Nematode-trapping (NT) fungi play a significant role in the biological control of plant- parasitic nematodes. NT fungi, as a predator, can differentiate into specialized structures called “traps” to capture, kill, and consume nematodes at a nutrient-deprived condition. Therefore, trap formation is also an important indicator that NT fungi transition from a saprophytic to a predacious lifestyle. With the development of gene knockout and multiple omics such as genomics, transcriptomics, and metabolomics, increasing studies have tried to investigate the regulation mechanism of trap formation in NT fungi. And elucidating the mechanism of trap formation will help to develop more effective anti-nematode agents by genetic modification.
- traps
- nematode-trapping fungi
- signaling pathways
- signal molecules
- pathway collaboration
- reverse genetics
1. Introduction
Plant-parasitic nematodes (PPNs) can cause direct damage to their host or act as virus vectors [1]. There are more than 4100 species of PPNs that have an impact on global agriculture and horticulture, causing an estimated annual loss of USD 173 billion [2][3]. Nematode-trapping (NT) fungi are potential biocontrol resources, which have the advantages of low toxicity, high efficiency, and environmental friendliness, so they have gradually become favored by people in recent years [3]. NT fungi can develop specialized structures called “traps”, an important indicator of the transition from saprophytic to predatory lifestyles, including constricting rings, adhesive networks, adhesive columns, and adhesive knobs, to capture, kill, and consume nematodes at a nutrient-deprived condition [4][5]. Therefore, trap formation is a crucial step in the lifestyle of NT fungi and indispensable for nematode predation. The factors involved in trap formation were variety, including multiple signal transduction pathways [6]; small molecular compounds [7][8][9][10][11][12]; intercellular communication [13]; adhesive protein [5]; nitrate assimilation [14]; woronin body [15]; peroxisome [16]; autophagy [17][18][19][20]; and pH-sensing receptors [21], according to the comprehensive research of genomics, transcriptomics, proteomics, metabolomics, and reverse genetics. Here, wresearchers review the recent progress in the regulatory mechanism of trap formation in the NT fungi based on phenotypes of various mutants and multi-omics analysis. WeResearchers especially focus on the latest studies in signal transduction pathways and small molecular compounds involved in trap formation. Elucidating the molecular mechanism of trap formation will not only provide a theoretical basis for the improvement of engineering biocontrol fungi, but contribute to understanding the adaptive evolution and lifestyle transition of NT fungi.
2. Multi-Omics Analysis Promotes Research on Trap Formation of NTematode-Trapping Fungi
Omics not only provides a macroscopic direction for the mechanism of trap formation but also provides specific targets. Comparative genomics studies have shown that many species-specific genes have expanded in the evolutionary process, and these genes may be related to the function specialization of NT fungi [5][6][22][23]. Arthrobotrys oligospora (teleomorph Orbilia auricolor), one of the typical NT fungi, was the first NT fungus whose genome and proteome were sequenced [6]. Comparative analysis showed that A. oligospora genome contains numerous pathogenicity-related genes, and a total of 398 homologous genes related to the pathogenicity of other fungi have been identified [6], while there were fewer lectin genes involved in fungus–nematode recognition in the Drechslerella stenobrocha genome [22]. This suggested that a different mechanism of trap formation may exist in various NT fungi. In addition, studies combined genome, proteome, and real-time PCR (RT-PCR) analyses to reveal that multiple signal transduction pathways have an integral part in trap formation [6]. Transcriptome sequencing and RT-PCR analysis showed that a large number of genes were significantly upregulated during the infection process of NT fungi, including genes involved in translation, amino acid metabolism, carbohydrate metabolism, cell wall and membrane biogenesis [6], secreted proteins [23], adhesion proteins [5], and the protein kinase C signal transduction pathway [22]. Comparative genomic analysis in the four representative trapping devices of NT fungi suggested that the simplification of the capture device was accompanied by the expansion of adhesion genes and the increase in adhesiveness on trap surfaces [5]. In conclusion, the above studies show that omics technologies have clarified the general direction for research on the regulation mechanism of traps formation. At the same time, the assembly of genomes in different fungi makes it possible to study the interaction between the NT fungi and nematodes at the molecular level. For instance, according to the genome sequence and annotation of Duddingtonia flagrans, a fluorescent protein system with native promoter was established, and a secretion protein PEFB was identified which was involved in the processes of infection against Caenorhabditis elegans [13]. Recently, nine species of NT fungi have been sequenced, such as adhesive-network-producing fungi A. oligospora and D. flagrans, adhesive-knob-producing fungi Monacrosporium haptotylum and Dactylellina entomopaga, constricting-ring-producing fungi D. stenobrocha and Drechslerella brochopaga, adhesive-columns-producing fungus Dactylellina cionopagum, and no trap producing fungus Dactylella cylindrospora (Table 1); these genomic information may help to elucidate the mechanism of trap formation of NT fungi.
Table 1.
Genomic features of different NT fungi.
|
Trapping Devices |
Fungi |
Genome Size |
GC Content (%) |
Number of Genes |
Reference |
|
Adhesive network |
A. oligospora ATCC24927 |
40.07 Mb |
44.45 |
11,479 |
[6] |
|
Adhesive network |
A. oligospora TWF154 |
39.62 Mb |
43.96 |
12,107 |
[24] |
|
Adhesive network |
D. flagrans |
36.64 Mb |
45.5 |
9927 |
[13] |
|
Adhesive knob |
M. haptotylum |
40.40 Mb |
45.24 |
10,959 |
[23] |
|
Adhesive knob |
D. entomopaga |
38.39 Mb |
44.9 |
11,130 |
[5] |
|
Constricting ring |
D. stenobrocha |
29.02 Mb |
52.5 |
5597 |
[22] |
|
Constricting ring |
D. brochopaga |
35.43 Mb |
49.42 |
10,234 |
[5] |
|
Adhesive column |
D. cionopagum |
43.12 Mb |
44.3 |
11,284 |
[5] |
|
no trapping device |
D. cylindrospora |
37.71 Mb |
46.02 |
10,785 |
[5] |
3. Overview of Signaling Pathways Involved in Trap Formation
3.1. G-Protein Signaling Pathway Involved in Trap Formation
3.2. Mitogen-Activated Protein Kinase (MAPK) Signaling Pathway Is Essential for Trap Formation
3.3. cAMP-Dependent Protein Kinase A (cAMP/PKA) Signaling Pathway Is Indispensable for Trap Formation
3.4. Ca2+-Related Signaling Pathway Regulates Trap Formation
4.Compounds as Signal Molecules to Regulate Trap Formation
There is an evolutionary arms race between predators and prey, and just as prey evolves specific strategies to avoid being hunted, predators also evolve stronger predation strategies, such as lure compounds, to ensure adequate food [48]. Normally, NT fungi are saprophytic, but they will become predators in order to maximize the chance of survival when nutrients are deficient, and their lifestyle transitions accordingly. However, NT fungi are non-motile predators, but nematodes can move at will. Hence, NT fungi evolved an ingenious way, “VOCs”, functioning as mimic pheromone, to lure nematodes. These volatile compounds (VOCs) include dimethyl disulfide, (±)2-methyl-1-butanol, 2,4-dithiapentane, S-methyl thioacetate, and methyl 3-methyl-2-butenoate, and especially, methyl 3-methyl-2-butenoate trigger strong sex- and stage-specific attraction in several Caenorhabditis species. Correspondingly, the olfactory neuron AWCs of C. elegans sensed the odors emanating from NT fungi and responded the attraction [49][7]. Interesting is that the ascarosides, an evolutionarily highly conserved family of small molecules produced by nematodes, downregulated the expression of polyketide synthase gene (artA), which in turn promoted the formation of new traps and then resulted in trapping networks. Therefore, the concentration of ascarosides increased as the nematodes approach, which in turn downregulated arthrosporol and 6-methyl-salicylic acid (6-MSA) formation, further causing the hunting ground to be covered with traps. The ascarosides disappear when the nematodes were completely digested, the contents of arthrosporol and 6-MSA returned to normal levels, and the mycelium switched to saprotrophic growth[50].
