Your browser does not fully support modern features. Please upgrade for a smoother experience.
Please note this is a comparison between Version 2 by Camila Xu and Version 1 by Rosaria Saletti.
Le proteine VDAC (canale selettivo anionico voltaggio-dipendente), note anche come suine mitocondriali, sono le proteine più abbondanti della membrana mitocondriale esterna (OMM), dove svolgono un ruolo vitale in vari processi cellulari, nella regolazione del metabolismo e nelle vie di sopravvivenza.
- voltage dependent anion channel
- cysteine overoxidation
- deamidation
1. Introduzctione
1.1. Isoforme VDAC: una famiglia di proteine hub
1.1. VDAC Isoforms: A Family of Hub Proteins
Le proteine VDAC (canale selettivovoltage-dependent anionico voltaggio-dipendente), note anche come suirine selective channel) proteins, also known as mitochondriali, sono le proteine più abbondanti della membrana porins, are the most abundant proteins of the outer mitochondriale esterna membrane (OMM) dove svolgono un ruolo vita where they play a vital role in vari processious cellulari, nella regolazione delr processes, in the regulation of metabolismo e nelle vie di sopravvivenza. Mediano lo scambio di ioni e , and in survival pathways. They mediate the ions and metaboliti tra ies exchange between mitocondri e il resto della cellula, garantendo una buona funzhondria and the rest of the cell, ensuring good functionalità dei complessiy of mitochondriali e la produzione complexes and energy production [1].
Negli eucarioti superiori, ci sono trekaryotes, there are three VDAC isoforme VDAC s (VDAC1, VDAC2, VDAC3) codificate da geniencoded by separati situati su ce genes located on different chromosomi diversies [2]. QuTheste proteine che formano i pori hanno un pesoe pore-forming proteins have a similar molecolare simileular weight (30 kDa) e sequenze altamenteand highly conservate di circaed sequences of about 280 amminoacidi ad eccezione delmino acids with the exception of VDAC2 che ha la frazionwhich has the N-terminale di moiety of 11 residui più lunga delle altrees longer than the other isoformes.
L'analisiThe evolutiva indicaonary analysis indicates VDAC3 come l' as the oldest isoforma più antica, mentr, while VDAC1 èis considerata la più giovane porina mitoced the youngest mitochondriale porin [3][4][3,4].
LThe strutture sexperimentali 3D dell'isoforma 3D structures of mouse and human VDAC1 di topo e umana sono state isoform have been determinate utilizzando la cried using X-ray crystallografia a raggi X e la phy and NMR [5][6][7]5,6,7]. QuTheste analisi hanno rivelato unyses revealed a struttura costituita dacture constituted by 19 β-filamenti disposti a formare unstrands arranged to form a trans-membrana β-barile e da une β-barrel and by a regione contenente α-elica al containing α-helix at the N-terminus della of the proteina. La canna è. The barrel is organizzata come una normale schieraed as a regular antiparallela di β fili ad eccezione dei fili 1 e 19 che corrono array of β-strands with the exception of strands 1 and 19 that run in parallelo. La coda anfipatica a α elica si trova all'interno del poro. Tuttavia, la posizione esatta e la struttura locale di questo. The amphipathic α-helix tail is located inside the pore. However, the exact position and local structure of this segmento sono ancora sfuggenti poiché queste caratteristiche non si sovrappongono perfettamente nelle strutture a raggi X e NMR disponibili are still elusive since these features are not perfectly overlapping in the available X-ray and NMR structures [5][6][7]5,6,7].
Recentemenly, te, la struttura dihe structure of zebrafish VDAC2 è stata risolta ad alta risoluzionewas solved at a high resolution, confermando la stessa disposizionirming the same β-barrel diarrangement as VDAC1 [8]. Zebrafish VDAC2 has unone cysteine residuo di cisteina nella e in the sequenza e manca della sequenza ce and lacks the 11 amino acid longer N-terminale più lunga di 11 amminoacidi sequence presente nelle VDAC d in mammalian VDACs.
Thei mammiferi.
LaVDAC3 structtura VDAC3 non è stata ancoraure has not yet been determinata. Diverse previsioni ed. Several bioinformatiche, basate sulla somiglianza di grandi predictions, based on the large sequenze, hannoce similarity, proposto un nucleo di botte come le altreed a barrel core such as the other VDAC isoforme VDAC [9]s [9]. NonoDestante l'elevata somiglianza di pite the high sequenza e l'omologia strutturale, lece similarity and structural homology, VDAC isoforme VDAC mostrano diverse proprietà funzionali all'interno della cellulas display different functional properties within the cell.
L'aAnalisi dei livelli di esysis of the expressione delle isoforme levels of human VDAC umane nelle cellule HeLaisoforms in HeLa cells, determinati mediante REAL-TIME ed by real-time PCR, suggerisce chests that VDAC1 è l'isoforma più abbondante, dieci volte più abbondante rispetto ais the most abundant isoform, ten times more abundant compared to VDAC2 e cento volte più abbondante rispetto a and hundred times more abundant compared to VDAC3, la meno caratterizzata dell the least characterized of the isoforme. Inoltre, la sovraess. In addition, the overexpressione di ogni singola of each single VDAC isoforma VDAC influisce sui livelli di mRNA delle altre due affects the mRNA levels of the other two isoformes, suggerendo che i rapporti tra le isoformesting that the ratios between VDAC sono sottoposti a un controlloisoforms are subjected to a reciproco che evita uno squilibrio tra questcal control that avoids an imbalance among these proteines [10].
SebbeneAlthough le trthe three isoforme mostrino un coinvolgimento comune nel mantenimento dellas show a common involvement in cellular bioenergetica cellulars maintenance, VDAC1 eand VDAC2 hanno funzioni ve specializzate nella morte cellulareed functions in programmata. Per l'ed cell death. For VDAC3 isoforma VDAC3, studi , recenti studies indicano un ruolo te a centrale nell role in ROS metabolismo dei ROS e nel controllo di and in mitochondrial qualità mitocondrialey control [11].
