Theoretical methods for finding and predicting new antimicrobial peptides (AMPs), based on the use of specially designed programs for these purposes, are making an increasing contribution to the development of new AMPs. Machine learning methods are also used for the prediction. Artificial neural networks perform highly accurate predictions based on rules from databases of antimicrobial peptides. Recently published data on the development of new AMP drugs based on a combination of molecular design and genetic engineering approaches are presented. This review examines AMP development strategies from the perspective of the current high prevalence of antibiotic-resistant bacteria, and the potential prospects and challenges of using AMPs against infection caused by severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2). In turn, we proposed another strategy for the development of new AMPs based on predicting amyloidogenic regions in a protein molecule sequence.
- antimicrobial peptides (AMPs)
- prediction of AMPs
- development of AMPs
- antibacterial peptides
- antiviral peptides
- SARS-CoV-2
- spike protein
- amyloidogenic regions
- protein aggregation
- coaggregation
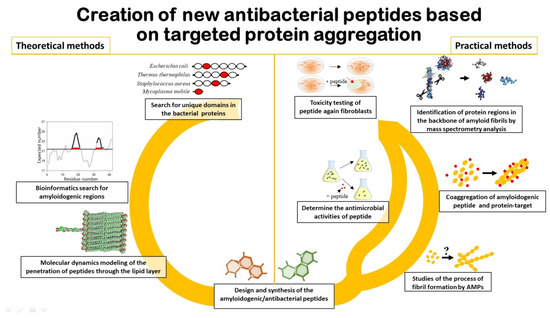
1. Types of Programs for Prediction and Development of New AMPs
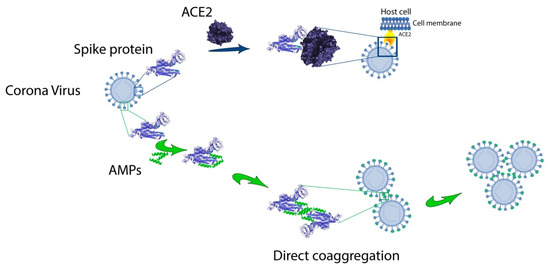
2. SARS-CoV-2 Like an Object for Prediction and Development of New AMPs
| Software | Amyloidogenic Regions for SARS-CoV-2 Spike Protein (https://www.uniprot.org/uniprot/P0DTC2) |
|---|---|
| AGGRESCAN (UAB, Barcelona, Spain http://bioinf.uab.es/aggrescan/) |
1–8, 10–16, 31–38, 40–47, 50–56, 58–70, 86–91, 100–109, 114–133, 140–145, 162–167, 189–208, 231–247, 260–271, 303–311, 331–336, 338–354, 362–383, 390–400, 428–437, 448–457, 484–490, 508–520, 584–588, 590–599, 610–617, 666–671, 688–698, 716–727, 729–745, 751–759, 761–768, 783–787, 797–805, 817–833, 853–861, 864–870, 872–914, 961–982, 1000–1013, 1044–1052, 1058–1068, 1097–1101, 1124–1137, 1171–1179, 1207–1249. |
| PASTA 2.0 (University of Padova, Padova, Italy http://protein.bio.unipd.it/pasta2/) |
1214–1244. |
| Waltz (Switch Laboratory, Leuven, Belgium http://waltz.switchlab.org/index.cgi) |
1–9, 88–95, 115–137, 141–146, 199–204, 261–271, 275–280, 365–378, 447–455, 485–496, 537–544, 608–618, 689–697, 703–727, 798–806, 913–940, 969–974, 1003–1015, 1063–1068, 1117–1122, 1175–1179, 1206–1233. |
| FoldAmyloid (Institute of Protein Research of the Russian Academy of Sciences, Pushchino, Russia http://bioinfo.protres.ru/fold-amyloid/) |
1–10, 34–38, 53–69,100–108,116–120, 126–135, 141–146, 174–178, 190–195, 235–244, 264–269, 274–278, 327–331, 348–353, 366–370, 389–395, 431– 436, 450–457, 487–492, 507–519, 539–543, 560–564, 582–586, 609–613, 691–697, 718–722, 738–743, 751– 755, 780–784, 799–804, 819–825, 875–880, 894–907, 993–999, 1005–1014, 1047–1051, 1060–1067, 1101–1105, 1127–1132, 1210–1239. |
References
- B. Agerberth; H. Gunne; J. Odeberg; P. Kogner; H. G. Boman; G. H. Gudmundsson; FALL-39, a putative human peptide antibiotic, is cysteine-free and expressed in bone marrow and testis.. Proceedings of the National Academy of Sciences 1995, 92, 195-199, 10.1073/pnas.92.1.195.