Trap formation is a highly energy-consuming process, and to conserve energy, NT fungi have to evolve a more effective strategy to regulate the triggering or closing of trap formation [8]: for example, triggering trap formation after sensing signals of nematodes during infection and terminating trap formation when nematodes were fully digested. Small molecular compounds played an important role in this conversion process, such as ascarosides from nematodes [8], and 6-MSA, oligosporons, arthrobotrisins, and arthrosporols isolated from A. oligospora and other NT fungi [50][51][52][53][54][55]. Recently, an increasing number of compounds and associated synthetic genes have been investigated. For instance, the latest research demonstrated that the chemical diversity of metabolites increased notably and exhibited species specificity in the process of changing lifestyle from saprophytic to predatory in NT fungi A. oligospora, A. thaumasia, and A. musiformis [56]. Volatile furanone and pyrone metabolites can help A. oligospora capture nematodes in the lifestyle transition[12], and abscisic acid was highly effective at enhancing trap formation of D. stenobrocha [57]. In addition, 6-MSA is a chemoattractant that can lure nematodes into the fungal mycelium. The artA expression can produce 6-MSA in hyphal tips, and was uncoupled from other enzymes required for the conversion of 6-MSA to arthrosporols; moreover, corresponding deletion strains produced more traps, suggesting a negative role of 6-MSA on trap formation in D. flagrans [50]. Furthermore, the gene cluster AOL_s00215 plays a key role in the production of arthrosporols in A. oligospora, and the number of traps was increased in deletion mutants of most genes in this gene cluster [10][11][58][59][60]. Arthrobotrisins were downregulated in Δric8, Δras2, and Δrheb mutants, indicating that G-proteins and small GTPases were involved in regulating the metabolism of arthrobotrisins [27][29]. Simultaneously, ammonia could function as a signaling molecule in NT fungi to trigger trap formation and kill nematodes, disrupting the gene involved in urea transport and metabolism, resulting in the abolition of urea-induced trap formation in A. oligospora [61]. Another study also demonstrated that ammonia can induce trap formation as a signal molecule in NT fungi A. oligospora, A. guizhouensis, D. phymatopaga, D. cionopaga, and D. brochopaga [62]. Furthermore, PKS−TPS hybrid pathway, for biosynthesis of sesquiterpenyl epoxy-cyclohexenoids, involved in trap formation via ammonia metabolism, deletion of most genes in the PKS−TPS hybrid pathway displayed significantly increase in trap formation [9]. Overall, the discovery of multiple compounds enriches our knowledge of the inducers in trap formation, which participate in trap formation as signaling molecules.
5. Multiple Cellular Processes Were Involved in Trap Formation
| Fungi | Mutated Genes |
Annotation | Phenotypic Traits | Reference | ||
|---|---|---|---|---|---|---|
| Traps | Conidiation | Mycelial Growth | ||||
| A. oligospora | gpb1 | G-protein β subunit | Y | N | N | [24] |
| A. oligospora | flbA | Regulator of G-protein signaling |
Y | Y | Y | [26] |
| A. oligospora | rgsA | Y | N | N | [26] | |
| A. oligospora | rgsB | Y | Y | Y | [26] | |
| A. oligospora | rgsB2-1 | Y | Y | Y | [26] | |
| A. oligospora | rgsB2-2 | Y | N | N | [26] | |
| A. oligospora | rgsB2-3 | Y | Y | N | [26] | |
| A. oligospora | rgsC | Y | Y | N | [26] | |
| A. oligospora | gas1 | GAS protein | Y | Y | N | [26] |
| A. oligospora | ras2 | RAS GTPase | Y | Y | Y | [29] |
| A. oligospora | ras3 | RAS GTPase | N | N | N | [29] |
| A. oligospora | rheb | RAS GTPase | Y | Y | Y | [29] |
| A. oligospora | rab-7A | RAB GTPase | Y | Y | Y | [28] |
| A. oligospora | rab-2 | RAB GTPase | N | Y | N | [28] |
| A. oligospora | rho2 | RHO GTPase | N | N | N | [30] |
| A. oligospora | rac | RHO GTPase | Y | Y | Y | [30] |
| A. oligospora | cdc42 | RHO GTPase | Y | Y | Y | [30] |
| A. oligospora | pex1 | Peroxisome biogenesis protein | Y | Y | Y | [16] |
| A. oligospora | pex6 | Peroxisome biogenesis protein | Y | Y | Y | [16] |
| A. oligospora | mkk1 | MAPK kinase MKK1 | Y | Y | Y | [37] |
| A. oligospora | ste7 | MAPK kinase STE7 | Y | Y | Y | [4] |
| A. oligospora | fus3 | MAPK FUS3 | Y | Y | Y | [4] |
| A. oligospora | ste12 | Tanscription factor | Y | N | Y | [4] |
| A. oligospora | slt2 | MAPK SLT2 | Y | Y | Y | [34] |
| A. oligospora | hog1 | MAPK HOG1 | Y | Y | N | [38] |
| A. oligospora | msb2 | Mucin protein | Y | N | Y | [38] |
| A. oligospora | ime2 | MAPK IME2 | Y | Y | Y | [41] |
| A. oligospora | bck1 | MAPK kinase kinase BCK1 | Y | Y | Y | [37] |
| A. oligospora | ric8 | Resistance to inhibitors of cholinesterase | Y | Y | Y | [27] |
| A. oligospora | stuA | Transcription factor | Y | Y | Y | [42] |
| A. oligospora | glo3 | ARF GTPase activator | Y | Y | Y | [32] |
| A. oligospora | camk | Ca2+/calmodulin-dependent protein kinases | Y | Y | Y | [43] |
| A. oligospora | ssk1 | Response regulator | Y | Y | Y | [39] |
| A. oligospora A. oligospora |
atg1 atg13 |
Autophagy protein Autophagy protein |
Y N |
Y N |
Y Y |
[20] [20] |
| A. oligospora | atg4 | Autophagy protein | Y | Y | Y | [18] |
| A. oligospora | atg5 | Autophagy protein | Y | Y | Y | [17] |
| A. oligospora | atg8 | Autophagy protein | Y | Y | Y | [19] |
| A. oligospora | hex1 | Woronin body major protein | Y | Y | Y | [15] |
| A. oligospora | gph1 | Glycogen phosphorylase | Y | Y | Y | [72] |
| A. oligospora | noxA | NADPH oxidase | Y | Y | Y | [71] |
| A. oligospora | niaD | Nitrate reductase | Y | - | Y | [14] |
