Since first being documented in ancient times, the relation of inflammation with injury and disease has evolved in complexity and causality. Early observations supported a cause (injury) and effect (inflammation) relationship, but the number of pathologies linked to chronic inflammation suggests that inflammation itself acts as a potent promoter of injury and disease. Additionally, results from studies over the last 25 ydearcades point to chronic inflammation and innate immune signaling as a critical link between stress (exogenous and endogenous) and adaptation. This brief review looks to highlight the role of the innate immune response in disease pathology, and recent findings indicating the innate immune response to chronic stresses as an influence in driving adaptation.
- inflammation
- innate immunity
- cancer
- autoinflammation
- neuro-muscular degeneration
- Fanconi anemia
- adaption
1. Overview
Since first being documented in ancient times by Greek and Egyptian physicians, the relationship of inflammation with injury and disease has evolved in complexity as well as causality. Initial characteristic descriptions by Hippocrates (5th century BC) included swelling or “edema” of the affected tissue. Later, Aulus Celsus (1st century BC/1st century AD) described the four main manifestations of inflammation: pain, edema, warmth, and redness of the interested tissue. The Roman physician/surgeon Galen later added the fifth hallmark of inflammation—loss of function of the affected tissue—a hallmark most closely associated with muscular, bone, or joint manifestations of inflammation visible, at the time, upon examination of the patient [1]. The empirical observations set the framework for detailed scientific studies over the centuries that resulted in the discovery and characterization of the immune system and its role in the response to not only injury, but also disease. These early studies supported a cause-and-effect theory whereby injury or disease gave rise to inflammation. However, is inflammation simply a case of the immune system responding to disease or can inflammation have a causative role in disease? The number of pathologies now linked to chronic inflammation would suggest inflammation may be able to both promote and be the initiating stimulus in multiple pathologies, but to this day the inflammation–disease relationship remains controversial. To clearly understand this relationship, one must first understand the relationship between inflammation, innate and acquired (or cell-based) immunity, DNA repair, and the assortment of mutations and epigenetic modifications accumulated during chronic inflammation.
12. Innate Immunity vs. Adaptive Immunity
23. Adaption vs. Disease in Chronic Inflammation
The interaction of organisms at almost every level with their surrounding environment is tutored by some form of immune response. Infectious agents, toxins, and other environmental stresses induce an innate immune inflammatory response, which in the acute phase has the objective of protecting the organism from the stressing agent by removing the stressor and repairing any damage, as stated above. These are transient stresses that the organism either is able to overcome and re-establish homeostasis or succumbs within a relatively short period of time. However, what happens when the stress is sub-lethal and yet cannot be resolved? As prolonged inflammation/inflammatory signaling generally has a negative impact on cells and tissues, chronic activation of the associated pathways can ultimately lead to chronic damage and cell death, with continual homeostatic imbalance and even death of the organism/species (in extreme cases depending on the stress). Thus, chronic inflammation can establish a selective pressure unless some means of adapting to the given stress can be achieved. Studies on a smaller scale have shown a similar scenario plays