- Nannette Y. Yount; Michael R. Yeaman; Multidimensional signatures in antimicrobial peptides. Proceedings of the National Academy of Sciences 2004, 101, 7363-7368, 10.1073/pnas.0401567101.
- Laurent Coquet; Jolanta Kolodziejek; Thierry Jouenne; Norbert Nowotny; Jay D. King; J. Michael Conlon; Peptidomic analysis of the extensive array of host-defense peptides in skin secretions of the dodecaploid frog Xenopus ruwenzoriensis (Pipidae). Comparative Biochemistry and Physiology Part D: Genomics and Proteomics 2016, 19, 18-24, 10.1016/j.cbd.2016.04.006.
- Xinwang Yang; Wen-Hui Lee; Yun Zhang; Extremely Abundant Antimicrobial Peptides Existed in the Skins of Nine Kinds of Chinese Odorous Frogs. Journal of Proteome Research 2011, 11, 306-319, 10.1021/pr200782u.
- Brian C. Schutte; Joseph P. Mitros; Jennifer A. Bartlett; Jesse D. Walters; Hong Peng Jia; Michael J. Welsh; Thomas L. Casavant; Paul B. McCray; Discovery of five conserved -defensin gene clusters using a computational search strategy. Proceedings of the National Academy of Sciences 2002, 99, 2129-2133, 10.1073/pnas.042692699.
- Margherita Zanetti; The role of cathelicidins in the innate host defenses of mammals.. Current Issues in Molecular Biology 2005, 7, 179-96.
- Yifeng Li; Xia Li; He Li; Oksana Lockridge; Guangshun Wang; A novel method for purifying recombinant human host defense cathelicidin LL-37 by utilizing its inherent property of aggregation. Protein Expression and Purification 2007, 54, 157-165, 10.1016/j.pep.2007.02.003.
- Christopher D. Fjell; Robert E.W. Hancock; Artem Cherkasov; AMPer: a database and an automated discovery tool for antimicrobial peptides. Bioinformatics 2007, 23, 1148-1155, 10.1093/bioinformatics/btm068.
- Alessandro Tossi; Molecular Diversity in Gene-Encoded, Cationic Antimicrobial Polypeptides. Current Pharmaceutical Design 2002, 8, 743-761, 10.2174/1381612023395475.
- Guangshun Wang; Structural Analysis of Amphibian, Insect, and Plant Host Defense Peptides Inspires the Design of Novel Therapeutic Molecules. Host Defense Peptides and Their Potential as Therapeutic Agents 2016, 9, 229-252, 10.1007/978-3-319-32949-9_9.
- Maire Begley; Paul D. Cotter; Colin Hill; R. Ross; Identification of a Novel Two-Peptide Lantibiotic, Lichenicidin, following Rational Genome Mining for LanM Proteins. Applied and Environmental Microbiology 2009, 75, 5451-5460, 10.1128/aem.00730-09.
- Mangal Singh; Dipti Sareen; Novel LanT Associated Lantibiotic Clusters Identified by Genome Database Mining. PLOS ONE 2014, 9, e91352, 10.1371/journal.pone.0091352.
- James T. Morton; Stefan D. Freed; Shaun W. Lee; Iddo Friedberg; A large scale prediction of bacteriocin gene blocks suggests a wide functional spectrum for bacteriocins. BMC Bioinformatics 2015, 16, 1-9, 10.1186/s12859-015-0792-9.