| A. oligospora | niiA | Nitrite reductase | Y | - | Y | [14] |
| A. oligospora | nrtB | Nitrate transporter | Y | - | Y | [14] |
| A. oligospora | nirA | nitrogen assimilation transcription factor | Y | - | Y | [14] |
| A. oligospora | mad1 | Adhesin protein | Y | - | - | [70] |
| A. oligospora | crn1 | Actin cytoskeleton and actin-associated protein | Y | Y | N | [68] |
| A. oligospora | palH | pH sensing receptor | Y | Y | Y | [21] |
| A. oligospora | fig1 | Low-affinity calcium system member | Y | Y | Y | [47] |
| A. oligospora | ubr1 | E3 ubiquitin-protein ligase | Y | - | Y | [63] |
| A. oligospora | vosA | Developmental regulator | N | Y | N | [65] |
| A. oligospora | velB | Developmental regulator | Y | Y | Y | [65] |
| A. oligospora | g276 | Fucose-specific lectin | Y | N | N | [67] |
| A. oligospora | g207 | F-box protein | Y | Y | Y | [64] |
| A. oligospora | AOL_s00215g277 | A putatively cupin-like family gene | Y | Y | N | [10] |
| A. oligospora | AOL_s00215g278 | Cytochrome P450 | Y | Y | Y | [60] |
| A. oligospora | AOL_s00215g279 | Oxidoreductase | Y | Y | Y | [10] |
| A. oligospora | AOL_s00215g280 | Cytochrome P450 | Y | Y | Y | [58] |
| A. oligospora | AOL_s00215g281 | Amidohydrolase | Y | N | Y | [11] |
| A. oligospora | AOL_s00215g282 | Cytochrome P450 oxidoreductase | Y | Y | Y | [11] |
| A. oligospora | AOL_s00215g283 | 6-methylsalicylic acid synthase | Y | - | N | [59] |
| A. oligospora | AOL_s00079g496 | Polyketide synthase | Y | Y | Y | [12] |
| D. flagrans | artA | Polyketide synthase | Y | - | N | [50] |
| D. flagrans | artB | Cytochrome P450 | Y | - | N | [50] |
| D. flagrans | artC | Amidohydrolase | N | - | N | [50] |
| D. flagrans | artD | Cytochrome P450 | Y | - | N | [50] |
| D. flagrans | sofT | Hyphal anastomosis gene | Y | - | Y | [13] |
| D. flagrans | sipC | STRIPAK complex component | Y | Y | Y | [69] |
| D. dactyloides | ste12 | Transcription factor | Y | Y | Y | [40] |
| M. haptotylum | slt2 | MAPK SLT2 | Y | Y | Y | [34] |
6.Summary and Perspectives
6.1. Multiple Signaling Pathways and Cellular Processes Co-Regulate Trap Formation
The development of multi-omics contributes to elucidating the mechanism of trap formation. Comparative genomics of nine NT fungi including two A. oligospora with different genetic backgrounds revealed commonalities and specificities of trap formation among different NT fungi [5][6][13][22][23][24]. As previously mentioned, the results of comparative genomics and multi-omics consistently showed that signal transduction pathways were closely related to the morphogenetic triggering process of traps. As a representative species of NT fungi, the formation mechanism of traps has been extensively investigated in A. oligospora, and trap formation is a complex process, which likely requires the coordination of multiple pathways. This was demonstrated by changes in the expression patterns of relevant genes in various mutant strains and interaction between some proteins (Table 2).
It is the nature of predators to sense and respond to prey. The G-protein signaling pathway was essential for the signal reception and transduction of nematodes in NT fungi. Deletion of gpb1 resulted in a remarkable reduction in trap formation [24]. Phosphorylation assay showed that phosphorylation signals were lost in both Δgpb1 and Δfus3 mutants upon nematode induction, suggesting that gpb1 may function upstream of fus3 [4]. RGSs negatively regulated G-protein signal transduction, and in the Δrgs mutants, the intracellular cAMP levels were significantly increased and the transcription levels of G-protein signaling genes were downregulated [26]. In addition, RIC8 regulated the cAMP level by interacting with Gα1, thereby participating in the fungal growth, environmental adaptation, and trap formation [27]. These findings show that G-protein regulates the cAMP/PKA and MAPK signaling pathways under the regulation of RGSs and other regulators.
In addition, the small GTPases family acts as switches in the signaling hub of molecular circuits [76]. A recent study showed that the expression levels of genes encoding regulatory subunits of PKA, MAPK, and P21-activated kinases were downregulated in Δcdc42 and Δrac mutants; thus, PKA, P21-activated kinases, and HOG1 may be downstream effectors of RHO GTPases in A. oligospora [30]. Similarly, RAS GTPases affected MAPK signaling by interacting with STE50 and directly regulating intracellular cAMP levels and the mTOR signaling pathway [29]. These indicate that small GTPases can directly or indirectly regulate MAPK, TOR, and cAMP/PKA signaling pathways. Transcription factors located downstream of these signaling pathways activate or repress the transcription of corresponding genes after receiving cascaded signals, thereby regulating the biological response of cells to external signals. STUA functions downstream of the cAMP/PKA signaling pathway, and the expression of genes involved in the G-protein signaling pathway were transcriptionally repressed in the mutants [42]. Meanwhile, reverse genetics demonstrated that STE12, which regulated the expression of “nematode-responsive genes” that trigger trap formation, was activated by the MAPK cascade in A. oligospora. GO enrichment analysis of STE12-dependent differentially expressed genes revealed that the two most significant terms were “integral component of membrane” and “oxidation–reduction process”, which were critical for hyphal growth and virulence in the NT fungi [4]. These results indicated that multiple signaling pathways co-regulate trap formation.