out in the cancer micro-environment, with the development or emergence of cancer stem cells [32]. The unrelated, random acquisition of a mutation at this point could be beneficial in establishing an adaptation to the stress, harmful by promoting disease, both, or neither. Most strict adaptations or silent modifications are not observed unless by chance, as healthy individuals are not often biomedical study subjects, and genetic longitudinal studies over a lifetime to catch these alterations are not practical. In contrast, there are multiple examples where mutations are associated with adaptation and disease or disease alone.2.1. Bone Marrow Failure Disorders and Acute Myelogenous Leukemia
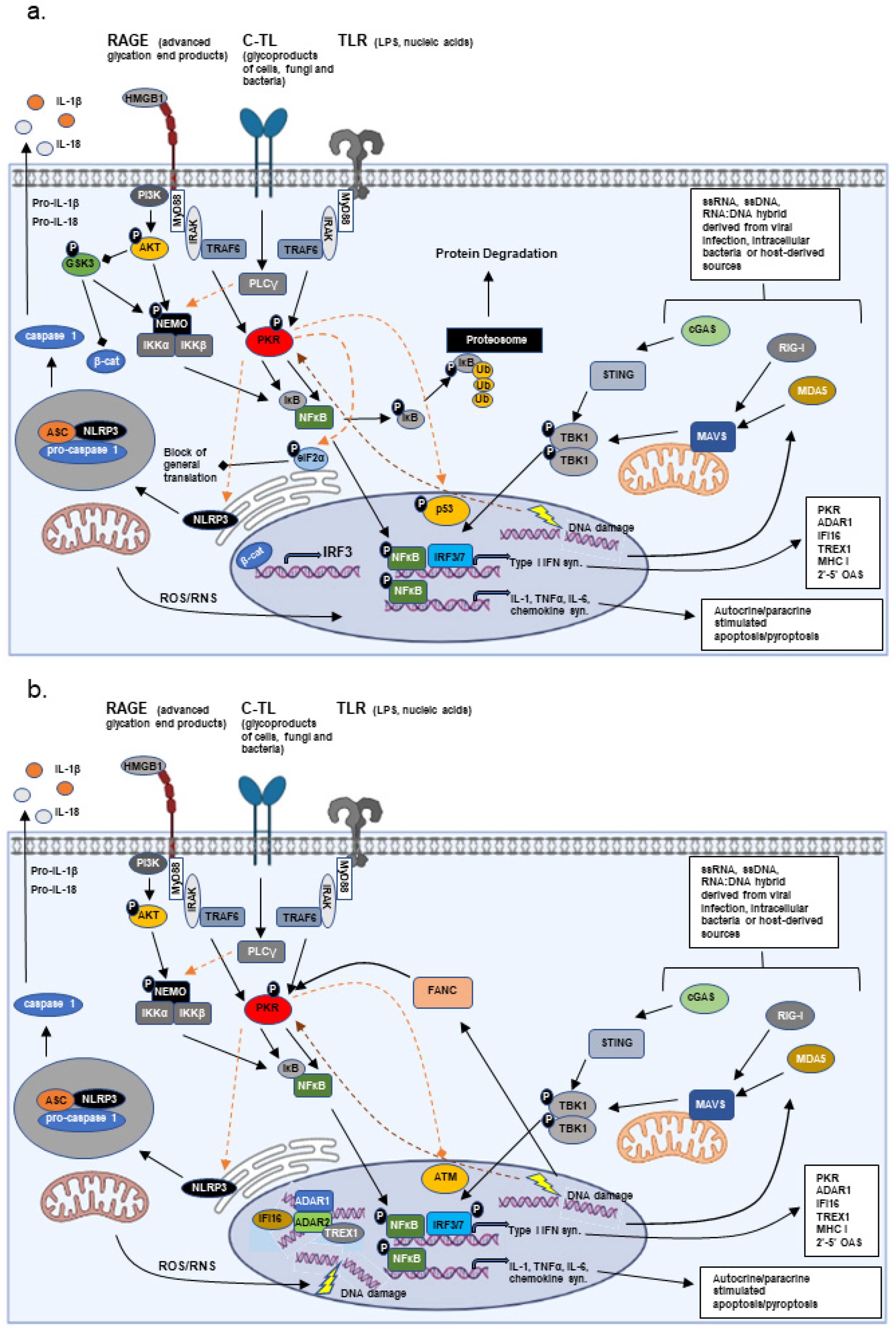
One of the more interesting adaptation/disease models is that provided by bone marrow failure disorders. The initial underlying etiology of these disorders is diverse, ranging from gene mutations (FA, myelodysplastic syndromes (MDS), Diamond-Blackfan Anemia (DBA), Shwachman-Diamond Syndrome (SDS), aplastic anemia (AA)), environmental effectors (AA), and unknown factors (MDS, AA) [12][33][34][35]. Age and prior therapeutic treatment with antineoplastics also plays an important role in MDS. In the early phase, there is an establishment of a pro-inflammatory/pro-apoptotic bone marrow environment, which initially leads to the cell death of hematopoietic progenitors and the associated anemia. In severe cases of anemia, affected individuals become dependent on transfusions for survival. At some point during disease progression, progenitor cells lose sensitivity to the pro-inflammatory/apoptotic environment and begin to expand to the peripheral blood, eventually culminating in AML. During this transition, adaptive changes/mutations occur to promote the resistance of hematopoietic progenitor cells to the pro-inflammatory environment [12][34][35][36]. This loss of apoptotic control happens to be one of the main initiating events in the formation of tumors, and under conditions of inflammation can result in a population of cells which no longer have the ability to respond to the growth control branch of inflammatory signaling while still maintaining the ability to respond to tissue repair, survival, and proliferative signals. Recent data demonstrated that innate immune/inflammatory signaling has a strong influence on AML progression. Elevated levels of active PKR in bone marrow derived progenitor cells from FA and MDS patients were first reported in 2004 and 2008, respectively [37][38]. In MDS, the number of bone marrow derived mononucleated cells (BMMCs) that contained elevated levels of active PKR were found to increase with increasing International Prognostic Scoring System (IPSS) risk categorization of the patient from which the BMMCs were derived. Not only was more active PKR present, but active PKR was localized primarily to the nucleus in BMMCs from high-risk patients [38]. Similarly, active PKR was observed in both the nucleus and cytosol in acute leukemic cells where PKR activity was determined to be necessary for leukemic cell growth [39][40]. These initial studies suggested that activated innate immune/inflammatory signaling could have a promoting effect on tumor development, but how this might occur was not clear until 2015. PKR, whose expression and/or activation is elevated in diverse tumors, was demonstrated to accelerate leukemic development [8][12][41]. In AML patients, PKR expression was directly correlated with poor overall survival and shorter time of remission, regardless of the presence of unfavorable cytogenetics. The authors determined that PKR activation enhanced the rate of mutagenesis by suppressing ataxia-telangiectasia mutated (ATM) kinase, an enzyme key to DNA double-strand break repair, thus promoting MDS progression to AML [41] (Figure 1b). Thus, activation of PKR could favor an enhanced rate of mutagenesis. Unfortunately, in this case, PKR activation and enhanced mutagenesis promoted continuous clonal survival and expansion, resulting in disease progression and death of the organism.2.2. Hemoglobin Beta
