6 Play Redundant Roles in Wax Synthesis. Int. J. Mol. Sci. 2022, 23, 4450.
- van Doorn, W.G.; Papini, A. Ultrastructure of autophagy in plant cells: A review. Autophagy 2013, 9, 1922–1936.
- Bolliet, V.; Labonne, J.; Olazcuaga, L.; Panserat, S.; Seiliez, I. Modeling of autophagy-related gene expression dynamics during long term fasting in European eel (Anguilla anguilla). Sci. Rep. 2017, 7, 17896.
- Lamark, T.; Johansen, T. Mechanisms of selective autophagy. Annu. Rev. Cell Dev. Biol. 2021, 37, 143–169.
- Li, F.; Chung, T.; Vierstra, R.D. AUTOPHAGY-RELATED11 plays a critical role in general autophagy- and senescence-induced mitophagy in Arabidopsis. Plant Cell 2014, 26, 788–807.
- Marion, J.; Le Bars, R.; Besse, L.; Batoko, H.; Satiat-Jeunemaitre, B. Multiscale and Multimodal Approaches to Study Autophagy in Model Plants. Cells 2018, 7, 5.
- Nakatogawa, H. Mechanisms governing autophagosome biogenesis. Nat. Rev. Mol. Cell Biol. 2020, 21, 439–458.
- Sieńko, K.; Poormassalehgoo, A.; Yamada, K.; Goto-Yamada, S. Microautophagy in Plants: Consideration of Its Molecular Mechanism. Cells 2020, 9, 887.
- Liaquat, F.; Munis, M.F.H.; Arif, S.; Haroon, U.; Shi, J.; Saqib, S.; Zaman, W.; Che, S.; Liu, Q. PacBio single-molecule long-read sequencing reveals genes tolerating manganese stress in Schima superba saplings. Front. Genet. 2021, 12, 635043.
- Ayaz, A.; Huang, H.; Zheng, M.; Zaman, W.; Li, D.; Saqib, S.; Zhao, H.; Lü, S. Molecular cloning and functional analysis of GmLACS2-3 reveals its involvement in cutin and suberin biosynthesis along with abiotic stress tolerance. Int. J. Mol. Sci. 2021, 22, 9175.
- Manghwar, H.; Hussain, A.; Ali, Q.; Liu, F. Brassinosteroids (BRs) role in plant development and coping with different stresses. Int. J. Mol. Sci. 2022, 23, 1012.
- Manghwar, H.; Hussain, A.; Ullah, A.; Gul, S.; Shaban, M.; Khan, A.H.; Ali, M.; Sani, S.G.A.S.; Chaudhary, H.J.; Munis, M.F.H. Expression analysis of defense related genes in wheat and maize against Bipolaris sorokiniana. Physiol. Mol. Plant Pathol. 2018, 103, 36–46.
- Ali, Q.; Yu, C.; Hussain, A.; Ali, M.; Ahmar, S.; Sohail, M.A.; Riaz, M.; Ashraf, M.F.; Abdalmegeed, D.; Wang, X. Genome engineering technology for durable disease resistance: Recent progress and future outlooks for sustainable agriculture. Front. Plant Sci. 2022, 13, 860281.
- Ali, Q.; Zheng, H.; Rao, M.J.; Ali, M.; Hussain, A.; Saleem, M.H.; Nehela, Y.; Sohail, M.A.; Ahmed, A.M.; Kubar, K.A. Advances, limitations, and prospects of biosensing technology for detecting phytopathogenic bacteria. Chemosphere 2022, 296, 133773.
- Liaquat, F.; Qunlu, L.; Arif, S.; Haroon, U.; Saqib, S.; Zaman, W.; Jianxin, S.; Shengquan, C.; Li, L.X.; Akbar, M. Isolation and characterization of pathogen causing brown rot in lemon and its control by using ecofriendly botanicals. Physiol. Mol. Plant Pathol. 2021, 114, 101639.
- Senft, D.; Ronai, Z.A. UPR, autophagy, and mitochondria crosstalk underlies the ER stress response. Trends Biochem. Sci. 2015, 40, 141–148.