- David J. Lynn; Rowan Higgs; Susan Gaines; Joanna Tierney; Tharappel James; Andrew T. Lloyd; Mario A. Fares; Grace Mulcahy; Cliona O’Farrelly; Bioinformatic discovery and initial characterisation of nine novel antimicrobial peptide genes in the chicken. Immunogenetics 2004, 56, 170-177, 10.1007/s00251-004-0675-0.
- André C. Amaral; Osmar N. Silva; Nathália C.C.R. Mundim; Maria J.A. De Carvalho; Ludovico Migliolo; Jose R.S.A. Leite; Maura V. Prates; Anamélia L. Bocca; Octávio L. Franco; Maria S.S. Felipe; et al. Predicting antimicrobial peptides from eukaryotic genomes: In silico strategies to develop antibiotics. Peptides 2012, 37, 301-308, 10.1016/j.peptides.2012.07.021.
- Marc Torrent; Victòria M. Nogués; Ester Boix; A theoretical approach to spot active regions in antimicrobial proteins. BMC Bioinformatics 2009, 10, 373-373, 10.1186/1471-2105-10-373.
- Roland Hellinger; Johannes Koehbach; Douglas E. Soltis; Eric J. Carpenter; Gane Ka-Shu Wong; Christian W Gruber; Peptidomics of Circular Cysteine-Rich Plant Peptides: Analysis of the Diversity of Cyclotides from Viola tricolor by Transcriptome and Proteome Mining. Journal of Proteome Research 2015, 14, 4851-4862, 10.1021/acs.jproteome.5b00681.
- Joon Ha Lee; Hoyong Chung; Yong Pyo Shin; In-Woo Kim; Sathishkumar Natarajan; Karpagam Veerappan; MinChul Seo; Junhyung Park; Jae Sam Hwang; Transcriptome Analysis of Psacothea hilaris: De Novo Assembly and Antimicrobial Peptide Prediction. Insects 2020, 11, 676, 10.3390/insects11100676.
- Sergei Yu. Grishin; Evgeniya I. Deryusheva; Andrey Machulin; Olga M. Selivanova; Anna V. Glyakina; Elena Yu. Gorbunova; Leila G. Mustaeva; Vyacheslav N. Azev; Valentina V. Rekstina; Tatyana S. Kalebina; et al.Alexey K. SurinOxana V. Galzitskaya Amyloidogenic Propensities of Ribosomal S1 Proteins: Bioinformatics Screening and Experimental Checking. International Journal of Molecular Sciences 2020, 21, 5199, 10.3390/ijms21155199.
- Oxana V. Galzitskaya; Sergiy O. Garbuzynskiy; Michail Yu. Lobanov; IS IT POSSIBLE TO PREDICT AMYLOIDOGENIC REGIONS FROM SEQUENCE ALONE?. Journal of Bioinformatics and Computational Biology 2006, 4, 373-388, 10.1142/s0219720006002004.
- Conchillo-Solé, O.; de Groot, N.S.; Avilés, F.X.; Vendrell, J.; Daura, X.; Ventura, S.; AGGRESCAN: A server for the prediction and evaluation of “hot spots” of aggregation in polypeptides. BMC Bioinform. 2007, 8, 65, 10.1186/1471-2105-8-65.
- Sergiy O. Garbuzynskiy; Michail Yu. Lobanov; Oxana V. Galzitskaya; FoldAmyloid: a method of prediction of amyloidogenic regions from protein sequence. Bioinformatics 2009, 26, 326-332, 10.1093/bioinformatics/btp691.
- Ian Walsh; Flavio Seno; Silvio C E Tosatto; Antonio Trovato; PASTA 2.0: an improved server for protein aggregation prediction. Nucleic Acids Research 2014, 42, W301-W307, 10.1093/nar/gku399.
- Mikael Oliveberg; Waltz, an exciting new move in amyloid prediction. Nature Methods 2010, 7, 187-188, 10.1038/nmeth0310-187.
- O. V. Galzitskaya; S. O. Garbuzynskiy; M. Yu. Lobanov; Prediction of natively unfolded regions in protein chains. Molecular Biology 2006, 40, 298-304, 10.1134/s0026893306020166.