Of course, there are also cases where the opposite is held true, where a random mutation’s emergence in a population is promoted by an environmental factor. Among one of the most prominent adaptation/disease mutations in this category is that involving hemoglobin beta (HBB) mutations associated with sickle cell anemia (HbS), in global regions prone to malaria. In these areas, the sickle cell mutation has been selected as an adaptation to an infective pathogen. While these mutations offer a level of resistance to malarial infections and disease, the associated phenotype can be deadly in individuals homozygous for HBB mutation. The general theory holds that the acquisition of HBB mutations occurred by random “happenchance” that, in regions with high incidence of malaria, became a favorable trait. In regions where malaria was not present, similar mutations would have been eventually deleted from the population pool. However, is it possible that the high incidence of malarial infection in these regions over the long-term may have favored the original emergence of these mutations in some manner? In a direct manner, no, but a recent study has shown that the acquisition of the HbS mutation in Africans is not totally random [42]. In the associated study, the authors reported that the mutations at this locus in Ghanaians occurred at a higher frequency than that observed in Europeans, suggesting that long-term environmental selective pressure exerted by the presence of malaria has favored mutagenesis at this locus.2.3. Gastro-Intestinal Microbiome
The importance of the relationship between the microbiome composition and host health has increasingly become more apparent. Individuals begin to acquire microbial species at birth, starting with maternal-associated oral–fecal/vaginal and later breast-feeding routes. Later in life, diet, behavioral activities, and lifestyle have major influences on the bacterial species represented in the microbiome. The interaction between the host immune system and the microbes present can establish a “balance of power” whereby immune inflammatory responses are controlled as well as the composition and expansion of microbial species present, creating a microenvironment beneficial to both host and microbe. Two of the most important routes of host control over the microbiome are the production of pancreatic-derived antimicrobial peptides (AMPs) and the synthesis of secretory IgA [43]. In contrast, the surface proteins and carbohydrates of the bacterial species present stimulate a specific set of Toll-like receptors, while other TLRs are completely absent, down-regulated, or transient in expression. Activation of the TLRs in the context of a healthy microbiome stimulates the concomitant activation of the phosphatidylinositol-3 kinase-AKT pathway, promoting the downstream synthesis of anti-inflammatory mediators. In addition, bacterial species composing a healthy microbiome appear to be adept at stimulating regulatory T lymphocytes, thus promoting immune suppression. Alterations in either of these host-mediated regulatory mechanisms can promote the appearance of a pro-inflammatory microbiome [43]. While healthy eating habits and lifestyle have a significant symbiotic effect on the bacterial species present, and these species tend to promote an anti-inflammatory environment, repressing innate immune inflammation and maintaining the health of the gut epithelium, poor eating habits and lifestyle, as well as antibiotic usage and physical and/or mental stress, alter the microbiome in a negative way. This change in the microbe composition likely relates to both changes in the host relation to the