- Manghwar, H.; Li, J. Endoplasmic Reticulum Stress and Unfolded Protein Response Signaling in Plants. Int. J. Mol. Sci. 2022, 23, 828.
- Manghwar, H.; Hussain, A. Mechanism of tobacco osmotin gene in plant responses to biotic and abiotic stress tolerance: A brief history. Biocell 2022, 46, 623.
- Yim, W.W.-Y.; Mizushima, N. Lysosome biology in autophagy. Cell Discov. 2020, 6, 6.
- Jin, M.; He, D.; Backues, S.K.; Freeberg, M.A.; Liu, X.; Kim, J.K.; Klionsky, D.J. Transcriptional regulation by Pho23 modulates the frequency of autophagosome formation. Curr. Biol. 2014, 24, 1314–1322.
- Smothers, D.B.; Kozubowski, L.; Dixon, C.; Goebl, M.G.; Mathias, N. The abundance of Met30p limits SCF(Met30p) complex activity and is regulated by methionine availability. Mol. Cell Biol. 2000, 20, 7845–7852.
- Feng, Y.; Ariosa, A.R.; Yang, Y.; Hu, Z.; Dengjel, J.; Klionsky, D.J. Downregulation of autophagy by Met30-mediated Atg9 ubiquitination. Proc. Natl. Acad. Sci. USA 2021, 118, e2005539118.
- Hu, G.; Rios, L.; Yan, Z.; Jasper, A.M.; Luera, D.; Luo, S.; Rao, H. Autophagy regulator Atg9 is degraded by the proteasome. Biochem. Biophys. Res. Commun. 2020, 522, 254–258.
- Camougrand, N.; Vigié, P.; Gonzalez, C.; Manon, S.; Bhatia-Kiššová, I. The yeast mitophagy receptor Atg32 is ubiquitinated and degraded by the proteasome. PLoS ONE 2020, 15, e0241576.
- Liu, C.C.; Lin, Y.C.; Chen, Y.H.; Chen, C.M.; Pang, L.Y.; Chen, H.A.; Wu, P.R.; Lin, M.Y.; Jiang, S.T.; Tsai, T.F.; et al. Cul3-KLHL20 Ubiquitin Ligase Governs the Turnover of ULK1 and VPS34 Complexes to Control Autophagy Termination. Mol. Cell 2016, 61, 84–97.
- Deng, R.; Zhang, H.L.; Huang, J.H.; Cai, R.Z.; Wang, Y.; Chen, Y.H.; Hu, B.X.; Ye, Z.P.; Li, Z.L.; Mai, J.; et al. MAPK1/3 kinase-dependent ULK1 degradation attenuates mitophagy and promotes breast cancer bone metastasis. Autophagy 2021, 17, 3011–3029.
- Shi, J.H.; Ling, C.; Wang, T.T.; Zhang, L.N.; Liu, W.W.; Qin, Y.; Tan, Y.H.; Cui, N.P.; Ni, Z.Y. TRK-fused gene (TFG) regulates ULK1 stability via TRAF3-mediated ubiquitination and protects macrophages from LPS-induced pyroptosis. Cell Death Dis. 2022, 13, 93.
- Nazio, F.; Carinci, M.; Valacca, C.; Bielli, P.; Strappazzon, F.; Antonioli, M.; Ciccosanti, F.; Rodolfo, C.; Campello, S.; Fimia, G.M.; et al. Fine-tuning of ULK1 mRNA and protein levels is required for autophagy oscillation. J. Cell Biol. 2016, 215, 841–856.
- Lee, J.; Kim, J.; Shin, J.; Kang, Y.; Choi, J.; Cheong, H. ATG101 Degradation by HUWE1-Mediated Ubiquitination Impairs Autophagy and Reduces Survival in Cancer Cells. Int. J. Mol. Sci. 2021, 22, 9182.
- Wan, W.; You, Z.; Zhou, L.; Xu, Y.; Peng, C.; Zhou, T.; Yi, C.; Shi, Y.; Liu, W. mTORC1-Regulated and HUWE1-Mediated WIPI2 Degradation Controls Autophagy Flux. Mol. Cell 2018, 72, 303–315.e6.