LThe funzioni dei VDC sono parecchie e alcune di queste dipendono o sono influenzate dall'ctions of VDACs are several-fold and some of these depend on, or are affected by, interazione con altre proteine ciction with other cytosoliche e and mitochondriali. A causa della loro proteins. Due to their localizzazione presso l'OMM, le VDAC sonozation at the OMM, VDACs are considerate ed to be hub proteine hub, che s, interagiscono con oltrecting with over 200 proteine al fine dis in order to integrare le funzioni mitocte mitochondriali con il resto delle attività cellulari functions with the rest of the cellular activities [12][13][12,13]. PertantoThus, leVDAC isoforme VDAC sembrano essere una giunzione per uns appears to be a junction for a varietà di segnaliy of signals associati a diversi percorsi legati alla sopravvivenza cellulare o alla morteed with different pathways related to cell survival or programmata. Inoltre, la funzione dei VDAC e le loroed death. Furthermore, the function of VDACs and their interazioni con altrections with other proteine sono influenzate da modificazionis are affected by post-traduzionali (PTMnslational modifications (PTMs) [14]. SUnfortunatamente, i PTM delleely, PTMs of VDAC proteine VDAC raps representano un campo poco esplorato, principalmente perché la scoperta e la caratterizzazione del a little explored field, mainly because discovery and characterization of PTM in questthese proteine è molto impegnativa, a causa della loro scarsas is very challenging, due to their poor solubilità ey and impossibilità diy to isolarte singole isoforme. Solo negli ultimi anni, il crescente numero di strumenti volti as. Only in recent years has the increasing number of tools aimed at identificare e ying and quantificare i PTM ha migliorato le conoscenze in questo campo e nei meccanismi che regolano le funzioni e le ying PTMs increased, improving the knowledge in this field and in the mechanisms that regulate functions and interazioni delle suipine mitocctions of mitochondriali porins. In particolare, lo sviluppo di metodi di cromatografiaular, the development of nano-reversed phase ultra-high-performance liquida ad altissime prestazioni chromatography (nanoRP-UHPLC) e di spettrometria di massa ad alta risoluzioneand ultra-sensitive high-resolution mass spectrometry (HRMS) a fase nano-invertita ha svolto un ruolo chiave in questo campo. I risultati ottenuti sui PTM VDAC utilizzando tali metmethods has played a key role in this field. The findings obtained on VDAC PTMs using such methodologie, che hanno permesso una caratterizzazione approfondita di queste proteine dei poris, which have permitted an in-depth characterization of these very hydrophobic trans-membrana molto idrofobiche, sono riassunti in questa recensionee pore proteins, are summarized in this review.
1.2. Le VDAC come attori principali nella mediazione e nella regolazione delle funzioni mitocondriali con attività cellulari
1.2. VDACs as Main Players in Mediating and Regulating Mitochondrial Functions with Cellular Activities
LaThe posizione nell'OMM consente allelocation in the OMM allows the VDAC proteine VDAC di agire come punti di ancoraggio pes to act as anchor points for diversi insiemi die sets of molecole cheules that interagiscono con ict with mitocondri. In questo modo, i VDC sono in grado di mediare e regolare l'hondria. In this way, VDACs are able to mediate and regulate the integrazione delle funzionition of mitochondriali con le attività functions with cellulari activities.
L'intTherattoma VDAC comprendeinteractome includes proteine situates located in OMM, membranainner mitochondriale interna membrane (IMM), spazio iintermembranae space (IMS), ciytosol, reticolo endoendoplasmic reticulum, plasmatico, membrana plasmatica ee and nucleo che sono coinvolti nelus that are involved in metabolismo, apoptosi, trasduzione del segnales, signal transduction, protezione controction against ROS, legame abinding to RNA o DNA e altro ancorar DNA, and more.
La mMobilità della regioney of VDAC N-terminale α-elica è α-helix region is importante per i for channel gating dei canali, ma anche per le but also for interazioni con proteinections with both pro-apoptotiche e and anti-apoptotiche come proteins such as Bax, Bak e, and Bcl-xL [15][16][17]15,16,17]. Il VDAC1 è coinvolto nel rilascio di fattoriis involved in the release of apoptotici localizzati nello spazio i factors located in the intermembrana grazie alla sua capacità die space due to its ability of oligomerizzarsizing in dimeri, esameri e strutture di ordine superiore, per formare un grande poro che consente il s, hexamers, and higher-order structures, to form a large pore that allows the passaggio del citocromo c e del fattore di induzione dell'apoptosie of cytochrome c and apoptosis inducing factor (AIF) al ci to the cytosol e diand conseguenza l'attivazione della morte cellulare programmata. Invecequently the activation of programmed cell death. Instead, VDAC2 funziona come fattorections as anti-apoptotico ed è sovraregolato in diverse malattie factor and it is upregulated in several debilitanti tra cui l'ting diseases including Alzheimer e il cancro’s and cancer [18]. QueThista proprietà è property is probabilmente dovuta alla capacità unica dily due to the unique ability of VDAC2 dito sequestrare la proteinaer the pro-apoptotica Bak nell'OMM e mantenerla nello stato inattivo protein Bak in the OMM and maintain it in the inactive state [11].
VDAC1 modistra siti di legame, situati nella sua porzione ciplays binding sites, located in its cytosolica, per molti enzimi moiety, for many metabolici, come la gli enzymes, such as glyceraldeide-3-fosfato deihyde 3-phosphate dehydrogenasi, lae, creatinchinasi, la glicerolo chinasi, la glucochinasi, la chinasi c-Raf e le kinase, glycerol kinase, glucokinase, c-Raf kinase, and hexokinase isoforme e l'esochinasi (I es (I and II), che necessitano di un accesso which need preferenziale all'ATPtial access to mitochondriale ATP [19].
L'Hesochinasixokinase interagisce attraverso la sua sequenza idrofobicacts through its hydrophobic N-terminale con sequence with Glu73 diof VDAC1, un sito di legamea binding site localizzato su un lato della parete della canna, sepolto nell'ambiente idrofobo died on one side of the barrel wall, buried in the hydrophobic environment of OMM [20].
ÈIt stato dimostrato che il trattamento dei mitocondri con dicicloesihas been demonstrated that treatment of mitochondria with dicyclohexylcarbodiimide (DCCD) inibisce l'hibits hexokinase–VDAC interazione esochinasi-VDAC a causa dellaction due to selective chemical modifica chimica selettiva della gtion of Glu73 [21].