microbiome (i.e., AMPs or other host metabolites synthesized) and the capacity of certain microbial species to better flourish under the altered conditions. The species present and the metabolites produced by this altered microbiome can often promote local inflammation, eliciting a stress response, chronic tissue damage, and disease. In addition to the gut microbiome, commensal microbes have also been observed to infiltrate and colonize other tissues of the gastrointestinal tract, such as the bile ducts, gall bladder, and liver, and to be important for the health of these tissues [44][45].2.4. Innate Immune Control of DNA Damage Repair and Epigentic Modifications
Current data from disease models are beginning to suggest that chronic stresses can actively favor mutagenesis and that the mutations that appear are not such a random happenchance after all, as they appear in particular regions of the genome at rates higher than would be predicted by random happenchance [46]. Thus, if chronic stress leads to chronic inflammation, what better way to control the rate of mutagenesis to favor adaptation than through components of the innate immune system? Other than PKR, a number of additional innate immune/inflammatory and interferon regulated factors have roles in both DNA damage repair and epigenetic modification, including ADAR1, the IFNγ-inducible protein 16 (IFI16), and the three prime repair exonuclease 1 (TREX1) [47][48][49][50][51] (Figure 1b). Moreover, a study by Tassinari et al. demonstrated that N6-adenosine methyltransferase METTL3, an epigenetic modifier shown to have a significant role in the DNA damage response and gametogenesis, methylates ADAR1 mRNA, enhancing ADAR1 protein synthesis [52][53][54]. The link between innate immune/inflammatory stress and DNA repair/modification may suggest that under conditions of constitutive/chronic stress, the immune response opens the door to a potential escape mechanism, allowing for enhanced mutation rates in the off-chance that an adaptation to the stress may be achieved [41][47][50][55]. In an individual organism this may occur throughout one’s life, but how could such a mechanism favor transmission to subsequent generations? The effect of stress on offspring has long been reported in models such as fruitflies, C.elegans, and even birds [56][57][58], but recent data suggest that stress and inflammation also influence the methylation and epigenetic modification of gamete DNA, especially that of sperm [59][60][61]. In addition, stress and inflammation are bound to affect the mother and the environment of the developing fetus. So, how could such selective mutagenesis result in degenerative diseases? In general, progression of degenerative disease is not measured by clonal evolution of cells in the affected tissue but by the degree of tissue destruction and patient scores from performance tests. This is not to say that clonal selection does not take place, it does, but almost strictly in cells belonging to the adaptive immune response (T and B cells) [62]. In pathological tissue specimens from patients with various degenerative diseases, the T cell population is pro-inflammatory. Moreover, in some cases there is a tendency for the appearance of B cells producing disease specific autoantibodies [63][64]. As regards the innate immune system, autoinflammatory and degenerative diseases tend to have an altered type I interferon response (IFR) or alterations in proteins belonging to the IFR [12][64][65][66]. Finally, an accumulation of data from SARS-CoV2 studies, as well as studies into AD, PD, and idopathic myocitis, suggest a role of altered PI3K/AKT/GSK3 signaling affecting the innate immune/interferon response in these pathologies [67].References
- Granger, D.N.; Senchenkova, E. Inflammation and the Microcirculation; Morgan & Claypool Life Sciences Publisher: San Rafael, CA, USA, 2010.