- Zhang, T.; Dong, K.; Liang, W.; Xu, D.; Xia, H.; Geng, J.; Najafov, A.; Liu, M.; Li, Y.; Han, X.; et al. G-protein-coupled receptors regulate autophagy by ZBTB16-mediated ubiquitination and proteasomal degradation of Atg14L. eLife 2015, 4, e06734.
- Platta, H.W.; Abrahamsen, H.; Thoresen, S.B.; Stenmark, H. Nedd4-dependent lysine-11-linked polyubiquitination of the tumour suppressor Beclin 1. Biochem. J. 2012, 441, 399–406.
- Xu, C.; Feng, K.; Zhao, X.; Huang, S.; Cheng, Y.; Qian, L.; Wang, Y.; Sun, H.; Jin, M.; Chuang, T.H.; et al. Regulation of autophagy by E3 ubiquitin ligase RNF216 through BECN1 ubiquitination. Autophagy 2014, 10, 2239–2250.
- Li, X.; Yang, K.B.; Chen, W.; Mai, J.; Wu, X.Q.; Sun, T.; Wu, R.Y.; Jiao, L.; Li, D.D.; Ji, J.; et al. CUL3 (cullin 3)-mediated ubiquitination and degradation of BECN1 (beclin 1) inhibit autophagy and promote tumor progression. Autophagy 2021, 17, 4323–4340.
- Shi, C.S.; Kehrl, J.H. TRAF6 and A20 regulate lysine 63-linked ubiquitination of Beclin-1 to control TLR4-induced autophagy. Sci. Signal. 2010, 3, ra42.
- Chen, Y.H.; Huang, T.Y.; Lin, Y.T.; Lin, S.Y.; Li, W.H.; Hsiao, H.J.; Yan, R.L.; Tang, H.W.; Shen, Z.Q.; Chen, G.C.; et al. VPS34 K29/K48 branched ubiquitination governed by UBE3C and TRABID regulates autophagy, proteostasis and liver metabolism. Nat. Commun. 2021, 12, 1322.
- Xiao, J.; Zhang, T.; Xu, D.; Wang, H.; Cai, Y.; Jin, T.; Liu, M.; Jin, M.; Wu, K.; Yuan, J. FBXL20-mediated Vps34 ubiquitination as a p53 controlled checkpoint in regulating autophagy and receptor degradation. Genes Dev. 2015, 29, 184–196.
- Scrivo, A.; Codogno, P.; Bomont, P. Gigaxonin E3 ligase governs ATG16L1 turnover to control autophagosome production. Nat. Commun. 2019, 10, 780.
- Jia, R.; Bonifacino, J.S. Negative regulation of autophagy by UBA6-BIRC6-mediated ubiquitination of LC3. eLife 2019, 8, e50034.
- Ma, K.; Fu, W.; Tang, M.; Zhang, C.; Hou, T.; Li, R.; Lu, X.; Wang, Y.; Zhou, J.; Li, X.; et al. PTK2-mediated degradation of ATG3 impedes cancer cells susceptible to DNA damage treatment. Autophagy 2017, 13, 579–591.
- Qi, H.; Xia, F.N.; Xie, L.J.; Yu, L.J.; Chen, Q.F.; Zhuang, X.H.; Wang, Q.; Li, F.; Jiang, L.; Xie, Q.; et al. TRAF Family Proteins Regulate Autophagy Dynamics by Modulating AUTOPHAGY PROTEIN6 Stability in Arabidopsis. Plant Cell 2017, 29, 890–911.
- Qi, H.; Li, J.; Xia, F.N.; Chen, J.Y.; Lei, X.; Han, M.Q.; Xie, L.J.; Zhou, Q.M.; Xiao, S. Arabidopsis SINAT Proteins Control Autophagy by Mediating Ubiquitylation and Degradation of ATG13. Plant Cell 2020, 32, 263–284.
- Leong, J.X.; Raffeiner, M.; Spinti, D.; Langin, G.; Franz-Wachtel, M.; Guzman, A.R.; Kim, J.G.; Pandey, P.; Minina, A.E.; Macek, B.; et al. A bacterial effector counteracts host autophagy by promoting degradation of an autophagy component. EMBO J. 2022, e110352.