What can be clearly seen in recent studies is the deletion of almost all of these genes affected redox processes [26][27][28][30][34][37][42][71] and partly influence heat shock response [29], cell walls [26][27][28][32][34][41][42], and mitochondria [29][30], indicating the genes related to these processes might be involved in trap formation, which was confirmed in our latest research (unpublished). At the same time, each gene does not influence the trap formation alone: They need to cooperate with each other (Table 2).
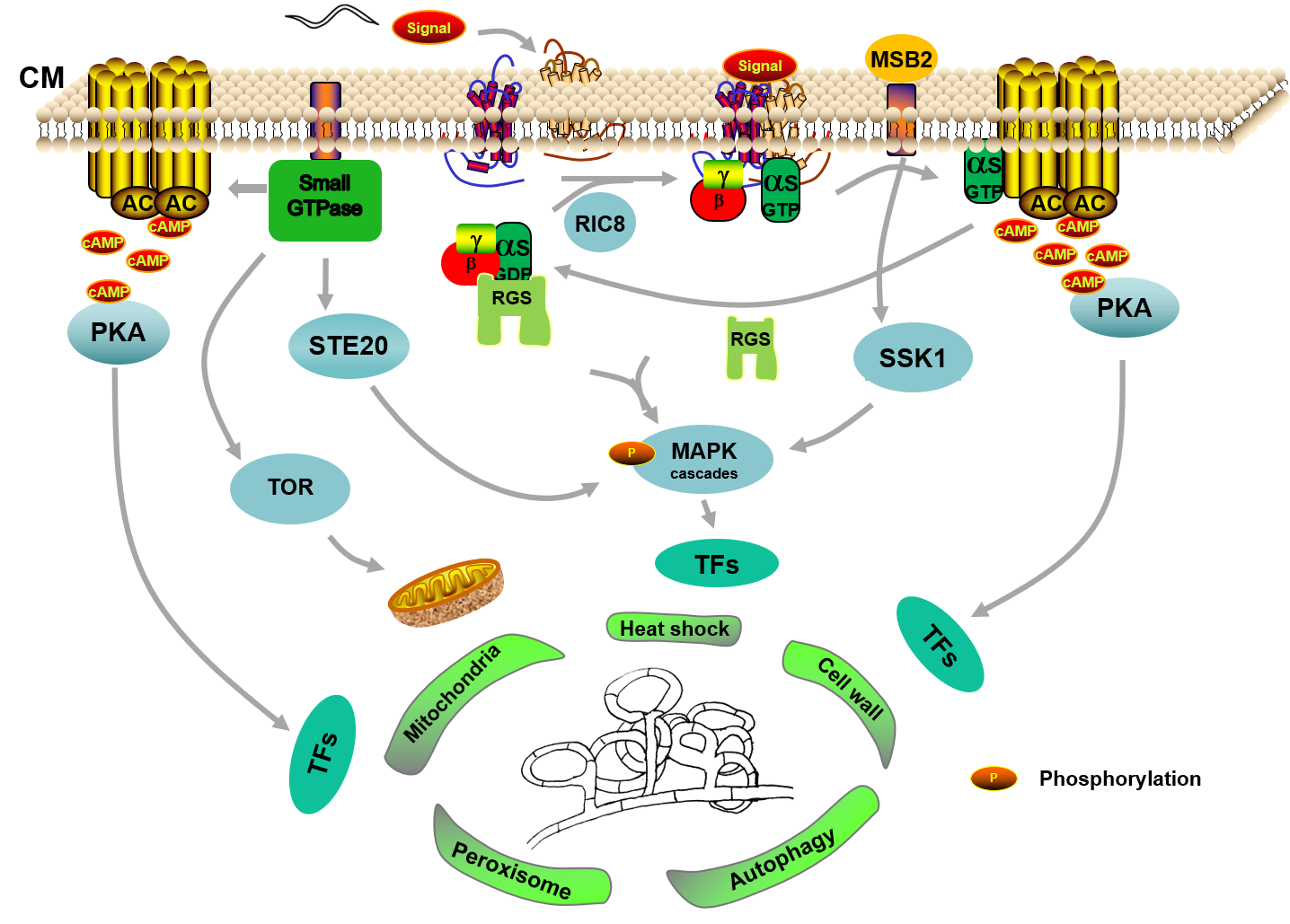
Collectively, given the recent progress in reverse genetics, knowledge obtained from different pathways based on the multi-omics and RT-PCR would contribute significantly to elucidating the specific mechanism of the trap formation, which we are only just beginning to understand. In general, it seems that GPCRs are the receptors to respond the nematode’s signals, and the G-protein signaling pathway accurately transform extracellular signals into intracellular signals with the regulation of RGSs, and then transfers to downstream cascade pathways, such as the cAMP/PKA pathway. Simultaneously, the small GTPase family also regulates the MAPK cascades by interacting with STE20 protein kinases. Subsequently, these cascade pathways further regulate a variety of cellular processes by regulating downstream transcription factors, thereby triggering trap formation (Figure 1).
6.2. The Regulatory Mechanism of Trap Formation May Vary in NT Fungi
Absolutely, it is not difficult to understand that there are differences in the mechanisms of traps formation due to the differences in the morphologies of traps and predatory ways. Comparative genomics of NT fungi with different types of traps have indicated that many species-specific genes and highly expressed uncharacterized orphan genes have expanded during evolution[5][6][22][23][77]. The whole genome blast analysis and the distribution analysis of orthologous gene showed that the putative pathogen–host interaction genes included 80 specific genes in A. oligospora. Furthermore, compared with other pathogenic and non-pathogenic fungi, A. oligospora contained 53.64% abundant orphan genes [6]. Another study showed that 20% of the genes in the M. haptotylum and A. oligospora genomes were shared, but importantly, as many as 16% in each genome genes were unique. Moreover, 15 species-specific genes, unique for the M. haptotylum, were found in the cohort of the 10-fold upregulated transcripts during infection [23]. Comparative analysis found that 12 multigene families and 23 gene domains were significantly expanded in the NT fungal genomes. Notably, nine of the expanded multigene families lacked significant matches in public databases, suggesting the existence of a potential novel mechanism underlying trap formation [5]. In addition, the traps of A. oligospora and most NT fungi must be induced, but the constricting rings of D. stenobrocha can form spontaneously, and its genome was more compact than that of A. oligospora with rare repeat-induced point mutation and transposon [22]. Therefore, a more suitable interpretation is that these may be involved in the functional specialization of different NT fungi, which means that the mechanism of trap formation may differ among various NT fungi.