- Gasteiger, G.; D’Osualdo, A.; Schubert, D.A.; Weber, A.; Bruscia, E.M.; Hartl, D. Cellular Innate Immunity: An Old Game with New Players. J. Innate Immun. 2017, 9, 111–125.
- Takeuchi, O.; Akira, S. Pattern recognition receptors and inflammation. Cell 2010, 140, 805–820.
- Prantner, D.; Nallar, S.; Vogel, S.N. The role of RAGE in host pathology and crosstalk between RAGE and TLR4 in innate immune signal transduction pathways. FASEB J. 2020, 34, 15659–15674.
- Rehwinkel, J.; Gack, M.U. RIG-I-like receptors: Their regulation and roles in RNA sensing. Nat. Rev. Immunol. 2020, 20, 537–551.
- Gan, Y.; Li, X.; Han, S.; Liang, Q.; Ma, X.; Rong, P.; Wang, W.; Li, W. The cGAS/STING Pathway: A Novel Target for Cancer Therapy. Front. Immunol. 2021, 12, 795401.
- Schroder, K.; Tschopp, J. The inflammasomes. Cell 2010, 140, 821–832.
- Gal-Ben-Ari, S.; Barrera, I.; Ehrlich, M.; Rosenblum, K. PKR: A Kinase to Remember. Front. Mol. Neurosci. 2018, 11, 480.
- Ito, T.; Yang, M.; May, W.S. RAX, a cellular activator for double-stranded RNA-dependent protein kinase during stress signaling. J. Biol. Chem. 1999, 274, 15427–15432.
- Patel, R.C.; Sen, G.C. PACT, a protein activator of the interferon-induced protein kinase, PKR. EMBO J. 1998, 17, 4379–4390.
- Blalock, W.L. Opposing forces fight over the same ground to regulate interferon signaling. Biochem. J. 2021, 478, 1853–1859.
- Piazzi, M.; Bavelloni, A.; Faenza, I.; Blalock, W. Glycogen synthase kinase (GSK)-3 and the double-strand RNA-dependent kinase, PKR: When two kinases for the common good turn bad. Biochim. Et Biophys. Acta. Mol. Cell Res. 2020, 1867, 118769.
- Lu, B.; Nakamura, T.; Inouye, K.; Li, J.; Tang, Y.; Lundback, P.; Valdes-Ferrer, S.I.; Olofsson, P.S.; Kalb, T.; Roth, J.; et al. Novel role of PKR in inflammasome activation and HMGB1 release. Nature 2012, 488, 670–674.
- Li, D.; Wu, M. Pattern recognition receptors in health and diseases. Signal Transduct. Target. Ther. 2021, 6, 291.
- Kok, K.H.; Lui, P.Y.; Ng, M.H.; Siu, K.L.; Au, S.W.; Jin, D.Y. The double-stranded RNA-binding protein PACT functions as a cellular activator of RIG-I to facilitate innate antiviral response. Cell Host Microbe 2011, 9, 299–309.
- Bennett, R.L.; Blalock, W.L.; Abtahi, D.M.; Pan, Y.; Moyer, S.A.; May, W.S. RAX, the PKR activator, sensitizes cells to inflammatory cytokines, serum withdrawal, chemotherapy, and viral infection. Blood 2006, 108, 821–829.
- Sokol, C.L.; Luster, A.D. The chemokine system in innate immunity. Cold Spring Harb. Perspect. Biol. 2015, 7, a016303.
- Eiz-Vesper, B.; Schmetzer, H.M. Antigen-Presenting Cells: Potential of Proven und New Players in Immune Therapies. Transfus. Med. Hemother. 2020, 47, 429–431.
- Kambayashi, T.; Laufer, T.M. Atypical MHC class II-expressing antigen-presenting cells: Can anything replace a dendritic cell? Nat. Rev. Immunol. 2014, 14, 719–730.
- Dong, C.; Flavell, R.A. Cell fate decision: T-helper 1 and 2 subsets in immune responses. Arthritis Res. 2000, 2, 179–188.
- Uribe-Querol, E.; Rosales, C. Phagocytosis: Our Current Understanding of a Universal Biological Process. Front. Immunol. 2020, 11, 1066.