IGlu73 residuoe di glu73is also è ancthe il sito di legame per le binding site for ceramidi, lipidi oncosoes, tumor suppressori in grado di agire direttamente sui mitocondri per innescare la morte delle cellule apoptotiche. È lipids able to act directly on mitochondria to trigger apoptotic cell death. It is interessante notare che sia ting that both VDAC1 che 2 possiedono, in una posizione simile, un and 2 own, in a similar position, a cysteine residuo di cisteinae (Cys127 inel human VDAC1 umano e and Cys138 inel human VDAC2 umano) sotto forma di acido so) in the form of sulfonico con una forte carica acid with a strong negativa simile a quella del residuo acido glutammatoe charge resembling that of the glutamate acid residue [22]. Invstece, l'ad, VDAC3 isoforma VDAC3 non mostra alcun does not show any residuo e homologo aus to Cys127/138 or Glu73 embedded incorporato nella porzione idrofobica dell' the hydrophobic moiety of the OMM.
VDAC1 presenta una tasca legante il c a cholesterolo formata, i binding pocket formed, in human isoforma umana, da residui di, by Ile123,, Leu144,, Tyr146,, Ala151e, and Val171 [ 23].
Mitochondriali formano porins form complessi con altrexes with other proteine, come ils, such as the adenine nucleotide translocase dell'adenina (ANT), la proteina trathe translocatriceor protein (TSPO), nota anche come recettore dellalso known as the peripheral-type benzodiazepine di tipo perifericoreceptor (PBR), HSP70 mitocmitochondriale e diverse HSP70, and several cytoskeletal proteine citoscheletriche comes such as tubulina, actina, catena leggera della dineina e , dynein light chain, and gelsolina [11].
La protTheina tra translocatriceor protein interagisce direttamente con tutte lects directly with all VDAC isoforme VDACs. In particolare, l'ular, interazione traction between TSPO eand VDAC1 contribuisce a regolare l'tes to regulate the efficienza dei meccanismi di controllo della cy of mitochondrial qualità mitocondriale e inibisce la mitofagiay control mechanisms and inhibits mitophagy, prevenendo l'ting ubiquitinazione delletion of proteine attraverso la s through downregulation della via of the PINK1/Parkin [24].pathway Il[24]. motivoThe GxxxG simotif presenta sias both in VDAC cheand in TSPO, ed èand is necessario per questay for this interazionection [25]. InMoltrereover, VDAC1 eand TSPO, in associazione contion with StAR (steroidogenic acute regulatory protein), formano il trasdusoma, un complesso multi the transduceosome, a multi-proteico coinvolto nel trasporto del cn complex involved in cholesterolo. In una precedente ipotesi transport. In a former hypothesis, VDAC1 eand TSPO in OMM, ANT in IMM e, and Cyclophilin D nella matricin the mitochondriale erano matrix were candidati a costituire il poro dies to constitute the permeability transizione di permeabilitàtion pore (PTP), un poro ad altaa high conduttanza e non ctance and non-specifico che consente il gonfiore pore that allows mitochondriale e il rilascio di proteine swelling and release of apoptogeniche. Più proteins. More recentemente, è statoly, it was proposto che il PTP potesse essere formato daed that PTP could be formed by dimeri del complessos of the ATP sintasiynthase complex [26].
Recenti studi hanno focalizzato l'attenzione sul ruolo delle proteinees have focused attention on the role of VDAC nella disfunzioneproteins in mitochondriale tipica di molte condizioni pat dysfunction typical of many pathologiche tra cui ictusal conditions including stroke, cancro, encefalomiopatie mitocer, mitochondriali e invecchiamento, nonché disturbi encephalomyopathies, and aging, as well as neurodegenerativie disorders [27][28][27,28].
L'Manalisi della spetss spectrometria di massa ha rivelato l'y analysis revealed the associazione tra VDAC e l'tion between VDACs and the ubiquitina ligasie Parkin. In presenza dice of damaged mitocondri danneggiati, come nel morbo di hondria, as in Parkinson, La’s disease, Parkina viene fosforilata da is phosphorylated by PINK1 e diand conseguenzaquently ubiquitina letes proteine che risiedono sull'OMM, prendendo di mira i mitocondri per la s that reside on the OMM, targeting the mitochondria for degradazionetion. Parkin è una is a cytosolic proteina citosolica ma trasloca nei but translocates to the mitocondri per partecipare ai meccanismi di controllo della hondria to participate in mitochondrial qualità mitocondriale. Le proteine VDAC rappresentano un sito di attracco di Parkin sui mitocondri difettosi [29]y control mechanisms.
Moreover, VDAC1 represents the main docking site at the mitochondrial level for misfolded and aggregated proteins, a common feature of neurodegenerative disorders known as proteinopathies, such as Alzheimer’s disease (AD), proteins represent a docking site of Parkinson’s disease (PD), Creutzfeldt–Jacob disease (CJD), dementia with Lewy bodies (DLB), Huntington disease (HD), and amyotrophic lateral sclerosis (ALS) on defective mitochondria [29].
[30].
For example, in AD post-mortem brains, in neuroblastoma cells and in an AD mouse model, a direct association was demonstrated between VDAC1, specifically its N-terminal region, and hyper-phosphorylated Tau but also with amyloid beta (Aβ), both in its monomeric and oligomeric forms [31]. These interactions can have a dramatic effect on mitochondrial functions in AD neuron because they block the PTP formation, disrupt the transport of mitochondrial proteins and metabolites, and impair gating, conductance, and physiological interactome of VDACs [32].
In Parkinson’s disease, α-synuclein directly interacts with mitochondria, blocks VDAC1, and impairs metabolite fluxes leading, consequently, to an energetic crisis able to compromise cell viability [33].
In ALS, several SOD1 mutants are able to bind VDAC1 [34]. This interaction impairs ATP/ADP exchange, VDAC1 conductance and mitochondrial membrane potential. Recently, the competition between SOD1G93A and HK1 was demonstrated in binding VDAC1, in NSC34 motor-neuron cell lines [35].