6.3. Current Status and Future Prospects
The completion of genome sequencing is a milestone for the study of the mechanism trap formation and interaction between the NT fungi and nematodes. Although the genomes of nine NT fungi, including two A. oligospora with different genetic backgrounds, have been sequenced [5][6][13][22][23][24], research on other NT fungi is still in its infancy, except for A. oligospora and D. flagrans. Differences in trap formation between diverse NT fungi remain an interesting topic for further study. There has been considerable progress in the mechanism of trap formation with the development of multi-omics and molecular biology techniques; however, more knowledge still needs to be accumulated for parsing more comprehensive connections in multiple pathways and organelles during trap formation. According to the phenotypes of these gene mutants, it can be found that mitochondria [29][30], peroxisomes [16], nucleus [17][30][39][46], woronin bodies [15], autophagosomes [17][18][19][20], and other organelles [39][46] have changed in varying degrees. So, here, the question arises: How do they cooperate with each other in trap formation? Are they actually involved in trap formation, or are these changes simply due to defects in growth?
NT fungi are important biocontrol resources, and the traps are the key devices to capture nematodes, which will be formed after receiving external signals. Small molecule compounds act as this signal, such as ascarosides [8], abscisic acid [57], ammonia [61][62], and 6-MSA [50], and arthrosporols [50][51][52][53][54][55] were also involved in regulating trap formation. However, the metabolomic analysis showed that a large number of compounds changed during trap formation and nematode infection [27][29][39][46][56]. So, what these compounds are and how they affect trap formation require further study. Could other compounds from nematodes or the environment also affect trap formation? Moreover, signaling pathways have been identified to be involved in trap formation in multiple NT fungi [4][26][27][28][29][30][32][34][37][38][39][41][43][48][78][79], whereas there is a lack of studies about downstream transcription factors, and further research is needed on the functions of upstream receptors located in the cell wall and membrane.
In summary, increasing knowledge about the regulation mechanism of trap formation of NT fungi has been acquired from the phenotypic traits of various mutants and multi-omics analysis in recent years, whereas elucidating the molecular mechanism of trap formation is still ongoing due to the complexity of trap production by multitudinous NT fungi. Discussing current pitfalls and future priorities of trap formation provides a solid basis for elucidating the mechanisms of trap formation and lifestyle transition of NT fungi, and will help to develop more effective anti-nematode agents by genetic modification.
Funding: This research described here is supported by the National Natural Science Foundation of China (no. 31960556), and the Applied Basic Research Foundation of Yunnan Province (no. 202001BB050004).
Institutional Review Board Statement: Not applicable.
Informed Consent Statement: Not applicable.
Data Availability Statement: Not applicable.
Conflicts of Interest: The authors declare no conflict of interest.
References
- Nicol, J.M.; Turner, S.J.; Coyne, D.L.; Den Nijs, L.D. ; Hockland, S.; Maafi, Z.T. Current nematode threats to world agriculture. In Genomics and Molecular Genetics of Plant-Nematode Interactions; Jones, J., Gheysen, G., Fenoll, C., Eds.; Springer: Dordrecht, The Netherlands, 2011; pp. 21–43.
- Elling, A.A. Major emerging problems with minor Meloidogyne species. Phytopathology 2013, 103, 1092–1102.
- Phani, V.; Khan, M.R.; Dutta, T.K. Plant-parasitic nematodes as a potential threat to protected agriculture: Current status and management options. Crop Prot. 2021, 144, 105573.
- Chen, S.A.; Lin, H.C.; Schroeder, F.C.; Hsueh, Y.P. Prey sensing and response in a nematode-trapping fungus is governed by the MAPK pheromone response pathway. Genetics 2021, 217, iyaa008.
- Ji, X.; Yu, Z.; Yang, J.; Xu, J.; Zhang, Y.; Liu, S.; Zou, C.; Li, J.; Liang, L.; Zhang, K.Q. Expansion of adhesion genes drives pathogenic adaptation of nematode-trapping fungi. IScience 2020, 23, 101057.
- Yang, J.; Wang, L.; Ji, X.; Feng, Y.; Li, X.; Zou, C.; Xu, J.; Ren, Y.; Mi, Q.; Wu, J.; et al. Genomic and proteomic analyses of the fungus Arthrobotrys oligospora provide insights into nematode-trap formation. PLoS Pathog. 2011, 7, e1002179.
- Hsueh, Y.P.; Gronquist, M.R.; Schwarz, E.M.; Nath, R.D.; Lee, C.H.; Gharib, S.; Schroeder, F.C.; Sternberg, P.W. Nema-tophagous fungus Arthrobotrys oligospora mimics olfactory cues of sex and food to lure its nematode prey. eLife 2017, 6, e20023.
- Hsueh, Y.P.; Mahanti, P.; Schroeder, F.C.; Sternberg, P.W. Nematode-trapping fungi eavesdrop on nematode pheromones. Curr. Biol. 2013, 23, 83–86.
- He, Z.Q.; Wang, L.J.; Wang, Y.J.; Chen, Y.H.; Wen, Y.; Zhang, K.Q.; Niu, X.M. Polyketide synthase-terpenoid synthase hybrid pathway regulation of trap formation through ammonia metabolism controls soil colonization of predominant nema-tode-trapping fungus. J. Agric. Food Chem. 2021, 69, 4464–4479.
- Chen, Y.H.; Liu, X.; Dai, R.; Ou, X.; Xu, Z.F.; Zhang, K.Q.; Niu, X.M. Novel polyketide-terpenoid hybrid metabolites and increased fungal nematocidal ability by disruption of genes 277 and 279 in nematode-trapping fungus Arthrobotrys oligospora. J. Agric. Food Chem. 2020, 68, 7870–7879.
- Xu, Z.F.; Chen, Y.H.; Song, T.Y.; Zeng, Z.J.; Yan, N.; Zhang, K.Q.; Niu, X.M. Nematicidal key precursors for the biosynthesis of morphological regulatory arthrosporols in the nematode-trapping fungus Arthrobotrys oligospora. J. Agric. Food Chem. 2016, 64, 7949–7956.
- Wang, B.L.; Chen, Y.H.; He, J.N.; Xue, H.X.; Yan, N.; Zeng, Z.J.; Bennett, J.W.; Zhang, K.Q.; Niu, X.M. Integrated metabolomics and morphogenesis reveal volatile signaling of the nematode-trapping fungus Arthrobotrys oligospora. Appl. Environ. Microbiol. 2018, 84, e02749-17.
- Youssar, L.; Wernet, V.; Hensel, N.; Yu, X.; Hildebrand, H.G.; Schreckenberger, B.; Kriegler, M.; Hetzer, B.; Frankino, P.; Dillin, A.; et al. Intercellular communication is required for trap formation in the nematode-trapping fungus Duddingtonia flagrans. PLoS Genet. 2019, 15, e1008029.
- Liang, L.; Liu, Z.; Liu, L.; Li, J.; Gao, H.; Yang, J.; Zhang, K.Q. The nitrate assimilation pathway is involved in the trap formation of Arthrobotrys oligospora, a nematode-trapping fungus. Fungal Genet. Biol. 2016, 92, 33–39.