- Ivashkiv, L.B.; Donlin, L.T. Regulation of type I interferon responses. Nat. Rev. Immunol. 2014, 14, 36–49.
- Forrester, S.J.; Kikuchi, D.S.; Hernandes, M.S.; Xu, Q.; Griendling, K.K. Reactive Oxygen Species in Metabolic and Inflammatory Signaling. Circ. Res. 2018, 122, 877–902.
- Liu, T.; Zhang, L.; Joo, D.; Sun, S.C. NF-kappaB signaling in inflammation. Signal Transduct. Target. Ther. 2017, 2, e17023.
- Liu, T.; Zhou, Y.; Ko, K.S.; Yang, H. Interactions between Myc and Mediators of Inflammation in Chronic Liver Diseases. Mediat. Inflamm. 2015, 2015, 276850.
- Bae, S.; Park, P.S.U.; Lee, Y.; Mun, S.H.; Giannopoulou, E.; Fujii, T.; Lee, K.P.; Violante, S.N.; Cross, J.R.; Park-Min, K.H. MYC-mediated early glycolysis negatively regulates proinflammatory responses by controlling IRF4 in inflammatory macrophages. Cell Rep. 2021, 35, 109264.
- Turner, M.D.; Nedjai, B.; Hurst, T.; Pennington, D.J. Cytokines and chemokines: At the crossroads of cell signalling and inflammatory disease. Biochim. Et Biophys. Acta 2014, 1843, 2563–2582.
- Cheroni, C.; Manganaro, L.; Donnici, L.; Bevilacqua, V.; Bonnal, R.J.P.; Rossi, R.L.; De Francesco, R. Novel interferon-sensitive genes unveiled by correlation-driven gene selection and systems biology. Sci. Rep. 2021, 11, 18043.
- Uzhachenko, R.V.; Shanker, A. CD8(+) T Lymphocyte and NK Cell Network: Circuitry in the Cytotoxic Domain of Immunity. Front. Immunol. 2019, 10, 1906.
- Muntjewerff, E.M.; Meesters, L.D.; van den Bogaart, G. Antigen Cross-Presentation by Macrophages. Front. Immunol. 2020, 11, 1276.
- Belizario, J.E.; Brandao, W.; Rossato, C.; Peron, J.P. Thymic and Postthymic Regulation of Naive CD4(+) T-Cell Lineage Fates in Humans and Mice Models. Mediat. Inflamm. 2016, 2016, 9523628.
- Van Niekerk, G.; Davids, L.M.; Hattingh, S.M.; Engelbrecht, A.M. Cancer stem cells: A product of clonal evolution? Int. J. Cancer 2017, 140, 993–999.
- Savage, S.A.; Dufour, C. Classical inherited bone marrow failure syndromes with high risk for myelodysplastic syndrome and acute myelogenous leukemia. Semin. Hematol. 2017, 54, 105–114.
- Blalock, W.L.; Bavelloni, A.; Piazzi, M.; Faenza, I.; Cocco, L. A role for PKR in hematologic malignancies. J. Cell. Physiol. 2010, 223, 572–591.
- Piazzi, M.; Bavelloni, A.; Gallo, A.; Faenza, I.; Blalock, W.L. Signal Transduction in Ribosome Biogenesis: A Recipe to Avoid Disaster. Int. J. Mol. Sci. 2019, 20, 2718.
- Fisher, D.A.C.; Fowles, J.S.; Zhou, A.; Oh, S.T. Inflammatory Pathophysiology as a Contributor to Myeloproliferative Neoplasms. Front. Immunol. 2021, 12, 683401.
- Zhang, X.; Li, J.; Sejas, D.P.; Rathbun, K.R.; Bagby, G.C.; Pang, Q. The Fanconi anemia proteins functionally interact with the protein kinase regulated by RNA (PKR). J. Biol. Chem. 2004, 279, 43910–43919.
- Follo, M.Y.; Finelli, C.; Mongiorgi, S.; Clissa, C.; Bosi, C.; Martinelli, G.; Blalock, W.L.; Cocco, L.; Martelli, A.M. PKR is activated in MDS patients and its subcellular localization depends on disease severity. Leukemia 2008, 22, 2267–2269.