- Tatyana B. Mamonova; Anna V. Glyakina; Oxana V. Galzitskaya; Maria G. Kurnikova; Stability and rigidity/flexibility—Two sides of the same coin?. Biochimica et Biophysica Acta (BBA) - Proteins and Proteomics 2013, 1834, 854-866, 10.1016/j.bbapap.2013.02.011.
- Niloofar Khairkhah; Mohammad Reza Aghasadeghi; Ali Namvar; Azam Bolhassani; Design of novel multiepitope constructs-based peptide vaccine against the structural S, N and M proteins of human COVID-19 using immunoinformatics analysis. PLoS ONE 2020, 15(10), e0240577, 10.1371/journal.pone.0240577.
- Peng Zhou; Xing-Lou Yang; Xian-Guang Wang; Ben Hu; Lei Zhang; Wei Zhang; Hao-Rui Si; Yan Zhu; Bei Li; Chao-Lin Huang; et al.Hui-Dong ChenJing ChenYun LuoHua GuoRen-Di JiangMei-Qin LiuYing ChenXu-Rui ShenXi WangXiao-Shuang ZhengKai ZhaoQuan-Jiao ChenFei DengLin-Lin LiuBing YanFa-Xian ZhanYan-Yi WangGeng-Fu XiaoZheng-Li Shi A pneumonia outbreak associated with a new coronavirus of probable bat origin. Nature Cell Biology 2020, 579, 270-273, 10.1038/s41586-020-2012-7.
- Hamed Memariani; Mojtaba Memariani; Hamideh Moravvej; Mohammad Shahidi-Dadras; Melittin: a venom-derived peptide with promising anti-viral properties. European Journal of Clinical Microbiology and Infections Diseases 2019, 39, 5-17, 10.1007/s10096-019-03674-0.
- Yangsheng Yu; Christopher L. Cooper; Guangshun Wang; M. Jane Morwitzer; Krishna Kota; Julie P. Tran; Steven B. Bradfute; Yan Liu; Jiayu Shao; Amanda K. Zhang; et al.Lindsey G. LuoSt. Patrick ReidSteven H. HinrichsKaihong Su Engineered Human Cathelicidin Antimicrobial Peptides Inhibit Ebola Virus Infection. iScience 2020, 23, 100999, 10.1016/j.isci.2020.100999.
- Yi Fan; Kai Zhao; Zhengli Shi; Peng Zhou; Bat Coronaviruses in China. Viruses 2019, 11, 210, 10.3390/v11030210.
- Yuan Huang; Chan Yang; Xin-Feng Xu; Wei Xu; Shuwen Liu; Structural and functional properties of SARS-CoV-2 spike protein: potential antivirus drug development for COVID-19. Acta Pharmacologica Sinica 2020, 41, 1141-1149, 10.1038/s41401-020-0485-4.
- Jiahua He; Huanyu Tao; Yumeng Yan; Sheng-You Huang; Yi Xiao; Molecular Mechanism of Evolution and Human Infection with SARS-CoV-2. Viruses 2020, 12, 428, 10.3390/v12040428.
- Alexandra C. Walls; Young-Jun Park; M. Alejandra Tortorici; Abigail Wall; Andrew T. McGuire; David Veesler; Structure, Function, and Antigenicity of the SARS-CoV-2 Spike Glycoprotein. Cell 2020, 181, 281-292.e6, 10.1016/j.cell.2020.02.058.
- Amanat Ali; Ranjit Vijayan; Dynamics of the ACE2–SARS-CoV-2/SARS-CoV spike protein interface reveal unique mechanisms. Scientific Reports 2020, 10, 1-12, 10.1038/s41598-020-71188-3.
- Jian Shang; Yushun Wan; Chuming Luo; Gang Ye; Qibin Geng; Ashley Auerbach; Fang Li; Cell entry mechanisms of SARS-CoV-2. Proceedings of the National Academy of Sciences 2020, 117, 11727-11734, 10.1073/pnas.2003138117.
- Jinyong Zhang; Huazong Zeng; Jiang Gu; Haibo Li; Lixin Zheng; Quan M. Zou; Progress and Prospects on Vaccine Development against SARS-CoV-2. Vaccines 2020, 8, 153, 10.3390/vaccines8020153.