- Liang, L.; Gao, H.; Li, J.; Liu, L.; Liu, Z.; Zhang, K.Q. The Woronin body in the nematophagous fungus Arthrobotrys oligospora is essential for trap formation and efficient pathogenesis. Fungal Biol. 2017, 121, 11–20.
- Liu, Q.; Li, D.; Jiang, K.; Zhang, K.Q.; Yang, J. AoPEX1 and AoPEX6 are required for mycelial growth, conidiation, stress response, fatty acid utilization, and trap formation in Arthrobotrys oligospora. Microbiol. Spectr. 2022, 10 (2) , e0027522
- Zhou, D.; Zhu, Y.; Bai, N.; Yang, L.; Xie, M.; Yang, J.; Zhu, M.; Zhang, K.Q.; Yang, J. AoATG5 plays pleiotropic roles in vegetative growth, cell nucleus development, conidiation, and virulence in the nematode-trapping fungus Arthrobotrys oli-gospora. Sci. China Life Sci. 2021, 65, 412–425.
- Zhou, D.; Xie, M.; Bai, N.; Yang, L.; Zhang, K.Q.; Yang, J. The autophagy-related gene Aolatg4 regulates hyphal growth, sporulation, autophagosome formation, and pathogenicity in Arthrobotrys oligospora. Front. Microbiol. 2020, 11, 592524.
- Chen, Y.L.; Gao, Y.; Zhang, K.Q.; Zou, C.G. Autophagy is required for trap formation in the nematode-trapping fungus Arthrobotrys oligospora. Environ. Microbiol. Rep. 2013, 5, 511–517.
- Zhou, D.; Zhu, Y.; Bai, N.; Xie, M.; Zhang, K.Q.; Yang, J. Aolatg1 and Aolatg13 regulate autophagy and play different roles in conidiation, trap formation, and pathogenicity in the nematode-trapping fungus Arthrobotrys oligospora. Front. Cell Infect. Microbiol. 2022, 11, 824407.
- Li, J.; Wu, R.; Wang, M.; Borneman, J.; Yang, J.; Zhang, K.Q. The pH sensing receptor AopalH plays important roles in the nematophagous fungus Arthrobotrys oligospora. Fungal Biol. 2019, 123, 547–554.
- Liu, K.; Zhang, W.; Lai, Y.; Xiang, M.; Wang, X.; Zhang, X.; Liu, X. Drechslerella stenobrocha genome illustrates the mechanism of constricting rings and the origin of nematode predation in fungi. BMC Genom. 2014, 15, 114 .
- Meerupati, T.; Andersson, K.M.; Friman, E.; Kumar, D.; Tunlid, A.; Ahren, D. Genomic mechanisms accounting for the adaptation to parasitism in nematode-trapping fungi. PLoS Genet. 2013, 9, e1003909.
- Yang, C.T.; de Ulzurrun, G.V.D.; Goncalves, A.P.; Lin, H.C.; Chang, C.W.; Huang, T.Y.; Chen, S.A.; Lai, C.K.; Tsai, I.J.; Schroeder, F.C.; et al. Natural diversity in the predatory behavior facilitates the establishment of a robust model strain for nematode-trapping fungi. Proc. Natl. Acad. Sci. USA 2020, 117, 6762–6770.
- Li, L.; Wright, S.J.; Krystofova, S.; Park, G.; Borkovich, K.A. Heterotrimeric G protein signaling in filamentous fungi. Annu. Rev. Microbiol. 2007, 61, 423–452.
- Ma, N.; Zhao, Y.; Wang, Y.; Yang, L.; Li, D.; Yang, J.; Jiang, K.; Zhang, K.Q.; Yang, J. Functional analysis of seven regulators of G protein signaling (RGSs) in the nematode-trapping fungus Arthrobotrys oligospora. Virulence 2021, 12, 1825–1840.
- Bai, N.; Zhang, G.; Wang, W.; Feng, H.; Yang, X.; Zheng, Y.; Yang, L.; Xie, M.; Zhang, K.Q.; Yang, J. Ric8 acts as a regulator of G-protein signalling required for nematode-trapping lifecycle of Arthrobotrys oligospora. Environ. Microbiol. 2021, in press.
- Yang, X.; Ma, N.; Yang, L.; Zheng, Y.; Zhen, Z.; Li, Q.; Xie, M.; Li, J.; Zhang, K.Q.; Yang, J. Two Rab GTPases play different roles in conidiation, trap formation, stress resistance, and virulence in the nematode-trapping fungus Arthrobotrys oligospora. Appl. Microbiol. Biotechnol. 2018, 102, 4601–4613.
- Yang, L.; Li, X.M.; Xie, M.H.; Bai, N.; Yang, J.L.; Jiang, K.X.; Zhang, K.Q.; Yang, J.K. Pleiotropic roles of Ras GTPases in the nematode-trapping fungus Arthrobotrys oligospora identified through multi-omics analyses. IScience 2021, 24, 102820.
- Yang, L.; Li, X.; Bai, N.; Yang, X.; Zhang, K.Q.; Yang, J. Transcriptomic analysis reveals that Rho GTPases regulate trap development and lifestyle transition of the nematode-trapping fungus Arthrobotrys oligospora. Microbiol. Spectr. 2022, 10, e0175921.
- Nie, Z.Z.; Randazzo, P.A. Arf GAPs and membrane traffic. J. Cell Sci. 2006, 119, 1203–1211.
- Ma, Y.; Yang, X.; Xie, M.; Zhang, G.; Yang, L.; Bai, N.; Zhao, Y.; Li, D.; Zhang, K.Q.; Yang, J. The Arf-GAP AoGlo3 regulates conidiation, endocytosis, and pathogenicity in the nematode-trapping fungus Arthrobotrys oligospora. Fungal Genet. Biol. 2020, 138, 103352.
- De Ulzurrun, G.V.D.; Hsueh, Y.P. Predator-prey interactions of nematode-trapping fungi and nematodes: Both sides of the coin. Appl. Microbiol. Biotechnol. 2018, 102, 3939–3949.
- Zhen, Z.; Xing, X.; Xie, M.; Yang, L.; Yang, X.; Zheng, Y.; Chen, Y.; Ma, N.; Li, Q.; Zhang, K.Q.; et al. MAP kinase Slt2 orthologs play similar roles in conidiation, trap formation, and pathogenicity in two nematode-trapping fungi. Fungal Genet. Biol. 2018, 116, 42–50.
- Chen, R.E.; Thorner, J. Function and regulation in MAPK signaling pathways: Lessons learned from the yeast Saccharomyces cerevisiae. Biochim. Biophys. Acta Mol. Cell Res. 2007, 1773, 1311–1340.
- Jiang, C.; Zhang, X.; Liu, H.; Xu, J.R. Mitogen-activated protein kinase signaling in plant pathogenic fungi. PLoS Pathog. 2018, 14, e1006875.
