The beneficial effects of nanotechnology in the field of disease diagnosis and therapy depends on the evolution of innovative approaches for cell tracking in living subjects. Recent developments in the use of nanotechnologies have contributed to advance of the high-resolution in vivo imaging methods as well as targeted disease approaches. In this context, the application of strategies for the biomimetic design and functionalization of nanoparticles (NP) to be used for directing cell labelling and their intracellular retention have received growing interest and require further investigation to improve direct cell tracking protocols allowing non-invasive long-term targeted monitoring through in vivo imaging.
- nanoparticles
- cell tracking
- cell labelling
- cell uptake
- stem cell homing
1. NP typology for in cell imaging
A literature survey covering research studies published over the last 10 years allowed to give an overview on (I) key nanocarrier-impacting elements, such as NP typology and functionalization, by analyzing their ability in tuning cell interaction mechanisms under in vitro and in vivo conditions (II) as well as sensitivity and specificity of the imaging devices supporting NP delivery assessment. Importantly, the identified nanocarrier elements were found to be beneficial for supporting both regenerative and cancer imaging. Below a description of the most relevant elements to be considered for the development of accurate cell tracking platforms.
1. NP typology for in cell imaging
Four distinctive classes of NP were identified to support cell tracking platforms: inorganic, polymeric, hybrid and lipid based. The inorganic category resulted the most represented NP typology in this survey, accounting for the 79.6% of the total collected citations, followed by polymeric (12.5%), hybrid (4.6%) and lipid based (3.3%), respectively (Figure 1). The prominent representation of the inorganic typology reflects the fact that these NP are the ones best suited to cell tracking through the most characterized imaging platforms to date.
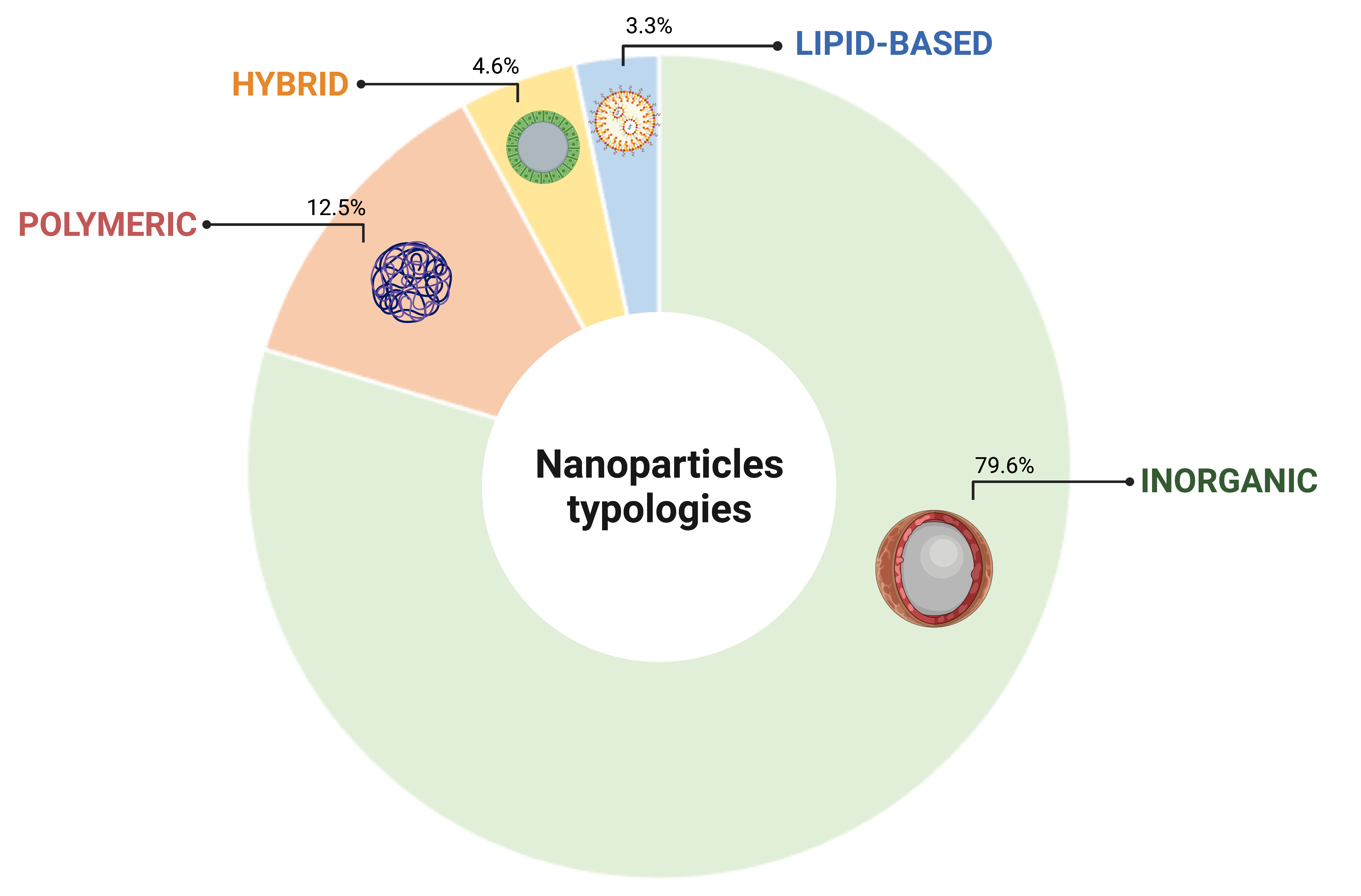
Figure 1.
Classification of NPs based on the constitutive element. Created with BioRender.
2. NP functionalization strategies directing targeted cell uptake, imaging performance and therapeutic tracking