In literature, the role of VDAC1 in neurodegeneration is rather well known; however, the involvement of the other two isoforms in these pathways remains poorly defined. This is likely associated with the relative abundance of VDAC1 compared to other isoforms which are more difficult to isolate in pure form.
Recent studies demonstrated that Cytoskeleton-associated protein 4 (CKAP4), a palmitoylated type II transmembrane protein localized to the endoplasmic reticulum (ER), regulates mitochondrial functions through an interaction with VDAC2 at ER-mitochondria contact sites [36].
VDAC2 binds inositol trisphosphate receptors (IP3R) and regulates the release of Ca2+ from the ER. In addition, several other interaction partners have been reported for VDAC2 isoform, which imply its effect in multiple cellular functions. Specifically, VDAC2 has been linked to many cellular proteins, including apoptotic factors as Bak and Bax, StAR, Metaxin2, eNOS (nitric oxide synthesize), GSK3β, tubulin, and Mcl1 [19]. In addition, VDAC2 and RACK1 (receptor of activated protein kinase C1) function as receptors for lymphocystis disease virus (LCDV) and for bursal disease virus in host cells [37].
VDAC2 together with VDAC3 binds Erastin, the activator of ferroptosis, a new pathway that regulates cell death characterized by the iron-dependent accumulation of lipid hydroperoxides. Interaction between VDAC2/3 and Erastin results in degradation of the channels following activation of ubiquitin protein ligase Nedd4 [38].
Finally, the VDAC3 isoform is associated with cytosolic proteins as tubulins and cytoskeletal proteins, stress sensors, chaperones, and proteasome components, redox-mediating enzymes such as protein disulfide isomerase [39].
2. Proteomics of VDAC Isoforms
2.1. Sample Preparation
Sample preparation has a profound effect on the final results of a proteomic workflow. Protein extraction methods and protein separation techniques should provide an unbiased and reliable map representative of all proteins present in a specific sample. The different extraction and fractionation approaches are based on proteins physicochemical and structural characteristics, such as molecular weight, solubility, hydrophobicity, and isoelectric point. A specific protocol has to be optimized for each particular sample, to maximize protein recovery and minimize the possible proteolysis and amino acid modifications. For these reasons, there is no universal extraction protocol and not a unique buffer composition. Regarding the extraction method, the different strategies available need to be compatible with both the amount of the processed material and the subsequent analytical approach (i.e., separation or MS).
The structural characterization of VDACs presents challenging issues due to their very high hydrophobicity, low solubility, and the impossibility to separate them from other mitochondrial proteins of similar hydrophobicity and to easily isolate each single isoform. In fact, isolation of VDACs has been possible exclusively for plant VDAC isoforms by chromatofocusing, thanks to the absence of phosphorylation sites in their structure [40]. Consequently, it is necessary to analyze them as components of a relatively complex mixture.
A bottom-up proteomic approach was used to investigate the VDAC3 from rat liver mitochondria (rVDAC3) [41]. According with a standard procedure [42], mitochondria were extracted and lysed with a buffer containing 3% Triton X-100 at pH 7.0. The VDAC proteins were partially purified by hydroxyapatite (HTP) chromatography, which allows to obtain a VDACs enriched fraction which comprises also other mitochondria hydrophobic proteins. After precipitation with cold acetone, the protein pellet was solubilized in SDS buffer and loaded on a 17% polyacrylamide gel (1D-SDS-PAGE). The bands in the range 30–35 kDa were manually excised from the gel, cut in small pieces, and subjected to reduction with DTT and alkylation by addition of IAA. Finally, the reduced and carboxyamidomethylated proteins were in gel-digested using trypsin and chymotrypsin, and the resulting peptide mixtures were analyzed by nUHPLC/HRMS [41]. MS data showed that rVDAC3 was found in the whole range 30–35 kDa, together with other proteins, mainly VDAC1, VDAC2, and several other mitochondrial proteins. The reason for VDAC3 electrophoretic heterogeneity probably stems from (i) the different pattern of cysteine oxidations that can modify the protein mobility; (ii) the different amount and quality of cysteine oxidations in various molecules (“redox isomers”).
The gel-digestion procedure shows some disadvantages: (i) larger peptides can get trapped between the gel meshes and lost during the extraction phase of the peptides from the gel; (ii) the electrophoretic procedure itself could damage the samples and alter the redox state of the sulfur amino acids (due to possible over heating generated by the applied voltage and to the presence of residual quantities of the catalysts used for the polyacrylamide polymerization). Furthermore, electrophoresis requires a relatively high amount of sample and the utilization of dyes and detergents. These last molecules could interfere with subsequent MS analyses because these compounds are difficult to eliminate from the sample.
2-DE could potentially represent a useful alternative to 1-DE to improve the separation of VDAC isoforms, but its utilization presents other problems. Actually, this kind of proteins has been under-represented in 2-DE gels due to difficulties in extracting and solubilizing them in the isoelectric focusing sample buffer. In fact, the most effective solubilizing agent for highly hydrophobic membrane proteins is SDS, but this detergent is incompatible with 2-DE. In addition to the difficulties in entering IPG (immobilized pH gradient) gels, membrane proteins tend to precipitate at their isoelectric point during IEF. Furthermore, their tendency to absorb the IPG matrix prevents their migration into the SDS-PAGE gel.
An improvement in the rVDAC3 mass spectrometric analysis was obtained following the introduction of a gel-free shotgun proteomic approach [41]. According to this procedure, to avoid any possible artefact due to air exposure and manipulations, reduction/alkylation was carried out before VDACs purification from the mitochondria. Afterwards, all the proteins present in the HTP eluate, without previous electrophoretic separation, were purified from non-protein contaminating molecules with the PlusOne 2-D Clean-Up kit, and the desalted protein pellet was then re-dissolved in ammonium bicarbonate containing RapiGest SF to improve the solubility. In fact, this surfactant makes the proteins more susceptible to enzymatic cleavage without modifying the sample or inhibiting endoprotease activity. Furthermore, the RapiGest SF is compatible with enzymes such as trypsin or chymotrypsin and does not influence subsequent MS analysis because it can be easily removed in acidic conditions.
Separate aliquots of reduced and alkylated proteins were then subjected to digestion with modified porcine trypsin and chymotrypsin. In this experiment, every protein in the HTP eluate was digested, producing a very complex peptide mixture, which was finally analyzed by nUHPLC/HR nESI-MS/MS.