- Rolando Cannalire; Irina Stefanelli; Carmen Cerchia; Andrea Rosario Beccari; Sveva Pelliccia; Vincenzo Summa; SARS-CoV-2 Entry Inhibitors: Small Molecules and Peptides Targeting Virus or Host Cells. International Journal of Molecular Sciences 2020, 21, 5707, 10.3390/ijms21165707.
- Debmalya Barh; Sandeep Tiwari; Bruno Silva Andrade; Marta Giovanetti; Eduardo Almeida Costa; Ranjith Kumavath; Preetam Ghosh; Aristóteles Góes-Neto; Luiz Carlos Junior Alcantara; Vasco Azevedo; et al. Potential chimeric peptides to block the SARS-CoV-2 spike receptor-binding domain. F1000Research 2020, 9, 576, 10.12688/f1000research.24074.1.
- Philippe Karoyan; Vincent Vieillard; Estelle Odile; Alexis Denis; Luis Gómez-Morales; Pascal Grondin; Olivier Lequin; An hACE2 peptide mimic blocks SARS-CoV-2 Pulmonary Cell Infection. bioRxiv 2020, none, none, 10.1101/2020.08.24.264077.
- Roujian Lu; Xiang Zhao; Juan Li; Peihua Niu; Bo Yang; Honglong Wu; Wenling Wang; Hao Song; Baoying Huang; Na Zhu; et al.Yuhai BiXuejun MaFaxian ZhanLiang WangTao HuHong ZhouZhenhong HuWeimin ZhouLi ZhaoJing ChenYao MengJi WangYang LinJianying YuanZhihao XieJinmin MaWilliam J LiuDayan WangWenbo XuEdward C HolmesGeorge F GaoGuizhen WuWeijun ChenWeifeng ShiWenjie Tan Genomic characterisation and epidemiology of 2019 novel coronavirus: implications for virus origins and receptor binding. The Lancet 2020, 395, 565-574, 10.1016/s0140-6736(20)30251-8.
- Frank L. Van De Veerdonk; Mihai G Netea; Marcel Van Deuren; Jos Wm Van Der Meer; Quirijn De Mast; Roger J Brüggemann; Hans Van Der Hoeven; Kallikrein-kinin blockade in patients with COVID-19 to prevent acute respiratory distress syndrome. eLife 2020, 9, e57555, 10.7554/elife.57555.
- Sandrine Belouzard; Victor C. Chu; Gary R. Whittaker; Activation of the SARS coronavirus spike protein via sequential proteolytic cleavage at two distinct sites. Proceedings of the National Academy of Sciences 2009, 106, 5871-5876, 10.1073/pnas.0809524106.
- Markus Hoffmann; Hannah Kleine-Weber; Stefan Pöhlmann; A Multibasic Cleavage Site in the Spike Protein of SARS-CoV-2 Is Essential for Infection of Human Lung Cells. Molecular Cell 2020, 78, 779-784.e5, 10.1016/j.molcel.2020.04.022.
- Kristian G. Andersen; Andrew Rambaut; W. Ian Lipkin; Edward C Holmes; Robert F. Garry; The proximal origin of SARS-CoV-2. Nature Medicine 2020, 26, 450-452, 10.1038/s41591-020-0820-9.
- Omid Tavassoly; Farinaz Safavi; Iman Tavassoly; Seeding Brain Protein Aggregation by SARS-CoV-2 as a Possible Long-Term Complication of COVID-19 Infection. ACS Chemical Neuroscience 2020, 11, 3704-3706, 10.1021/acschemneuro.0c00676.
- Alexey K. Surin; Sergei Yu. Grishin; Oxana V. Galzitskaya; Identification of Amyloidogenic Regions in the Spine of Insulin Fibrils. Biochemistry (Moscow) 2019, 84, 47-55, 10.1134/s0006297919010061.
- Shruti Mukherjee; Dipita Bhattacharyya; Anirban Bhunia; Host-membrane interacting interface of the SARS coronavirus envelope protein: Immense functional potential of C-terminal domain. Biophysical Chemistry 2020, 266, 106452-106452, 10.1016/j.bpc.2020.106452.