Firstly, accurate cell tracking occurs if the NP are correctly taken up by the cell to be targeted. The targetability and efficiency of the uptake are governed by the interactions between the NP surface groups and the plasma membrane antigens and/or receptors, which in turn depend on the density of the ligands and the antigens/receptors present on an NP and a cell, respectively [1]. NP functionalization with specific bioactive molecules have been investigated to that purpose and those collected with the bibliographic survey were classified on the basis of their ability to capture specific cell biomarkers such as antigens/receptors (moieties with active action such as antibodies, peptides, aptamers, carbohydrates, glycoproteins and vitamins) or to enhance NP permeation and retention (moieties with passive action such as amino acids, polymers, polypeptides, heparin and quaternary ammonium cations) (Figure 2; targeting and uptake moieties).
Secondly, NP functionalization with contrast agents to be followed with imaging devices is strictly required for the identification of tissue alterations or lesion characterization as well as the diagnosis and treatment of any disease. Different molecules with optical properties such as fluorophore and bioluminescent dyes, isotopes or chemical elements with high molecular weight have been investigated as contrast agents and applied for imaging of cells bearing NP [2] (Figure 2; labeling moieties).
Lastly, strategies to track therapy delivery through imaging assume a central role for the implementation of currently available diagnostic and therapeutic protocols [3]. In this context, several therapeutic NP that are capable of both self-reporting disease and/or tissue damage and delivering therapy have been developed to date. Functionalization allowing the use of these NPs as therapeutic agents, referred to as therapeutic moieties consist of molecules managing the proliferative and inflammation status of the target cell. Therapeutic moieties identified by the present literature survey include miRNA, siRNA, genes and drugs or compounds with key roles in the modulation of cell proliferation and differentiation (Figure 2; uptake moieties).