- Hosokawa, N.; Hara, T.; Kaizuka, T.; Kishi, C.; Takamura, A.; Miura, Y.; Iemura, S.; Natsume, T.; Takehana, K.; Yamada, N.; et al. Nutrient-dependent mTORC1 association with the ULK1-Atg13-FIP200 complex required for autophagy. Mol. Biol. Cell 2009, 20, 1981–1991.
- Mizushima, N.; White, E.; Rubinsztein, D.C. Breakthroughs and bottlenecks in autophagy research. Trends Mol. Med. 2021, 27, 835–838.
- Zhao, Y.G.; Chen, Y.; Miao, G.; Zhao, H.; Qu, W.; Li, D.; Wang, Z.; Liu, N.; Li, L.; Chen, S.; et al. The ER-Localized Transmembrane Protein EPG-3/VMP1 Regulates SERCA Activity to Control ER-Isolation Membrane Contacts for Autophagosome Formation. Mol. Cell 2017, 67, 974–989.e6.
- Thurston, T.L.; Boyle, K.B.; Allen, M.; Ravenhill, B.J.; Karpiyevich, M.; Bloor, S.; Kaul, A.; Noad, J.; Foeglein, A.; Matthews, S.A.; et al. Recruitment of TBK1 to cytosol-invading Salmonella induces WIPI2-dependent antibacterial autophagy. EMBO J. 2016, 35, 1779–1792.
- Dikic, I.; Elazar, Z. Mechanism and medical implications of mammalian autophagy. Nat. Rev. Mol. Cell Biol. 2018, 19, 349–364.
- Shizukuishi, S.; Ogawa, M.; Ryo, A.; Ohnishi, M. Streptococcus pneumoniae promotes its own survival via choline-binding protein CbpC-mediated degradation of ATG14. Autophagy 2020, 16, 1529–1531.
- Kang, R.; Zeh, H.J.; Lotze, M.T.; Tang, D. The Beclin 1 network regulates autophagy and apoptosis. Cell Death Differ. 2011, 18, 571–580.
- Funderburk, S.F.; Wang, Q.J.; Yue, Z. The Beclin 1-VPS34 complex--at the crossroads of autophagy and beyond. Trends Cell Biol. 2010, 20, 355–362.
- Han, T.; Guo, M.; Gan, M.; Yu, B.; Tian, X.; Wang, J.B. TRIM59 regulates autophagy through modulating both the transcription and the ubiquitination of BECN1. Autophagy 2018, 14, 2035–2048.
- van der Veen, A.G.; Ploegh, H.L. Ubiquitin-like proteins. Annu. Rev. Biochem. 2012, 81, 323–357.
- Haller, M.; Hock, A.K.; Giampazolias, E.; Oberst, A.; Green, D.R.; Debnath, J.; Ryan, K.M.; Vousden, K.H.; Tait, S.W. Ubiquitination and proteasomal degradation of ATG12 regulates its proapoptotic activity. Autophagy 2014, 10, 2269–2278.
- Li, X.; He, S.; Ma, B. Autophagy and autophagy-related proteins in cancer. Mol. Cancer 2020, 19, 12.
- Zhang, N.; Yang, Y.; Lu, H.; Xiang, Y.; Huang, X.; Hu, R.; Chen, Z.; Yuan, W.; Peng, R.; Peng, J.; et al. Spodoptera litura autophagy-related protein 1 interacts with autophagy-related protein 5 and enhances its degradation. Insect Mol. Biol. 2017, 26, 190–203.
- Xie, X.; Bi, H.L.; Lai, S.; Zhang, Y.L.; Li, N.; Cao, H.J.; Han, L.; Wang, H.X.; Li, H.H. The immunoproteasome catalytic β5i subunit regulates cardiac hypertrophy by targeting the autophagy protein ATG5 for degradation. Sci. Adv. 2019, 5, eaau0495.