The new “in solution-digestion” protocol associated with nUHPLC/HR ESI-MS/MS allowed to extend the coverage of the rat and human VDACs sequences with respect to that obtained with the previous procedure [43], so that it was possible to completely cover the rat and human VDAC1 sequences and almost completely the rat and human VDAC2 and VDAC3 sequences [22][41][44]. It should be noted that the short regions not identified in VDAC2 and VDAC3 correspond to small tryptic or chymotryptic peptides or even to single amino acids, which cannot be detected in LC/MS analysis.
Moreover, by means of this new procedure a detailed characterization of PTMs of the three VDACs was obtained (see next paragraphs).
2.2. Mass Spectrometry Analysis of Post-Translational Modifications
The mammalian proteome is vastly more complex than the related genome. The reasons for this difference reside both in the molecular mechanisms that allow a single gene to encode for multiple proteins (genomic recombination, transcription initiation at alternative promoters, differential transcription termination, and alternative splicing of the transcript) and in the post-translational modifications (PTMs) which represent a wide range of chemical changes that proteins can undergo after synthesis. They include the specific cleavage of protein precursors, the covalent addition or removal of low-molecular weight groups (i.e., acetylation, glycosylation, hydroxylation, phosphorylation, ubiquitination) and the formation of disulfide bonds or other redox modifications [45][46][47].
PTMs play crucial roles in cell biology since they can change protein physical or chemical properties, activity, localization, and/or stability. Traditionally, PTMs have been identified by Edman degradation, amino acid analysis, isotopic labeling, or immunochemistry. Within recent years, MS has proven to be extremely useful in PTM discovery. Post-translationally modified amino acids always have a different molecular mass than the original, unmodified residues and this mass increment or deficit is usually the basis for the detection and characterization of PTM by MS (commonly by LC-ESI-MS/MS).
MS has several advantages for characterization of PTMs, including (i) very high sensitivity; (ii) discovery of novel PTMs; (iii) ability to identify PTMs and the modified sites, even in complex protein mixtures; and (iv) ability to quantify the relative changes in PTM occupancy at distinct sites. None of the other techniques provide all these features, so the greater majority of the known PTMs have been described by MS [48].
To improve sensibility, several methods have been developed to enrich the samples in proteins or peptides with specific PTMs prior to MS/MS analysis, such as anti-pY antibodies, IMAC (immobilized metal affinity chromatography) and TiO2 for phosphorylation [49][50], affinity capture with lectins for glycosylated proteins [51], and resin coupled with anti-acetyl-lysine for acetylated proteins [52]. Although, as previously described, isolation of single isoforms of VDACs cannot be obtained, application of combined HPLC and high-resolution ESI-MSMS analysis has resulted in the identification of several PTM in these proteins. In the following, a summary of the MS-based PTMs characterized in VDACs is reported and the respective biological significance discussed. These results are resumed in Table 1 and Figure 1.
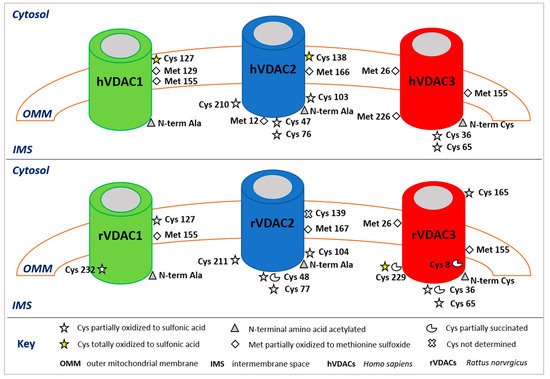

Figure 1. Post-translational modifications of human (upper panel) and rattus (lower panel) VDAC isoforms. The image shows only the modified amino acids and their positions with respect to the cytosol, the outer mitochondrial membrane (OMM), and the intermembrane space (IMS). In the rVDAC1 Cys232 faces the aqueous inside of the pore; in the rVDAC3 Cys8 is located inside of the pore.
Table 1. Post-translational modifications in VDAC isoforms obtained using mass spectrometry. PTM type, mass shift (Da), source of the sample, modified residue, MS method and relative reference are reported. Studies are described by listing first author + year.
| ISOFORM | PTM Type | ΔMass (Da) | Source | Residue | Method | Study |
|---|---|---|---|---|---|---|
| VDAC1 | Protein N-terminal acetylation | 42.0106 | Rat liver | Ala 2 | nUHPLC/high resolution nESI-MS/MS in a Q-QT-qIT MS |
Saletti et al., 2018 |
| HAP1 cells | Ala 2 | Pittalà et al., 2020 | ||||
| Acetylation | 42.0106 | Mouse liver | Lys 33, 41, 74, 234 | nHPLC MS/MS in an LTQ MS | Kim et al., 2006 | |
| Lys 41, 122, 132 | nHPLC MS/MS in an LTQ 2D ion-trap MS |
Schwer et al., 2009 | ||||
| Mouse liver and heart | Lys 237 | UPLC Velos-FT MS | Yang et al., 2011 | |||
| Human liver | Lys 28 | LC/LC-MS/MS in an FTICR/MS | Zhao et al., 2010 | |||