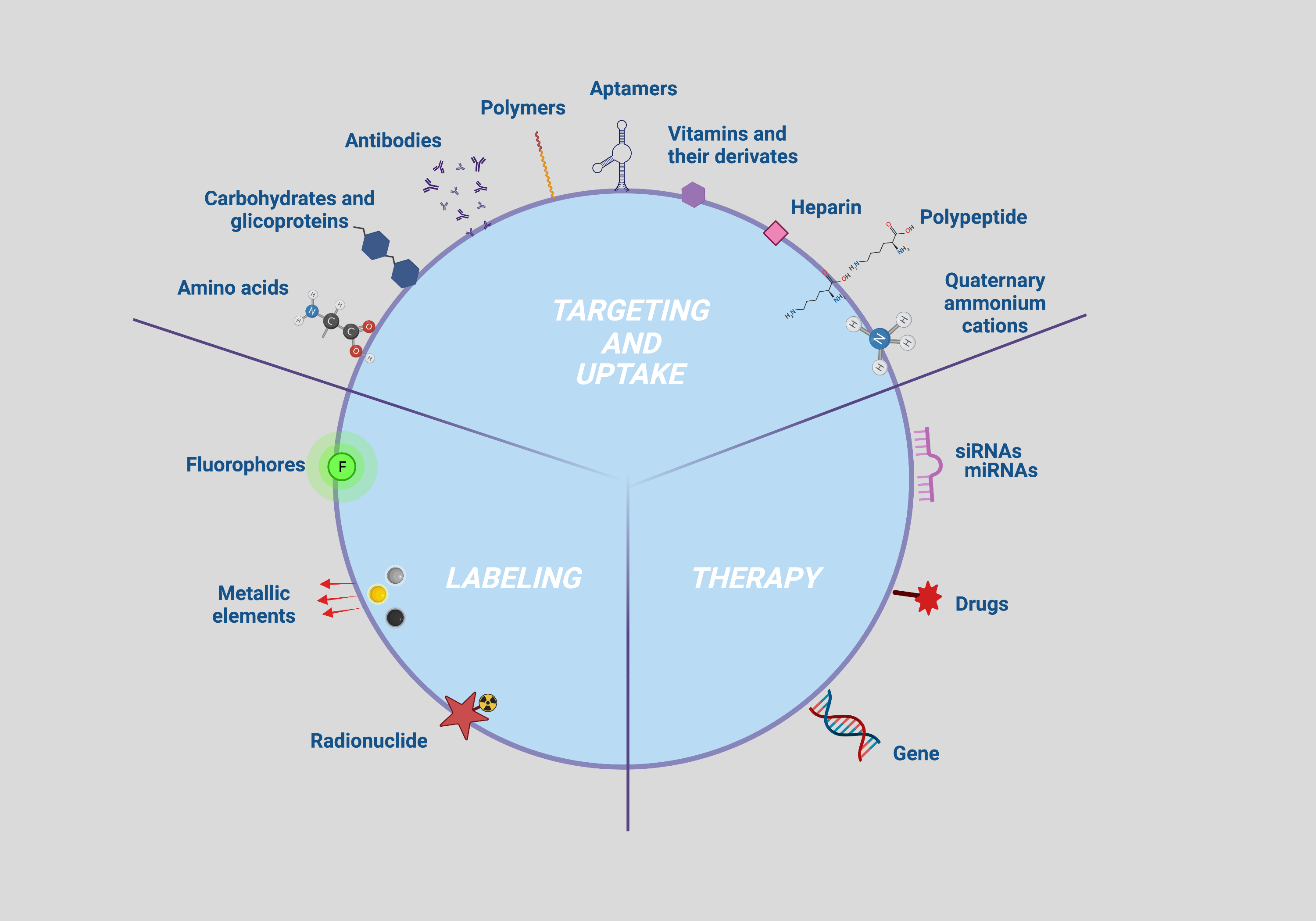
Figure 2.
Multimodal NP surface modifications applied for enhancing targeting, uptake, tracking, and therapy. Created with BioRender.