- Mallik, A.; Yammani, R.R. Saturated fatty acid palmitate negatively regulates autophagy by promoting ATG5 protein degradation in meniscus cells. Biochem. Biophys. Res. Commun. 2018, 502, 370–374.
- Rehman, N.U.; Zeng, P.; Mo, Z.; Guo, S.; Liu, Y.; Huang, Y.; Xie, Q. Conserved and Diversified Mechanism of Autophagy between Plants and Animals upon Various Stresses. Antioxidants 2021, 10, 1736.
- Kabeya, Y.; Mizushima, N.; Yamamoto, A.; Oshitani-Okamoto, S.; Ohsumi, Y.; Yoshimori, T. LC3, GABARAP and GATE16 localize to autophagosomal membrane depending on form-II formation. J. Cell Sci. 2004, 117 Pt 13, 2805–2812.
- Tanida, I.; Sou, Y.S.; Minematsu-Ikeguchi, N.; Ueno, T.; Kominami, E. Atg8L/Apg8L is the fourth mammalian modifier of mammalian Atg8 conjugation mediated by human Atg4B, Atg7 and Atg3. FEBS J. 2006, 273, 2553–2562.
- Jiang, T.X.; Zou, J.B.; Zhu, Q.Q.; Liu, C.H.; Wang, G.F.; Du, T.T.; Luo, Z.Y.; Guo, F.; Zhou, L.M.; Liu, J.J.; et al. SIP/CacyBP promotes autophagy by regulating levels of BRUCE/Apollon, which stimulates LC3-I degradation. Proc. Natl. Acad. Sci. USA 2019, 116, 13404–13413.
- Liu, Y.; Levine, B. Autosis and autophagic cell death: The dark side of autophagy. Cell Death Differ. 2015, 22, 367–376.
- Lu, X.; Zhang, J.; Li, Y.Q.; Liu, Q.X.; Zhou, D.; Deng, X.F.; Qiu, Y.; Chen, Q.; Li, M.Y.; Liu, X.Q.; et al. Plasmodium Circumsporozoite Protein Enhances the Efficacy of Gefitinib in Lung Adenocarcinoma Cells by Inhibiting Autophagy via Proteasomal Degradation of LC3B. Front. Cell Dev. Biol. 2022, 10, 830046.
- Besteiro, S.; Brooks, C.F.; Striepen, B.; Dubremetz, J.F. Autophagy protein Atg3 is essential for maintaining mitochondrial integrity and for normal intracellular development of Toxoplasma gondii tachyzoites. PLoS Pathog. 2011, 7, e1002416.
- Metlagel, Z.; Otomo, C.; Takaesu, G.; Otomo, T. Structural basis of ATG3 recognition by the autophagic ubiquitin-like protein ATG12. Proc. Natl. Acad. Sci. USA 2013, 110, 18844–18849.
- Liu, Y.; Bassham, D.C. Autophagy: Pathways for self-eating in plant cells. Annu. Rev. Plant Biol. 2012, 63, 215–237.
- Suttangkakul, A.; Li, F.; Chung, T.; Vierstra, R.D. The ATG1/ATG13 protein kinase complex is both a regulator and a target of autophagic recycling in Arabidopsis. Plant Cell 2011, 23, 3761–3779.
- Liu, F.; Hu, W.; Li, F.; Marshall, R.S.; Zarza, X.; Munnik, T.; Vierstra, R.D. AUTOPHAGY-RELATED14 and Its Associated Phosphatidylinositol 3-Kinase Complex Promote Autophagy in Arabidopsis. Plant Cell 2020, 32, 3939–3960.
- Kim, D.Y.; Scalf, M.; Smith, L.M.; Vierstra, R.D. Advanced proteomic analyses yield a deep catalog of ubiquitylation targets in Arabidopsis. Plant Cell 2013, 25, 1523–1540.
- Bao, Y.; Wang, C.; Jiang, C.; Pan, J.; Zhang, G.; Liu, H.; Zhang, H. The tumor necrosis factor receptor-associated factor (TRAF)-like family protein SEVEN IN ABSENTIA 2 (SINA2) promotes drought tolerance in an ABA-dependent manner in Arabidopsis. New Phytol. 2014, 202, 174–187.