| Oxidation | 15.9949 | Rat liver | Met 155 | LC/LC-MS/MS in an FTICR/MS | Guan et al., 2003 | |
| nUHPLC/high resolution nESI-MS/MS in a Q-QT-qIT MS |
Saletti et al., 2018 | |||||
| HAP1 cells | Met 129, 155 | Pittalà et al., 2020 | ||||
| Trioxidation | 47.9847 | Rat liver | Cys 127, 232 | Saletti et al., 2018 | ||
| HAP1 cells | Cys 127 | Pittalà et al., 2020 | ||||
| Phosphorylation | 79.9663 | Rat liver | Ser 12, 136 | HPLC MS/MS in an LTQ MS | Distler et al., 2007 | |
| Mouse liver | Ser 117 | nHPLC MS/MS in an LTQ MS | Lee et al., 2007 | |||
| HeLa cells | Ser 101, 102, 104, Thr 107 |
nHPLC MS/MS in an LTQ-Orbitrap MS |
Olsen et al., 2006 | |||
| Mouse brain | Tyr 80, 208 | LC-MS/MS in an LTQ FT MS | Ballif et al., 2008 | |||
| VDAC2 | Protein N-terminal acetylation | 42.0106 | Rat liver | Ala 2 | nUHPLC/high resolution nESI-MS/MS in a Q-QT-qIT MS |
Saletti et al., 2018 |
| HAP1 cells | Ala 2 | Pittalà et al., 2020 | ||||
| Acetylation | 42.0106 | Mouse liver | Lys 32, 75 | nHPLC MS/MS in an LTQ MS | Kim et al., 2006 | |
| Lys 121 | nHPLC MS/MS in an LTQ 2D ion-trap MS |
Schwer et al., 2009 | ||||
| Oxidation | 15.9949 | Rat liver | Met 167 | nUHPLC/high resolution nESI-MS/MS in a Q-QT-qIT MS |
Saletti et al., 2018 | |
| HAP1 cells | Met 12, 166 | Pittalà et al., 2020 | ||||
| Trioxidation | 47.9847 | Rat liver | Cys 48, 77, 104, 211 | Saletti et al., 2018 | ||
| HAP1 cells | Cys 47, 76, 103, 138, 210 | Pittalà et al., 2020 | ||||
| Succination | 116.0110 | Mouse brain | Cys 48, 77 | LC-nESI-MS/MS in an LTQ-Orbitrap MS |
Piroli et al., 2016 | |
| Rat liver | Cys 48 | nUHPLC/high resolution nESI-MS/MS in a Q-QT-qIT MS |
Saletti et al., 2018 | |||
| Phosphorylation | 79.9663 | HeLa cells | Ser 115, Thr 118 | nHPLC MS/MS in an LTQ-Orbitrap MS |
Olsen et al., 2006 | |
| Rat liver | Thr 109 | SCX-RP-MS/MS in an LTQ-Orbitrap MS |
Deng et al., 2010 | |||
| Rat liver | Tyr 237 | HPLC MS/MS in an LTQ MS | Distler et al., 2007 | |||
| Mouse brain | Tyr 207 | LC-MS/MS in an LTQ FT MS | Ballif et al., 2008 | |||
| VDAC3 | Protein N-terminal acetylation | 42.0106 | Rat liver | Cys 2 | nUHPLC/high resolution nESI-MS/MS in a Q-QT-qIT MS |
Saletti et al., 2016 |
| HAP1 cells | Cys 2 | Pittalà et al., 2020 | ||||
| Acetylation | 42.0106 | Mouse liver | Lys 20, 61, 226 | nHPLC MS/MS in an LTQ MS | Kim et al., 2006 | |
| Lys 63, 109 | nHPLC MS/MS in an LTQ 2D ion-trap MS |
Schwer et al., 2009 | ||||
| Human liver | Lys 28 | LC/LC-MS/MS in an FTICR-MS | Zhao et al., 2010 | |||
| Oxidation | 15.9949 | Rat liver | Met 26, 155 | nUHPLC/high resolution nESI-MS/MS in a Q-QT-qIT MS |
Saletti et al., 2016 | |
| HAP1 cells | Met 26, 155, 226 | Pittalà et al., 2020 | ||||
| Trioxidation | 47.9847 | Rat liver | Cys 36, 65, 165, 229 | Saletti et al., 2016 | ||
| HAP1 cells | Cys 36, 65 | Pittalà et al., 2020 | ||||
| Succination | 116.0110 | Rat liver | Cys 8, 36, 229 | Saletti et al., 2018 | ||
| Phosphorylation | 79.9663 | Rat liver | Ser 241, Thr 33 | HPLC MS/MS in an LTQ MS | Distler et al., 2007 | |
| Mouse brain | Tyr 49 | LC-MS/MS in an LTQ FT MS | Ballif et al., 2008 |
References
- Shoshan-Barmatz, V.; De Pinto, V.; Zweckstetter, M.; Raviv, Z.; Keinan, N.; Arbel, N. VDAC, a multi-functional mitochon-drial protein regulating cell life and death. Mol. Aspects Med. 2010, 31, 227–285.
- Messina, A.; Reina, S.; Guarino, F.; De Pinto, V. VDAC isoforms in mammals. Biochim. Biophys. Acta 2012, 1818, 1466–1476.
- Young, M.J.; Bay, D.C.; Hausner, G.; Court, D.A. The evolutionary history of mitochondrial porins. BMC Evol. Biol. 2007, 7, 31.
- De Pinto, V.; Reina, S.; Gupta, A.; Messina, A.; Mahalakshmi, R. Role of cysteines in mammalian VDAC isoforms’ function. Biochim. Biophys. Acta Bioenerg. 2016, 1857, 1219–1227.
- Hiller, S.; Garces, R.G.; Malia, T.J.; Orekhov, V.Y.; Colombini, M.; Wagner, G. Solution structure of the integral human membrane protein VDAC-1 in detergent micelles. Science 2008, 321, 1206–1210.
- Bayrhuber, M.; Meins, T.; Habeck, M.; Becker, S.; Giller, K.; Villinger, S.; Vonrhein, C.; Griesinger, C.; Zweckstetter, M.; Zeth, K. Structure of the human voltage-dependent anion channel. Proc. Natl. Acad. Sci. USA 2008, 105, 15370–15375.
- Ujwal, R.; Cascio, D.; Colletier, J.P.; Faham, S.; Zhang, J.; Toro, L.; Ping, P.; Abramson, J. The crystal structure of mouse VDAC1 at 2.3 angstrom resolution reveals mechanistic insights into metabolite gating. Proc. Natl. Acad. Sci. USA 2008, 105, 17742–17747.
- Schredelseker, J.; Paz, A.; Lopez, C.J.; Altenbach, C.; Leung, C.S.; Drexler, M.K.; Chen, J.N.; Hubbell, W.L.; Abramson, J. High resolution structure and double electron-electron resonance of the zebrafish voltage-dependent anion channel 2 reveal an oligomeric population. J. Biol. Chem. 2014, 289, 12566–12577.