- Blalock, W.L.; Grimaldi, C.; Fala, F.; Follo, M.; Horn, S.; Basecke, J.; Martinelli, G.; Cocco, L.; Martelli, A.M. PKR activity is required for acute leukemic cell maintenance and growth: A role for PKR-mediated phosphatase activity to regulate GSK-3 phosphorylation. J. Cell. Physiol. 2009, 221, 232–241.
- Blalock, W.L.; Bavelloni, A.; Piazzi, M.; Tagliavini, F.; Faenza, I.; Martelli, A.M.; Follo, M.Y.; Cocco, L. Multiple forms of PKR present in the nuclei of acute leukemia cells represent an active kinase that is responsive to stress. Leukemia 2011, 25, 236–245.
- Cheng, X.; Byrne, M.; Brown, K.D.; Konopleva, M.Y.; Kornblau, S.M.; Bennett, R.L.; May, W.S. PKR inhibits the DNA damage response, and is associated with poor survival in AML and accelerated leukemia in NHD13 mice. Blood 2015, 126, 1585–1594.
- Melamed, D.; Nov, Y.; Malik, A.; Yakass, M.B.; Bolotin, E.; Shemer, R.; Hiadzi, E.K.; Skorecki, K.L.; Livnat, A. De novo mutation rates at the single-mutation resolution in a human HBB gene region associated with adaptation and genetic disease. Genome Res. 2022, 32, 488–498.
- Zheng, D.; Liwinski, T.; Elinav, E. Interaction between microbiota and immunity in health and disease. Cell Res. 2020, 30, 492–506.
- Suppli, M.P.; Bagger, J.I.; Lelouvier, B.; Broha, A.; Demant, M.; Konig, M.J.; Strandberg, C.; Lund, A.; Vilsboll, T.; Knop, F.K. Hepatic microbiome in healthy lean and obese humans. JHEP Rep. 2021, 3, 100299.
- Molinero, N.; Ruiz, L.; Milani, C.; Gutierrez-Diaz, I.; Sanchez, B.; Mangifesta, M.; Segura, J.; Cambero, I.; Campelo, A.B.; Garcia-Bernardo, C.M.; et al. The human gallbladder microbiome is related to the physiological state and the biliary metabolic profile. Microbiome 2019, 7, 100.
- Monroe, J.G.; Srikant, T.; Carbonell-Bejerano, P.; Becker, C.; Lensink, M.; Exposito-Alonso, M.; Klein, M.; Hildebrandt, J.; Neumann, M.; Kliebenstein, D.; et al. Mutation bias reflects natural selection in Arabidopsis thaliana. Nature 2022, 602, 101–105.
- Shiromoto, Y.; Sakurai, M.; Minakuchi, M.; Ariyoshi, K.; Nishikura, K. ADAR1 RNA editing enzyme regulates R-loop formation and genome stability at telomeres in cancer cells. Nat. Commun. 2021, 12, 1654.
- Ka, N.L.; Lim, G.Y.; Hwang, S.; Kim, S.S.; Lee, M.O. IFI16 inhibits DNA repair that potentiates type-I interferon-induced antitumor effects in triple negative breast cancer. Cell Rep. 2021, 37, 110138.
- Schoggins, J.W.; Rice, C.M. Interferon-stimulated genes and their antiviral effector functions. Curr. Opin. Virol. 2011, 1, 519–525.
- Jimeno, S.; Balestra, F.R.; Huertas, P. The Emerging Role of RNA Modifications in DNA Double-Strand Break Repair. Front. Mol. Biosci. 2021, 8, 664872.
- Tao, S.S.; Wu, G.C.; Zhang, Q.; Zhang, T.P.; Leng, R.X.; Pan, H.F.; Ye, D.Q. TREX1 As a Potential Therapeutic Target for Autoimmune and Inflammatory Diseases. Curr. Pharm. Des. 2019, 25, 3239–3247.
- Tassinari, V.; Cesarini, V.; Tomaselli, S.; Ianniello, Z.; Silvestris, D.A.; Ginistrelli, L.C.; Martini, M.; De Angelis, B.; De Luca, G.; Vitiani, L.R.; et al. ADAR1 is a new target of METTL3 and plays a pro-oncogenic role in glioblastoma by an editing-independent mechanism. Genome Biol. 2021, 22, 51.