- Stanislav R. Kurpe; Sergei Yu. Grishin; Alexey K. Surin; Olga M. Selivanova; Roman S. Fadeev; Ylyana F. Dzhus; Elena Yu. Gorbunova; Leila G. Mustaeva; Vyacheslav N. Azev; Oxana V. Galzitskaya; et al. Antimicrobial and Amyloidogenic Activity of Peptides Synthesized on the Basis of the Ribosomal S1 Protein from Thermus Thermophilus. International Journal of Molecular Sciences 2020, 21, 6382, 10.3390/ijms21176382.
- Shuai Xia; Yun Zhu; Meiqin Liu; Qiaoshuai Lan; Wei Xu; Yanling Wu; TianLei Ying; Shuwen Liu; Zhengli Shi; Shibo Jiang; et al.Lu Lu Fusion mechanism of 2019-nCoV and fusion inhibitors targeting HR1 domain in spike protein. Cellular & Molecular Immunology 2020, 17, 765-767, 10.1038/s41423-020-0374-2.
- Shuai Xia; Meiqin Liu; Chao Wang; Wei Xu; Qiaoshuai Lan; Siliang Feng; Feifei Qi; Linlin Bao; Lanying Du; Shuwen Liu; et al.Chuan QinFei SunZhengli ShiYun ZhuShibo JiangLu Lu Inhibition of SARS-CoV-2 (previously 2019-nCoV) infection by a highly potent pan-coronavirus fusion inhibitor targeting its spike protein that harbors a high capacity to mediate membrane fusion. Cell Research 2020, 30, 343-355, 10.1038/s41422-020-0305-x.
- Kehu Yuan; Ling Yi; Jian Chen; Xiuxia Qu; Tingting Qing; Xi Rao; Pengfei Jiang; Jianhe Hu; Zikai Xiong; Yuchun Nie; et al.Xuanling ShiWei WangChen LingXiaolei YinKeqiang FanLuhua LaiMingxiao DingHongkui Deng Suppression of SARS-CoV entry by peptides corresponding to heptad regions on spike glycoprotein. Biochemical and Biophysical Research Communications 2004, 319, 746-752, 10.1016/j.bbrc.2004.05.046.
- Dong P. Han; Adam Penn-Nicholson; Michael W. Cho; Identification of critical determinants on ACE2 for SARS-CoV entry and development of a potent entry inhibitor. Virology 2006, 350, 15-25, 10.1016/j.virol.2006.01.029.
- Genwei Zhang; Sebastian Pomplun; Alexander Robert Loftis; Xuyu Tan; Andrei Loas; Bradley L. Pentelute; Investigation of ACE2 N-terminal fragments binding to SARS-CoV-2 Spike RBD. bioRxiv 2020, none, none, 10.1101/2020.03.19.999318.
- Borong Lin; Xue Qing; Jinling Liao; Kan Zhuo; Role of Protein Glycosylation in Host-Pathogen Interaction. Cells 2020, 9, 1022, 10.3390/cells9041022.
- Anneke Engering; Teunis B. H. Geijtenbeek; Yvette Van Kooyk; Immune escape through C-type lectins on dendritic cells. Trends in Immunology 2002, 23, 480-485, 10.1016/s1471-4906(02)02296-2.
- Alessandra Cambi; Marjolein Koopman; Carl G. Figdor; How C-type lectins detect pathogens. Cellular Microbiology 2005, 7, 481-488, 10.1111/j.1462-5822.2005.00506.x.
- Murat Topuzoğullari; Tayfun Acar; Pelin Pelit Arayıcı; Burcu Uçar; Erennur Ugurel; Emrah Şefik Abamor; Tülin Arasoğlu; Dilek Turgut-Balik; Serap Derman; An insight into the epitope-based peptide vaccine design strategy and studies against COVID-19. TURKISH JOURNAL OF BIOLOGY 2020, 44, 215-227, 10.3906/biy-2006-1.