- Amodeo, G.F.; Scorciapino, M.A.; Messina, A.; De Pinto, V.; Ceccarelli, M. Charged residues distribution modulates selec-tivity on the open state of human isoforms of the voltage dependent anion-selective channel. PLoS ONE 2014, 9, e103879.
- De Pinto, V.; Guarino, F.; Guarnera, A.; Messina, A.; Reina, S.; Tomasello, M.F.; Palermo, V.; Mazzoni, C. Characterization of human VDAC isoforms: A peculiar function for VDAC3? Biochim. Biophys. Acta-Bioenergetics 2010, 1797, 1268-1275.
- Naghdi, S.; Várnai, P.; Hajnóczky, G. Motifs of VDAC2 required for mitochondrial Bak import and tBid-induced apoptosis. Proc. Natl. Acad. Sci. USA 2015, 112, E5590–E5599.
- Shoshan-Barmatz, V.; Maldonado, E.N.; Krelin, Y. VDAC1 at the crossroads of cell metabolism, apoptosis and cell stress. Cell stress 2017, 1, 1.
- Shoshan-Barmatz, V.; Pittala, S.; Mizrachi, D. VDAC1 and the TSPO: Expression, Interactions, and Associated Functions in Health and Disease States. Int. J. Mol. Sci. 2019, 20, 3348.
- Kerner, J.; Lee, K.; Tandler, B.; Hoppel, C.L. VDAC proteomics: Post-translation modifications. Biochim. Biophys. Acta 2012, 1818, 1520–1525.
- Geula, S.; Ben-Hail, D.; Shoshan-Barmatz, V. Structure-based analysis of vdac1: N-terminus location, translocation, channel gating and association with anti-apoptotic proteins. Biochem. J. 2012, 444, 475–485.
- Abu-Hamad, S.; Arbel, N.; Calo, D.; Arzoine, L.; Israelson, A.; Keinan, N.; Ben-Romano, R.; Friedman, O.; Shoshan-Barmatz, V. The vdac1 n-terminus is essential both for apoptosis and the protective effect of anti-apoptotic proteins. J. Cell Sci. 2009, 122, 1906–1916.
- Shi, Y.; Chen, J.;Weng, C.; Chen, R.; Zheng, Y.; Chen, Q.; Tang, H. Identification of the protein-protein contact site and inter-action mode of human vdac1 with bcl-2 family proteins. Biochem. Biophys. Res. Commun. 2003, 305, 989–996.
- Naghdi, S.; Hajnóczky, G. VDAC2-specific cellular functions and the underlying structure. Biochim. Biophys. Acta 2016, 1863, 2503–2514.
- Shoshan-Barmatz, V.; Ben-Hail, D. VDAC, a multi-functional mitochondrial protein as a pharmacological target. Mitochon-drion 2012, 12, 24–34.
- Abu-Hamad, S.; Zaid, H.; Israelson, A.; Nahon, E.; Shoshan-Barmatz, V. Hexokinase-I protection against apoptotic cell death is mediated via interaction with the voltage-dependent anion channel-1: Mapping the site of binding. J. Biol. Chem. 2008, 283, 13482–13490.
- De Pinto, V.; al Jamal, J.A.; Palmieri, F. Location of the dicyclohexylcarbodiimide-reactive glutamate residue in the bovine heart mitochondrial porin. J. Biol. Chem. 1993, 268, 12977–12982.
- Pittalà, M.G.G.; Saletti, R.; Reina, S.; Cunsolo, V.; De Pinto, V.; Foti, S. A High Resolution Mass Spectrometry Study Reveals the Potential of Disulfide Formation in Human Mitochondrial Voltage-Dependent Anion Selective Channel Isoforms (hVDACs). Int. J. Mol. Sci. 2020, 21, 1468.
- Budelier, M.M.; Cheng, W.W.L.; Bergdoll, L.; Chen, Z.W.; Janetka, J.W.; Abramson, J.; Krishnan, K.; Mydock-McGrane, L.; Covey, D.F.; Whitelegge, J.P.; et al. Photoaffinity labeling with cholesterol analogues precisely maps a cholesterol-binding site in voltage-dependent anion channel-1. J. Biol. Chem. 2017, 292, 9294–9304.
- Gatliff, J.; East, D.; Crosby, J.; Abeti, R.; Harvey, R.; Craigen, W.; Parker, P.; Campanella, M. Tspo interacts with vdac1 and triggers a ros-mediated inhibition of mitochondrial quality control. Autophagy 2015, 10, 2279–2296.
- Mueller, B.K.; Subramaniam, S.; Senes, A. A frequent, GxxxG-mediated, transmembrane association motif is optimized for the formation of interhelical Cα-H hydrogen bonds. Proc. Natl. Acad. Sci. USA 2014, 111, E888–E895.
- Bernardi, P.; Di Lisa, F.; Fogolari, F.; Lippe, G. From ATP to PTP and back: A dual function for the mitochondrial ATP syn-thase. Circ. Res. 2015, 116, 1850–1862.
- Reina, S.; Guarino, F.; Magrì, A.; De Pinto, V. VDAC3 as a potential marker of mitochondrial status is involved in cancer and pathology. Front. Oncol. 2016, 6, 264.
- Magrì, A.; Messina, A. Interactions of VDAC with Proteins Involved in Neurodegenerative Aggregation: An Opportunity for Advancement on Therapeutic Molecules. Curr. Med. Chem. 2017, 24, 4470–4487.
- Sun, Y.; Vashisht, A.A.; Tchieu, J.; Wohlschlegel, J.A.; Dreier, L. Voltage-dependent Anion Channels (VDACs) recruit Parkin to defective mitochondria to promote mitochondrial autophagy. J. Biol. Chem. 2012, 287, 40652–40660.
- Sheikh, S.; Safia; Haque, E.; Snober, S.M. Neurodegenerative diseases: Multifactorial conformational diseases and their therapeutic interventions. J. Neurodegen. Dis. 2013, 2013, 563481.
- Smilansky, A.; Dangoor, L.; Nakdimon, I.; Ben-Hail, D.; Mizrachi, D.; Shoshan-Barmatz, V. The voltage-dependent anion channel 1 mediates amyloid beta toxicity and represents a potential target for Alzheimer's disease therapy. J. Biol. Chem. 2015, 290, 30670–30683.