- Xie, M.H.; Yang, J.L.; Jiang, K.X.; Bai, N.; Zhu, M.C.; Zhu, Y.M.; Zhang, K.Q.; Yang, J.K. AoBck1 and AoMkk1 are necessary to maintain cell wall integrity, vegetative growth, conidiation, stress resistance, and pathogenicity in the nematode-trapping fungus Arthrobotrys oligospora. Front. Microbiol. 2021, 12, 649582.
- Kuo, C.Y.; Chen, S.A.; Hsueh, Y.P. The high osmolarity glycerol (HOG) pathway functions in osmosensing, trap morpho-genesis and conidiation of the nematode-trapping fungus Arthrobotrys oligospora. J. Fungi 2020, 6, 191.
- Jiang, K.X.; Liu, Q.Q.; Bai, N.; Zhu, M.C.; Zhang, K.Q.; Yang, J.K. AoSsk1, a response regulator required for mycelial growth and development, stress responses, trap formation, and the secondary metabolism in Arthrobotrys oligospora. J. Fungi 2022, 8, 260.
- Fan, Y.; Zhang, W.; Chen, Y.; Xiang, M.; Liu, X. DdaSTE12 is involved in trap formation, ring inflation, conidiation, and vegetative growth in the nematode-trapping fungus Drechslerella dactyloides. Appl. Microbiol. Biotechnol. 2021, 105, 7379–7393.
- Xie, M.; Bai, N.; Yang, J.; Jiang, K.; Zhou, D.; Zhao, Y.; Li, D.; Niu, X.; Zhang, K.Q.; Yang, J. Protein kinase Ime2 is required for mycelial growth, conidiation, osmoregulation, and pathogenicity in nematode-trapping fungus Arthrobotrys oligospora. Front. Microbiol. 2020, 10, 3065.
- Xie, M.; Wang, Y.; Tang, L.; Yang, L.; Zhou, D.; Li, Q.; Niu, X.; Zhang, K.Q.; Yang, J. AoStuA, an APSES transcription factor, regulates the conidiation, trap formation, stress resistance and pathogenicity of the nematode-trapping fungus Arthrobotrys oligospora. Environ. Microbiol. 2019, 21, 4648–4661.
- Zhen, Z.; Zhang, G.; Yang, L.; Ma, N.; Li, Q.; Ma, Y.; Niu, X.; Zhang, K.Q.; Yang, J. Characterization and functional analysis of calcium/calmodulin-dependent protein kinases (CaMKs) in the nematode-trapping fungus Arthrobotrys oligospora. Appl. Microbiol. Biotechnol. 2019, 103, 819–832.
- Khalil, H.B.; Wang, Z.; Wright, J.A.; Ralevski, A.; Donayo, A.O.; Gulick, P.J. Heterotrimeric Gα subunit from wheat (Triticum aestivum), GA3, interacts with the calcium-binding protein, Clo3, and the phosphoinositide-specific phospholipase C, PI-PLC1. Plant Mol. Biol. 2011, 77, 145–158.
- Meijer, H.J.G.; Munnik, T. Phospholipid-based signaling in plants. Annu. Rev. Plant Biol. 2003, 54, 265–306.
- Xie, M.; Ma, N.; Bai, N.; Zhu, M.; Zhang, K.Q.; Yang, J. Phospholipase C (AoPLC2) regulates mycelial development, trap morphogenesis, and pathogenicity of the nematode-trapping fungus Arthrobotrys oligospora. J. Appl. Microbiol. 2022, 132, 2144–2156.
- Zhang, W.; Hu, C.; Hussain, M.; Chen, J.; Xiang, M.; Liu, X. Role of low-affinity calcium system member Fig1 homologous proteins in conidiation and trap-formation of nematode-trapping fungus Arthrobotrys oligospora. Sci. Rep. 2019, 9, 4440.
- Dawkins, R.; Krebs, J.R. Arms races between and within species. Proc. R. Soc. B 1979, 205, 489–511.
- Zhu, M.; Chen, Y.; Zhao, N.; Bai, H.; Zhang, K.; Huang, X. Multiple olfactory pathways contribute to the lure process of Caenorhabditis elegans by pathogenic bacteria. Sci. China Life Sci. 2021, 64, 1346–1354.
- Yu, X.; Hu, X.; Pop, M.; Wernet, N.; Kirschhofer, F.; Brenner-Weiss, G.; Keller, J.; Bunzel, M.; Fischer, R. Fatal attraction of Caenorhabditis elegans to predatory fungi through 6-methyl-salicylic acid. Nat. Commun. 2021, 12, 5462.
- Stadler, M.; Sterner, O.; Anke, H. New biologically active compounds from the nematode-trapping fungus Arthrobotrys oli-gospora Fresen. Z. Naturforsch. C 1993, 48, 843–850.
- Anderson, M.G.; Jarman, T.B.; Rickards, R.W. Structures and absolute configurations of antibiotics of the oligosporon group from the nematode-trapping fungus Arthrobotrys oligospora. J. Antibiot. 1995, 48, 391–398.
- Wei, L.X.; Zhang, H.X.; Tan, J.L.; Chu, Y.S.; Li, N.; Xue, H.X.; Wang, Y.L.; Niu, X.M.; Zhang, Y.; Zhang, K.Q. Arthrobotrisins A-C, oligosporons from the nematode-trapping fungus Arthrobotrys oligospora. J. Nat. Prod. 2011, 74, 1526–1530.
- Zhang, H.X.; Tan, J.L.; Wei, L.X.; Wang, Y.L.; Zhang, C.P.; Wu, D.K.; Zhu, C.Y.; Zhang, Y.; Zhang, K.Q.; Niu, X.M. Morphology regulatory metabolites from Arthrobotrys oligospora. J. Nat. Prod. 2012, 75, 1419–1423.
- He, Z.Q.; Tan, J.L.; Li, N.; Zhang, H.X.; Chen, Y.H.; Wang, L.J.; Zhang, K.Q.; Niu, X.M. Sesquiterpenyl epoxy-cyclohexenoids and their signaling functions in nematode-trapping fungus Arthrobotrys oligospora. J. Agric. Food Chem. 2019, 67, 13061–13072.
- Kuo, T.H.; Yang, C.T.; Chang, H.Y.; Hsueh, Y.P.; Hsu, C.C. Nematode-trapping fungi produce diverse metabolites during predator-prey interaction. Metabolites 2020, 10, 117.
- Xu, L.L.; Lai, Y.L.; Wang, L.; Liu, X.Z. Effects of abscisic acid ad nitric oxide on trap formation and trapping of nematodes by the fungus Drechslerella stenobrocha AS6.1. Fungal Biol. 2011, 115, 97–101.