- Lin, Z.; Hsu, P.J.; Xing, X.; Fang, J.; Lu, Z.; Zou, Q.; Zhang, K.J.; Zhang, X.; Zhou, Y.; Zhang, T.; et al. Mettl3-/Mettl14-mediated mRNA N(6)-methyladenosine modulates murine spermatogenesis. Cell Res. 2017, 27, 1216–1230.
- Zhang, C.; Chen, L.; Peng, D.; Jiang, A.; He, Y.; Zeng, Y.; Xie, C.; Zhou, H.; Luo, X.; Liu, H.; et al. METTL3 and N6-Methyladenosine Promote Homologous Recombination-Mediated Repair of DSBs by Modulating DNA-RNA Hybrid Accumulation. Mol. Cell 2020, 79, 425–442.e427.
- Jimeno, S.; Prados-Carvajal, R.; Fernandez-Avila, M.J.; Silva, S.; Silvestris, D.A.; Endara-Coll, M.; Rodriguez-Real, G.; Domingo-Prim, J.; Mejias-Navarro, F.; Romero-Franco, A.; et al. ADAR-mediated RNA editing of DNA:RNA hybrids is required for DNA double strand break repair. Nat. Commun. 2021, 12, 5512.
- Geffroy, B.; Douhard, M. The Adaptive Sex in Stressful Environments. Trends Ecol. Evol. 2019, 34, 628–640.
- Kishimoto, S.; Uno, M.; Okabe, E.; Nono, M.; Nishida, E. Environmental stresses induce transgenerationally inheritable survival advantages via germline-to-soma communication in Caenorhabditis elegans. Nat. Commun. 2017, 8, 14031.
- Cheng, C.; Kirkpatrick, M. Environmental Plasticity in the Intersexual Correlation and Sex Bias of Gene Expression. J. Hered. 2017, 108, 754–758.
- Sharma, U. Paternal Contributions to Offspring Health: Role of Sperm Small RNAs in Intergenerational Transmission of Epigenetic Information. Front. Cell Dev. Biol. 2019, 7, 215.
- Donkin, I.; Barres, R. Sperm epigenetics and influence of environmental factors. Mol. Metab. 2018, 14, 1–11.
- Marcho, C.; Oluwayiose, O.A.; Pilsner, J.R. The preconception environment and sperm epigenetics. Andrology 2020, 8, 924–942.
- Yang, Q.; Wang, G.; Zhang, F. Role of Peripheral Immune Cells-Mediated Inflammation on the Process of Neurodegenerative Diseases. Front. Immunol. 2020, 11, 582825.
- Sim, K.Y.; Im, K.C.; Park, S.G. The Functional Roles and Applications of Immunoglobulins in Neurodegenerative Disease. Int. J. Mol. Sci. 2020, 21, 5295.
- Piazzi, M.; Bavelloni, A.; Cenni, V.; Faenza, I.; Blalock, W.L. Revisiting the Role of GSK3, A Modulator of Innate Immunity, in Idiopathic Inclusion Body Myositis. Cells 2021, 10, 3255.
- Lin, B.; Goldbach-Mansky, R. Pathogenic insights from genetic causes of autoinflammatory inflammasomopathies and interferonopathies. J. Allergy Clin. Immunol. 2022, 149, 819–832.
- Nazmi, A.; Field, R.H.; Griffin, E.W.; Haugh, O.; Hennessy, E.; Cox, D.; Reis, R.; Tortorelli, L.; Murray, C.L.; Lopez-Rodriguez, A.B.; et al. Chronic neurodegeneration induces type I interferon synthesis via STING, shaping microglial phenotype and accelerating disease progression. Glia 2019, 67, 1254–1276.
- Long, H.Z.; Cheng, Y.; Zhou, Z.W.; Luo, H.Y.; Wen, D.D.; Gao, L.C. PI3K/AKT Signal Pathway: A Target of Natural Products in the Prevention and Treatment of Alzheimer’s Disease and Parkinson’s Disease. Front. Pharmacol. 2021, 12, 648636.