3. Approaches and devices for in vitro and in vivo tracking of NP delivery
In vitro and in vivo protocols for cell imaging rely on physical chemical properties of NP core element and NP functionalization moieties.In vitro tracking of the NP core element can be achieved by means of transmission electron microscopy (TEM) and scanning electron microscopy (SEM), whereas NP surface-labeling moieties such as fluorophores and cell-labeling moieties such as luciferin can be followed in vitro by using confocal laser microscope and luminometer, respectively (Figure 3A). On the other hand, in vivo tracking of the core element can be accomplished with MRI and tomography imaging such as computed tomography (CT), microcomputed tomography (µCT), photoacoustic tomography (PA) and X-ray fluorescence tomography (XFT) imaging. NP functionalization with fluorophores have been also applied for in vivo fluorescence tracking with imaging devices such as the NIR fluorescence IVIS Spectrum system, fluorescence molecular tomography imaging and single- and two-photon fluorescence. Of note, according to NP classification, the present literature survey identified the MRI (52%), followed by CT (25%), optical (19%) and radionuclide (4%) imaging as the most used in vivo imaging devices (Figure 3B).
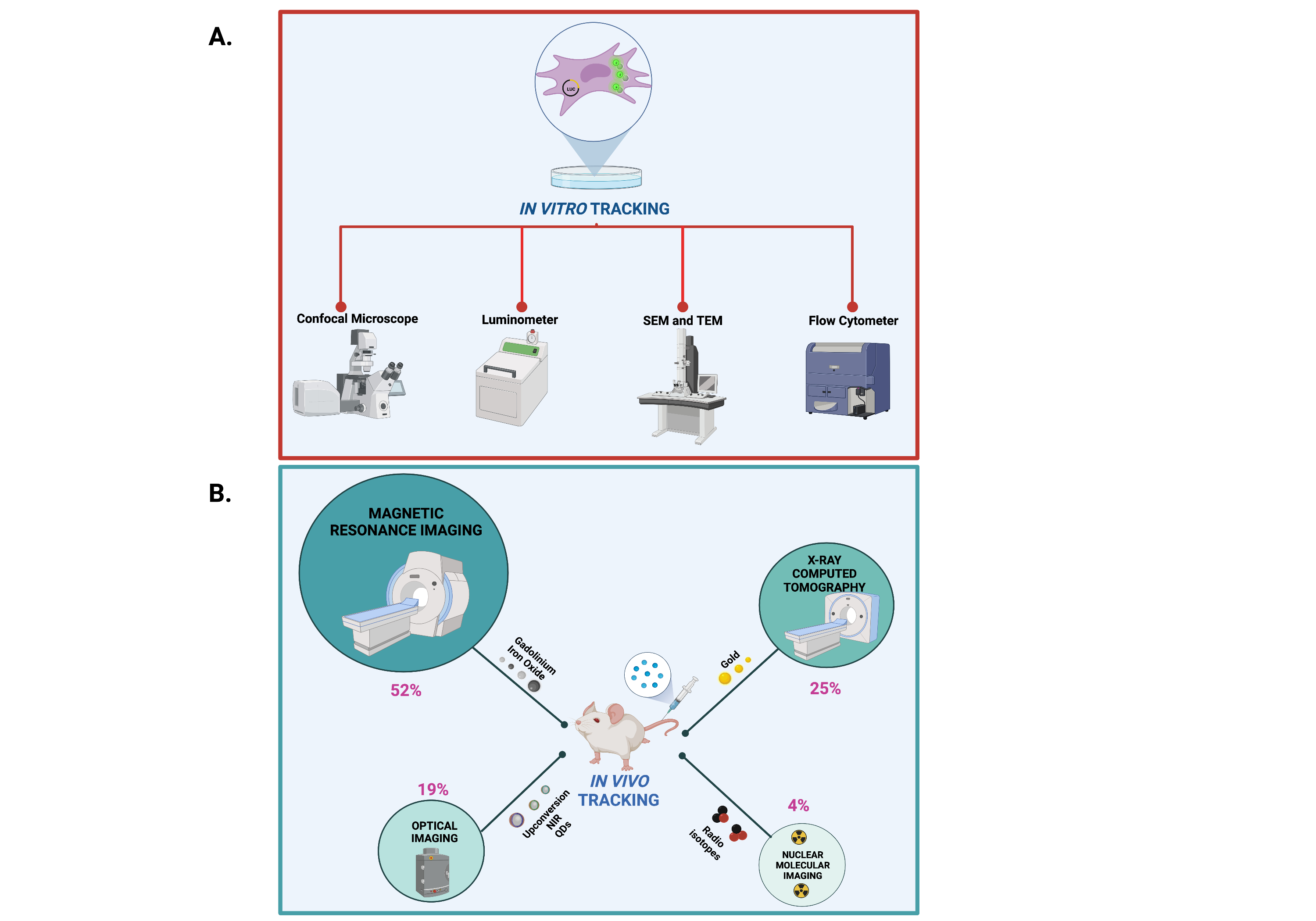
Figure 3.