- Song, T.Y.; Xu, Z.F.; Chen, Y.H.; Ding, Q.Y.; Sun, Y.R.; Miao, Y.; Zhang, K.Q.; Niu, X.M. Potent nematicidal activity and new hybrid metabolite production by disruption of a cytochrome P450 gene involved in the biosynthesis of morphological reg-ulatory arthrosporols in nematode-trapping fungus Arthrobotrys oligospora. J. Agric. Food Chem. 2017, 65, 4111–4120.
- Xu, Z.F.; Wang, B.L.; Sun, H.K.; Yan, N.; Zeng, Z.J.; Zhang, K.Q.; Niu, X.M. High trap formation and low metabolite production by disruption of the polyketide synthase gene involved in the biosynthesis of arthrosporols from nematode-trapping fungus Arthrobotrys oligospora. J. Agric. Food Chem. 2015, 63, 9076–9082.
- Teng, L.L.; Song, T.Y.; Chen, Y.H.; Chen, Y.G.; Zhang, K.Q.; Li, S.H.; Niu, X.M. Novel polyketide-terpenoid hybrid metabolites from a potent nematicidal Arthrobotrys oligospora mutant delta AOL_s00215g278. J. Agric. Food Chem. 2020, 68, 11449–11458.
- Wang, X.; Li, G.H.; Zou, C.G.; Ji, X.L.; Liu, T.; Zhao, P.J.; Liang, L.M.; Xu, J.P.; An, Z.Q.; Zheng, X.; et al. Bacteria can mobilize nematode-trapping fungi to kill nematodes. Nat. Commun. 2014, 5, 5776.
- Su, H.N.; Xu, Y.Y.; Wang, X.; Zhang, K.Q.; Li, G.H. Induction of trap formation in nematode-trapping fungi by bacte-ria-released ammonia. Lett. Appl. Microbiol. 2016, 62, 349–353.
- Zhang, W.; Chen, J.; Fan, Y.; Hussain, M.; Liu, X.; Xiang, M. The E3-ligase AoUBR1 in N-end rule pathway is involved in the vegetative growth, secretome, and trap formation in Arthrobotrys oligospora. Fungal Biol. 2021, 125, 532–540.
- Peng, H.; Dong, X.; Lu, H.; Kong, X.; Zha, X.; Wang, Y. A putative F-box-domain-encoding gene AOL_s00076g207 regulates the development and pathogenicity of Arthrobotrys oligospora. J. Basic Microbiol. 2022, 62, 74–81.
- Zhang, G.; Zheng, Y.; Ma, Y.; Yang, L.; Xie, M.; Zhou, D.; Niu, X.; Zhang, K.Q.; Yang, J. The velvet proteins VosA and VelB play different roles in conidiation, trap formation, and pathogenicity in the nematode-trapping fungus Arthrobotrys oligospora. Front. Microbiol. 2019, 10, 1917.
- Zhou, L.; Li, M.; Cui, P.; Tian, M.; Xu, Y.; Zheng, X.; Zhang, K.; Li, G.; Wang, X. Arrestin-coding genes regulate endocytosis, sporulation, pathogenicity, and stress resistance in Arthrobotrys oligospora. Front. Cell Infect. Microbiol. 2022, 12, 754333.
- Si, J.; Dong, X.; Zhang, G.; Lu, H.; Tang, K.; Zhang, L.; Kong, X.; Sheng, K.; Wang, J.; Zha, X.; et al. The fucose-specific lectin gene AOL_s00054g276 affects trap formation and nematocidal activity of the nematophagous fungus Arthrobotrys oligospora. FEMS Microbiol. Lett. 2022, 369, fnac013.
- Zhang, D.; Zhu, X.; Sun, F.; Zhang, K.; Niu, S.; Huang, X. The roles of actin cytoskeleton and actin-associated protein Crn1p in trap formation of Arthrobotrys oligospora. Res. Microbiol. 2017, 168, 655–663.
- Wernet, V.; Waeckerle, J.; Fischer, R. The STRIPAK component SipC is involved in morphology and cell-fate determination in the nematode-trapping fungus Duddingtonia flagrans. Genetics 2022, 220, iyab153.
- Liang, L.; Shen, R.; Mo, Y.; Yang, J.; Ji, X.; Zhang, K.Q. A proposed adhesin AoMad1 helps nematode-trapping fungus Ar-throbotrys oligospora recognizing host signals for life-style switching. Fungal Genet. Biol. 2015, 81, 172–181.
- Li, X.; Kang, Y.Q.; Luo, Y.L.; Zhang, K.Q.; Zou, C.G.; Liang, L.M. The NADPH oxidase AoNoxA in Arthrobotrys oligospora functions as an initial factor in the infection of Caenorhabditis elegans. J. Microbiol. 2017, 55, 885–891.
- Wu, Q.Y.; Zhu, Y.Y.; Zou, C.G.; Kang, Y.Q.; Liang, L.M. GPH1 is involved in glycerol accumulation in the three-dimensional networks of the nematode-trapping fungus Arthrobotrys oligospora. J. Microbiol. 2016, 54, 768–773.
- Ji, X.; Li, H.; Zhang, W.; Wang, J.; Liang, L.; Zou, C.; Yu, Z.; Liu, S.; Zhang, K.Q. The lifestyle transition of Arthrobotrys oligospora is mediated by microRNA-like RNAs. Sci. China Life Sci. 2020, 63, 543–551.
- Han, X.; Altegoer, F.; Steinchen, W.; Binnebesel, L.; Schuhmacher, J.; Glatter, T.; Giammarinaro, P.I.; Djamei, A.; Rensing, S.A.; Reissmann, S.; et al. A kiwellin disarms the metabolic activity of a secreted fungal virulence factor. Nature 2019, 565, 650–650.
- Lo Presti, L.; Kahmann, R. How filamentous plant pathogen effectors are translocated to host cells. Curr. Opin. Plant Biol. 2017, 38, 19–24.
- Takai, Y.; Kaibuchi, K.; Kikuchi, A.; Kawata, M. Small GTP-Binding Proteins. In International Review of Cytology; Jeon, K.W., Friedlander, M., Eds.; Academic Press: Cambridge, MA, USA, 1992; Volume 133, pp. 187–230.
- Andersson, K.M.; Kumar, D.; Bentzer, J.; Friman, E.; Ahren, D.; Tunlid, A. Interspecific and host-related gene expression patterns in nematode-trapping fungi. BMC Genom. 2014, 15, 1–15.
- Liu, Q.; Li, D.; Jiang, K.; Zhang, K.Q.; Yang, J. AoPEX1 and AoPEX6 are required for mycelial growth, conidiation, stress response, fatty acid utilization, and trap formation in Arthrobotrys oligospora. Microbiol. Spectr. 2022, in press.
- Liu, Q.; Li, D.; Jiang, K.; Zhang, K.Q.; Yang, J. AoPEX1 and AoPEX6 are required for mycelial growth, conidiation, stress response, fatty acid utilization, and trap formation in Arthrobotrys oligospora. Microbiol. Spectr. 2022, in press.