- Ge Liu; Brandon Carter; Trenton Bricken; Siddhartha Jain; Mathias Viard; Mary Carrington; David K. Gifford; Computationally Optimized SARS-CoV-2 MHC Class I and II Vaccine Formulations Predicted to Target Human Haplotype Distributions. Cell Systems 2020, 11, 131-144.e6, 10.1016/j.cels.2020.06.009.
- Abhishek Singh; Mukesh Thakur; Lalit Kumar Sharma; Kailash Chandra; Designing a multi-epitope peptide based vaccine against SARS-CoV-2. Scientific Reports 2020, 10, 1-12, 10.1038/s41598-020-73371-y.
- Anjali Dhall; Sumeet Patiyal; Neelam Sharma; Salman Sadullah Usmani; Gajendra P.S. Raghava; Computer-aided prediction and design of IL-6 inducing peptides: IL-6 plays a crucial role in COVID-19. Briefings in Bioinformatics 2020, 1, bbaa259, 10.1093/bib/bbaa259.
- Leticia R. Cruz; Idania Baladron; Aliusha Rittoles; Pablo A. Diaz; Carmen Valenzuela; Raul Santana; Maria M. Vazquez; Ariadna Garcia; Deyli Chacon; Delvin Thompson; et al. Treatment with an Anti-CK2 Synthetic Peptide Improves Clinical Response in Covid-19 Patients with Pneumonia. A Randomized and Controlled Clinical Trial. medRxiv 2020, none, none, 10.1101/2020.09.03.20187112.
- Maria Del Carmen Dominguez Horta Sr.; CIGB-258 immunomodulatory peptide: a novel promising treatment for critical and severe COVID-19 patients. medRxiv 2020, none, none, 10.1101/2020.05.27.20110601.
- Andre Gustavo Bonavita; Ac2-26 mimetic peptide of annexin A1 to treat severe COVID-19: A hypothesis. Medical Hypotheses 2020, 145, 110352-110352, 10.1016/j.mehy.2020.110352.
- Vladimir Khavinson; N. S. Linkova; Anastasiia Dyatlova; B. I. Kuznik; Roman Umnov; Peptides: Prospects for Use in the Treatment of COVID-19. Molecules 2020, 25, 4389, 10.3390/molecules25194389.
- Suyash Pant; Meenakshi Singh; V. Ravichandiran; U. S. N. Murty; Hemant Kumar Srivastava; Peptide-like and small-molecule inhibitors against Covid-19. Journal of Biomolecular Structure and Dynamics 2020, none, 1-10, 10.1080/07391102.2020.1757510.
- Linlin Zhang; Daizong Lin; Xinyuanyuan Sun; Ute Curth; Christian Drosten; Lucie Sauerhering; Stephan Becker; Katharina Rox; Rolf Hilgenfeld; Crystal structure of SARS-CoV-2 main protease provides a basis for design of improved α-ketoamide inhibitors. Science 2020, 368, 409-412, 10.1126/science.abb3405.
- Ritika Kabra; Shailza Singh; Evolutionary Artificial Intelligence Based Peptide Discoveries for Effective Covid-19 Therapeutics. Biochimica et Biophysica Acta (BBA) - Molecular Basis of Disease 2020, 1867(1), 165978, 10.1016/j.bbadis.2020.165978.
- Surid Mohammad Chowdhury; Shafi Ahmad Talukder; Akib Mahmud Khan; Nadia Afrin; Ackas Ali; Rajib Islam; Rimon Parves; Abdulla Al Mamun; Abu Sufian; Nayeem Hossain; et al. Antiviral Peptides as Promising Therapeutics against SARS-CoV-2. The Journal of Physical Chemistry B 2020, 124, 9785-9792, 10.1021/acs.jpcb.0c05621.
- Andre Watson; Leonardo M R Ferreira; Peter Hwang; Jinbo Xu; Robert M. Stroud; Peptide Antidotes to SARS-CoV-2 (COVID-19). bioRxiv 2020, none, none, 10.1101/2020.08.06.238915.
- Arun Suria Karnan Mahendran; Yin Sze Lim; Chee-Mun Fang; Hwei-San Loh; Cheng Foh Le; The Potential of Antiviral Peptides as COVID-19 Therapeutics. Frontiers in Pharmacology 2020, 11, 575444, 10.3389/fphar.2020.575444.