- Hemachandra Reddy, P. Is the mitochondrial outer membrane protein VDAC1 therapeutic target for Alzheimer’s disease? Biochim. Biophys. Acta 2013, 1832, 67–75.
- Rostovtseva, T.K.; Gurnev, P.A.; Protchenko, O.; Hoogerheide, D.P.; Yap, T.L.; Philpott, C.C.; Lee, J.C.; Bezrukov, S.M. α-Synuclein shows high affinity interaction with Voltage-dependent Anion Channel, suggesting mechanisms of mitochon-drial regulation and toxicity in Parkinson Disease. J. Biol. Chem. 2015, 290, 18467–18477.
- Israelson, A.; Arbel, N.; Da Cruz, S.; Ilieva, H.; Yamanaka, K.; Shoshan-Barmatz, V.; Cleveland, D.W. Misfolded mutant SOD1 directly inhibits VDAC1 conductance in a mouse model of inherited ALS. Neuron 2010, 67, 575–587.
- Magrì, A.; Belfiore, R.; Reina, S.; Tomasello, M.F.; Di Rosa, M.C.; Guarino, F.; Leggio, L.; De Pinto, V.; Messina, A. Hexoki-nase I N-terminal based peptide prevents the VDAC1-SOD1G93A interaction and re-establishes ALS cell viability. Sci. Rep. 2016, 6, 34802.
- Harada, T.; Sada, R.; Osugi, Y.; Matsumoto, S.; Matsuda, T.; Hayashi-Nishino, M.; Nagai, T.; Harada, A.; Kikuchi, A. Pal-mitoylated CKAP4 regulates mitochondrial functions through an interaction with VDAC2 at ER–mitochondria contact sites. J. Cell Sci. 2020, 133, jcs249045.
- Zhong, Y.; Tang, X.; Sheng, X.; Xing, J.; Zhan, W. Voltage-Dependent Anion Channel Protein 2 (VDAC2) and Receptor of Ac-tivated Protein C Kinase 1 (RACK1) Act as Functional Receptors for Lymphocystis Disease Virus Infection. J. Virol. 2019, 93, e00122-19.
- Yang, Y.; Luo, M.; Zhang, K.; Zhang, J.; Gao, T.; O’Connell, D.; Yao, F.; Mu, C.; Cai, B.; Shang, Y.; Chen, W. Nedd4 ubiq-uitylates VDAC2/3 to suppress erastin-induced ferroptosis in melanoma. Nat. Commun. 2020, 11, 433.
- Messina, A.; Reina, S.; Guarino, F.; Magrì, A.; Tomasello, F.; Clark, R.E.; Ramsayc, R.R.; De Pinto, V. Live cell interactome of the human voltage dependent anion channel 3 (VDAC3) revealed in HeLa cells by affinity purification tag technique. Mol. BioSyst. 2014, 10, 2134–2145.
- Abrecht, H.; Wattiez, R.; Ruysschaert, J.M.; Homblé, F. Purification and characterization of two Voltage-Dependent Anion Channel Isoforms from plant seeds. Plant Physiol. 2000, 124, 1181–1190.
- Saletti, R.; Reina, S.; Pittalà, M.G.G.; Belfiore, R.; Cunsolo, V.; Messina, A.; De Pinto, V.; Foti, S. High resolution mass spec-trometry characterization of the oxidation pattern of methionine and cysteine residues in rat liver mitochondria Volt-age-Dependent Anion selective Channel 3 (VDAC3). Biochim. Biophys. Acta-Biomembr. 2017, 1859, 301–311.
- De Pinto, V.; Prezioso, G.; Palmieri, F. A simple and rapid method for the purification of the mitochondrial porin from mammalian tissues. Biochim. Biophys. Acta 1987, 905, 499–502.
- Distler, A.M.; Kerner, J.; Peterman, S.M.; Hoppel, C.L. A targeted proteomic approach for the analysis of rat liver mito-chondrial outer membrane proteins with extensive sequence coverage. Anal. Biochem. 2006, 356, 18–29.
- Saletti, R.; Reina, S.; Pittalà, M.G.G.; Magrì, A.; Cunsolo, V.; Foti, S.; De Pinto, V. Post-translational modifications of VDAC1 and VDAC2 cysteines from rat liver mitochondria. Biochim. Biophys. Acta-Bioenerg. 2018, 1859, 806–816.
- Wang, Y.; Peterson, S.; Loring, J. Protein post-translational modifications and regulation of pluripotency in human stem cells. Cell Res. 2014, 24, 143–160.
- Duan, G.; Walther, D. The roles of post-translational modifications in the context of protein interaction networks. PLoS Comput. Biol. 2015, 11, e1004049.
- Reina, S.; Pittalà, M.G.G.; Guarino, F.; Messina, A.; De Pinto, V.; Foti, S.; Saletti, R. Cysteine oxidations in mitochondrial membrane proteins: The case of VDAC isoforms in mammals. Front. Cell Dev. Biol. 2020, 8, 397.
- Jensen, O.N. Modification-specific proteomics: Characterization of posttranslational modifications by mass spectrometry. Curr. Opin. Chem. Biol. 2004, 8, 33–41.
- Corthals, G.L.; Aebersold, R.; Goodlett, D.R. Identification of phosphorylation sites using microimmobilized metal affinity chromatography. Meth. Enzymol. 2005, 405, 66–81.
- Larsen, M.R.; Thingholm, T.E.; Jensen, O.N.; Roepstorff, P.; Jørgensen, T.J. Highly selective enrichment of phosphorylated peptides from peptide mixtures using titanium dioxide microcolumns. Mol. Cell. Proteomics 2005, 4, 873–886.
- Yang, Z.; Hancock, W.S. Approach to the comprehensive analysis of glycoproteins isolated from human serum using a mul-ti-lectin affinity column. J. Chromatogr. A 2004, 1053, 79–88.
- Kim, S.C.; Sprung, R.; Chen, Y.; Xu, Y.; Ball, H.; Pei, J.; Cheng, T.; Kho, Y.; Xiao, H.; Xiao, L.; et al. Substrate and functional diversity of lysine acetylation revealed by a proteomics survey. Mol. Cell. 2006, 23, 607–618.
More