Methods and devices for in vivo and in vitro imaging of NPs. Created with BioRender.
4. NP uptake strategies and mechanisms
The effects of the physicochemical properties of NP, such as size, shape, charge and surface chemistry, are crucial for determining the cellular uptake mechanism [4] consequently influencing NP fate upon delivery [1] and the success of any NP-based cell tracking approach. OuResearchers' bibliographic survey identified the endocytosis mechanism as the most representative for the internalization of NP, whereases a limited number of studies find electroporation and transfection as alternative strategies for an effective NP delivery into the cell.
5. Cell Types Suitable for Safe NP Delivery
Several cell models have been investigated in the last decade as potential carriers for an efficient NP release.
Stem cells, with their intrinsic ability to home in on cancer or injury sites, represent the most suitable cell type source allowing NP delivery, with a 60% of the reviewed studies investigating their potential for biomedical application for cancer and regenerative medicine purposes. Conversely, a smaller percentage of studies (31%), focusing on the development of clinical protocols for precision cancer medicine, adopted cancer cells as the target of NP delivery. Only a few original research articles, pointing to the elucidation of inflammatory homing mechanisms, used other cell sources such as macrophages (7%) or T cells (2%).
Take-home message
Issues determining the effectiveness of NP-assisted cell tracking such as NP design, material properties and functionalization with specific targeting, labeling (tracking) and/or uptake-enhancing moieties have been identified and almost exhaustively characterized during the last decade. Nevertheless, an in-depth analysis of mechanisms determining the efficacy of NP incorporation