- Biplab K. Maiti; Potential Role of Peptide-Based Antiviral Therapy Against SARS-CoV-2 Infection. ACS Pharmacology & Translational Science 2020, 3, 783-785, 10.1021/acsptsci.0c00081.
- Saad Salman; Fahad Hassan Shah; Maham Chaudhry; Muniba Tariq; Muhammad Yasir Akbar; Muhammad Adnan; In silico analysis of protein/peptide-based inhalers against SARS-CoV-2. Future Virology 2020, 15, 557-564, 10.2217/fvl-2020-0119.
- I-Ni Hsieh; Kevan L. Hartshorn; The Role of Antimicrobial Peptides in Influenza Virus Infection and Their Potential as Antiviral and Immunomodulatory Therapy. Pharmaceuticals 2016, 9, 53, 10.3390/ph9030053.
- Bhattacharya, R.; Gupta, A.M.; Mitra, S.; Mandal, S.; Biswas, S.R.; A natural food preservative peptide nisin can interact with the SARS-CoV-2 spike protein receptor human ACE2. Virology 2021, 552, 107–111, 10.1016/j.virol.2020.10.002.
- Sherif Elnagdy; Maha AlKhazindar; The Potential of Antimicrobial Peptides as an Antiviral Therapy against COVID-19. ACS Pharmacology & Translational Science 2020, 3, 780-782, 10.1021/acsptsci.0c00059.
- Jianshe Lang; Ning Yang; Jiejie Deng; Kangtai Liu; Peng Yang; Guigen Zhang; Chengyu Jiang; Inhibition of SARS Pseudovirus Cell Entry by Lactoferrin Binding to Heparan Sulfate Proteoglycans. PLOS ONE 2011, 6, e23710, 10.1371/journal.pone.0023710.
- Ahmed Salah Gouda; Bruno Mégarbane; Snake venom‐derived bradykinin‐potentiating peptides: A promising therapy for COVID ‐19?. Drug Development Research 2020, none, none, 10.1002/ddr.21732.
- Yogesh Kumar Verma; Ranjan Verma; Nishant Tyagi; AmanPreet Behl; Subodh Kumar; G. U. Gurudutta; COVID-19 and its Therapeutics: Special Emphasis on Mesenchymal Stem Cells Based Therapy. Stem Cell Reviews and Reports 2020, none, 1-19, 10.1007/s12015-020-10037-2.
- Giulia Abate; Maurizio Memo; Daniela Uberti; Impact of COVID-19 on Alzheimer’s Disease Risk: Viewpoint for Research Action. Healthcare 2020, 8, 286, 10.3390/healthcare8030286.
- Rasoul Mirzaei; Pedram Goodarzi; Muhammad Asadi; Ayda Soltani; Hussain Ali Abraham Aljanabi; Ali Salimi Jeda; Shirin Dashtbin; Saba Jalalifar; Rokhsareh Mohammadzadeh; Ali Teimoori; et al.Kamran TariMehdi SalariSima GhiasvandSima KazemiRasoul YousefimashoufHossein KeyvaniSajad Karampoor Bacterial co‐infections with SARS‐CoV ‐2. IUBMB Life 2020, 72, 2097-2111, 10.1002/iub.2356.
- Hui Xuan Lim; Jianhua Lim; Seyed Davoud Jazayeri; Sibrandes Poppema; Chit Laa Poh; Development of multi-epitope peptide-based vaccines against SARS-CoV-2. Biomedical Journal 2020, none, none, 10.1016/j.bj.2020.09.005.
- Huy Xuan Luong,Tung Truong Thanh, Tuan Hiep Tran; Antimicrobial peptides – Advances in development of therapeutic applications.. Life Sciences 2020, 260, 118407-118407, http://doi.org/10.1016/j.lfs.2020.118407.
- Huy Xuan Luong,Tung Truong Thanh, Tuan Hiep Tran; Antimicrobial peptides – Advances in development of therapeutic applications.. Life Sciences 2020, 260, 118407-118407, http://doi.org/10.1016/j.lfs.2020.118407.
