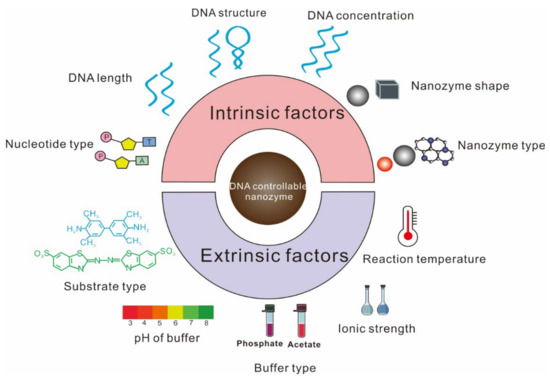
With helps from nanotechnology advancement, researchers have found that ssDNA could enhance the catalytic activity of nanomaterials, including noble-metal-based nanomaterials
[31][33], transition-metal-based nanomaterials, carbon-based nanomaterials
[48][50], MOFs, and their hybrids. As mentioned above, the adsorption of ssDNA onto the surface of nanomaterials was a prerequisite for ssDNA to regulate the nanozyme activity. For example, ssDNA on the surface of nanozymes could adsorb more substrate (e.g., 3,3′,5,5′-tetramethylbenzidine (TMB)) around the nanozyme due to electrostatic attraction and π–π stacking interaction between TMB and ssDNA, further accelerating electron transfer from TMB to the target sensing molecules (e.g., H
2O
2) and enhancing the nanozyme activity
[43][49][45,51]. Thus, factors affecting both ssDNA–nanomaterial and ssDNA–TMB affinity are important for the enhancement of nanozyme activity by ssDNA. The usual trend of nanozyme activity was to first increase and then decrease with the increase in ssDNA concentration
[50][51][52][52,53,54]. However, some data support a different variation trend of nanozyme activity, which is to first increase and then keep stable with the increase of ssDNA concentration
[44][48][49][53][54][46,50,51,55,56]. The reason for the two different trends was unclear.
ThWe
researchers sspeculate that the presence of excessive ssDNA may hinder the contact of the nanozyme and TMB in the former trend since it had a wide range of ssDNA concentration. DNA molecules with different structures were also reported to affect the peroxidase-like activity of nanozymes. The order of enhancement of the ability of catalytic activity was: hybridization chain reaction (HCR) products (long dsDNA) > hairpin DNA > ssDNA > dsDNA
[51][53]. Moreover, different DNA nucleotides had different abilities to enhance the nanozyme activity. The order of enhancement of Fe
3O
4 nanozyme activity was: C > G > T > A
[43][45]. Cytosine in buffer at pH 4.0 was protonated, which could assist charge neutralization on the surface of Fe
3O
4 nanoparticles and reduce repulsion among DNA, leading to the adsorption of more DNA. The same variation trend was observed by Wang et al.
[48][50] using protonated graphitic carbon nitride (Pg-C
3N
4) nanosheets as nanozymes. The peroxidase-like activity of 20 bp homo ssDNA (A
20, C
20, G
20, and T
20) modified MoS
2 nanozymes followed the trend of G
20 ≈ T
20 > A
20 > C
20 > no DNA. The explanation from another perspective was given for the weakest activity of C
20-modified MoS
2 nanozymes. Protonated cytosine nucleotides in buffer at pH 4.0 increased the electrostatic repulsion between cytosine and positively charged TMB, resulting in a lower affinity of MoS
2 nanozymes
[49][51]. Guanine (G) was also reported to significantly enhance the peroxidase-like activity of WS
2 nanosheets
[55][57], iron-based MOFs modified with acidized carbon nanotubes (MOF/CNTs)
[50][52], and MIL-53(Fe)
[44][46]. Purine (A, G) modification demonstrated a remarkable enhancement of the peroxidase-like activity of AuNPs, while pyrimidine (T, C) modification enhanced it slightly, which was attributed to the difference in the interaction between TMB and the surface-adsorbed nucleobases
[56][58]. In addition, the effect of DNA length on the enhancement of nanozyme activity was also investigated. The longest DNA (ploy C
30) demonstrated the largest enhancement of peroxidase-like activity of Fe
3O
4 nanoparticles due to the presence of more binding sites
[43][45]. Increasing the number of cytosine nucleotides from 30 to 40, the peroxidase-like activity of Pg-C
3N
4 nanosheets kept steady, which indicated the saturation of ssDNA on the surface of the Pg-C
3N
4 nanosheets
[48][50]. A slight inhibition of the catalytic activity of MoS
2 nanozymes was found when the number of thymidine nucleotides increased from 30 to 40
[49][51].
Besides the intrinsic factors mentioned above, the reaction conditions were also very important for DNA to regulate the nanozyme activity. The buffer pH at 4.0 had been found to be the optimum condition for both the catalytic activity of nanozymes and the inhibition of nanozyme activity by ssDNA
[43][44][54][55][45,46,56,57]. TMB was the most-used substrate in those literatures. It carries positive charges at pH 4.0 that is below its pKa, which is beneficial to the reaction between ssDNA and TMB
[43][45]. However, continuously reducing the pH resulted in the protonation of two amino acids of TMB, which made them insusceptible to oxidation
[57][59]. Moreover, buffer type and concentration were also reported to affect the ability of ssDNA to enhance the peroxidase-like activity of nanozymes. In a phosphate buffer, low-concentration DNA moderately increased the peroxidase-like activity of CeO
2, while DNA in acetate buffer had no effect on the catalytic activity of CeO
2, except for the high-concentration DNA (more than 10 µM), which inhibited the CeO
2 nanozyme activity
[58][60]. This was because phosphate could compete with ssDNA to bind to CeO
2 nanozymes and enhance their peroxidase-like activity. Low ionic strength increased the peroxidase-like activity of ssDNA-modified WS
2 nanosheets
[55][57], while high ionic strength could shield the electrostatic reaction, which reduced the interaction among ssDNA, nanozymes, and TMB
[59][61].
More amazingly, the enhancement or inhibition of the peroxidase-like activity of AuNPs by ssDNA can be controlled by H
2O
2 concentration. In a 10 mM H
2O
2 reaction system, DNA inhibited the peroxidase-like activity of AuNPs in the first 10 min and enhanced the catalytic activity of AuNPs in the next 20 min of the reaction time, while at a lower concentration of H
2O
2 (5 mM), it prolonged the inhibition time to 30 min and enhancement occurred after 30 min
[56][58]. Moreover, by changing the substrate from the positively charged TMB to the negatively charged 2,2′-azino-bis (3-ethyl benzothiazoline-6-sulfonic acid) (ABTS), the enhanced ability of ssDNA disappeared, and the inhibiting ability of ssDNA appeared
[43][55][45,57].
As previously mentioned, ssDNA can either enhance or inhibit the catalytic activity of nanozymes, and the deciding factor for enhancement or inhibition was to increase or decrease the affinity of substrate (e.g., TMB or ABTS) and nanozymes, respectively, by the introduction of ssDNA. A small change of these factors caused the interaction of ssDNA and nanozymes to be altered, which resulted in variations of the nanozyme activity. Thus, the process of designing aptamer-assisted nanozyme sensing needs to be thoughtful since various factors can affect the regulation of ssDNA on nanozyme activities.