by the cell holds a prominent role in NP-assisted cell tracking as it keeps the potential clinical traceability for a targeted, sensitive, and long monitoring of cells with the current clinically approved imaging devices improving the performance of the currently used iodinated, barium or gadolinium-based contrast agents [5].
Based on these premises, this conceptual synthesis of the literature data aims to create an easily employable knowledge platform to be exploited for the optimization of NP delivery solutions for cell tracking, finally providing future perspectives for a bench to bedside translation (Figure 4).
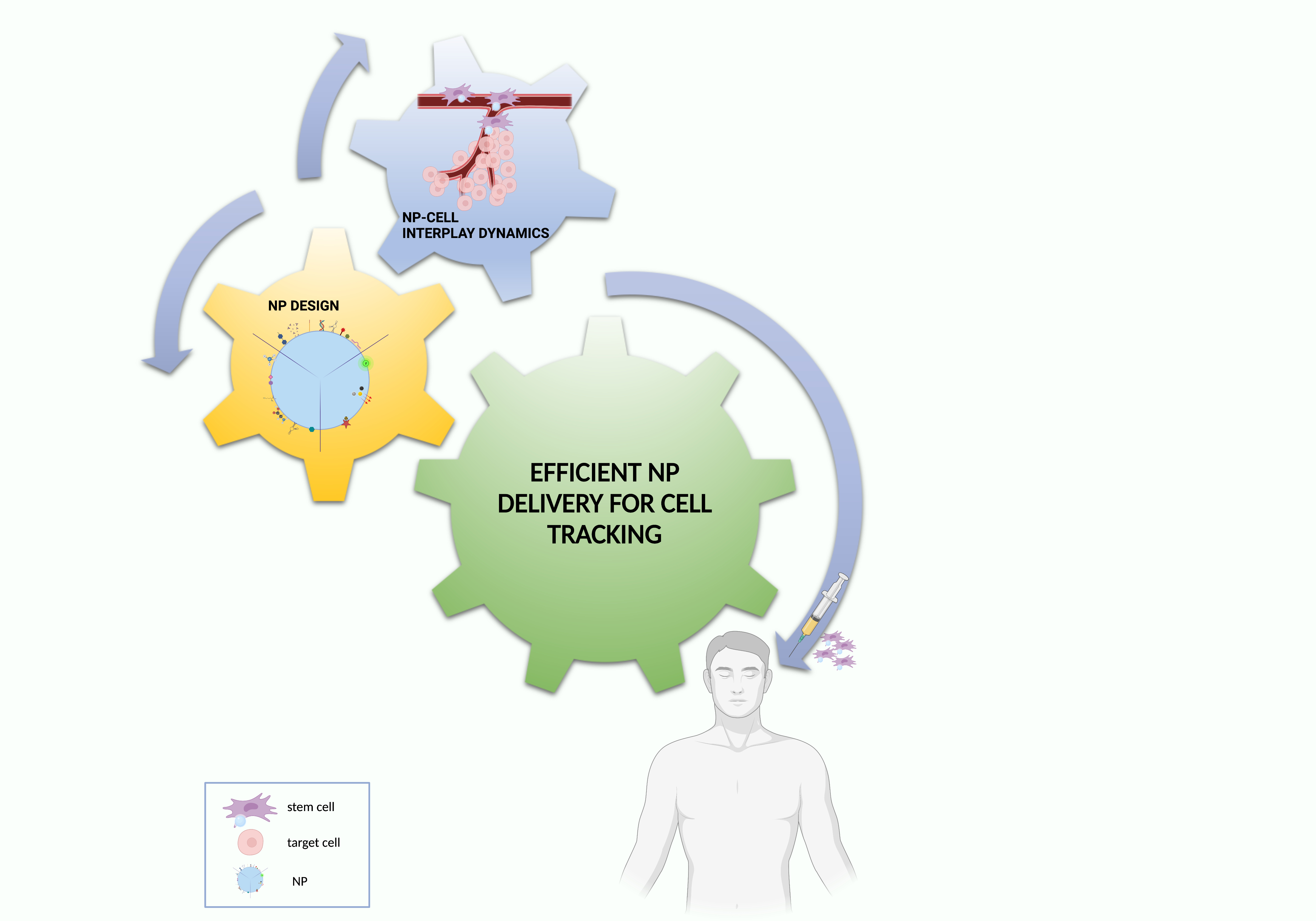
Figure 4. Essential ingredients for effective NP–cell tracking. Created with BioRender.
References
- Parisa Foroozandeh; Azlan Abdul Aziz; Insight into Cellular Uptake and Intracellular Trafficking of Nanoparticles. Nanoscale Research Letters 2018, 13, 1-12, 10.1186/s11671-018-2728-6.
- Benjamin P. Burke; Christopher Cawthorne; Stephen J. Archibald; Multimodal nanoparticle imaging agents: design and applications. Philosophical Transactions of the Royal Society of London. Series A: Mathematical, Physical and Engineering Sciences 2017, 375, 20170261, 10.1098/rsta.2017.0261.
- Cristina Zavaleta; Dean Ho; Eun Ji Chung; Theranostic Nanoparticles for Tracking and Monitoring Disease State. SLAS Technology 2017, 23, 281-293, 10.1177/2472630317738699.
- Heather Herd; Nicole Daum; Arwyn T. Jones; Hanno Huwer; Hamidreza Ghandehari; Claus-Michael Lehr; Nanoparticle Geometry and Surface Orientation Influence Mode of Cellular Uptake. ACS Nano 2013, 7, 1961-1973, 10.1021/nn304439f.
- M.R. (Monique) Bernsen; J. (Jamal) Guenoun; S.T. (Sandra) Van Tiel; G.P. (Gabriel) Krestin; Nanoparticles and clinically applicable cell tracking. The British Journal of Radiology 2015, 88, 20150375, 10.1259/bjr.20150375.
