Cytokinesis, as the last stage of the cell division cycle, is a tightly controlled process amongst all eukaryotes, with defective division leading to severe cellular consequences and implicated in serious human diseases and conditions such as cancer. Both mammalian cells and the fission yeast Schizosaccharomyces pombe use binary fission to divide into two equally sized daughter cells. Similar to mammalian cells, in S. pombe, cytokinetic division is driven by the assembly of an actomyosin contractile ring (ACR) at the cell equator between the two cell tips. The ACR is composed of a complex network of membrane scaffold proteins, actin filaments, myosin motors and other cytokinesis regulators. The contraction of the ACR leads to the formation of a cleavage furrow which is severed by the endosomal sorting complex required for transport (ESCRT) proteins, leading to the final cell separation during the last stage of cytokinesis, abscission.
- cytokinesis
- contractile ring
- fission yeast
1. The Use of Fission Yeast to Study Eukaryotic Cytokinesis
2. Actin–Myosin Contractile Ring (ACR) Assembly in Fission Yeast
2.1. Positioning of the Cell Division Plane
In S. pombe, cellular growth occurs throughout a longer interphase period, with this ceasing during the shorter mitosis and cytokinesis periods after a certain cell length is achieved. During the cell cycle, the “middle” and “end or tip” locations are specified by two spatial axes. The “middle” location is defined by the nucleus, which is positioned at the cell center by a microtubule-pushing mechanism, where a force is produced by the cytoplasmic microtubule bundles and acts on the nucleus [10,11][10][11]. Furthermore, this force is able to efficiently re-center the nuclei of cells exposed to nuclear displacement [12,13][12][13]. The dynamic interplay between the nucleus and the microtubule cytoskeleton is illustrated and reviewed by Gallardo et al. [14]. The “end or tip” location is defined by a formin-mediated actin assembly mechanism at cell tips [15], and polarity factors including the DYRK kinase Pom1p gradient at cell poles [16]. Pom1p gradients are tightly controlled at the “end or tip”, with the dephosphorylation of Pom1p enhancing a lipid-binding activity, whereas autophosphorylation promotes Pom1p’s detachment from the plasma membrane [16]. Pom1p has an established role in regulating the timing of mitotic entry, as it phosphorylates the membrane-binding C-terminal region of the ACR scaffold protein, Cdr2p, at the cell “middle” through preventing its plasma membrane binding and the formation of nodes [17]. Research has revealed a strong correlation between the cell size at division and Pom1p medial levels; however, such a correlation between the cytosolic or cell tip levels of Pom1p and the cell length is inconsistent, indicating that Pom1p may interact with Cdr2p in the cytosol or at the cell tips [18]. For example, the Pom1p gradient model [16] is opposed by the findings of Pan et al. [19], in which they proposed a novel cell size control mechanism in which cells use Cdr2p to monitor their size. This model therefore implicates a relationship between both Pom1p and Cdr2p with cell size. Much evidence shows that the Anillin-like protein Mid1p localizes to the “middle” location and initiates ACR assembly [20,21,22,23][20][21][22][23]. Mid1p has two membrane binding domains, the pleckstrin homology domain (PH) and the cryptic domain (C2) [24]. However, it only binds the plasma membrane after it is activated and released from the nucleus [20,25][20][25]. The roles of Mid1p in positioning the ACR are now well understood in S. pombe and are reviewed in Rezig et al. [26], with the mechanism of the medial positioning of the ACR schematically described in Figure 1.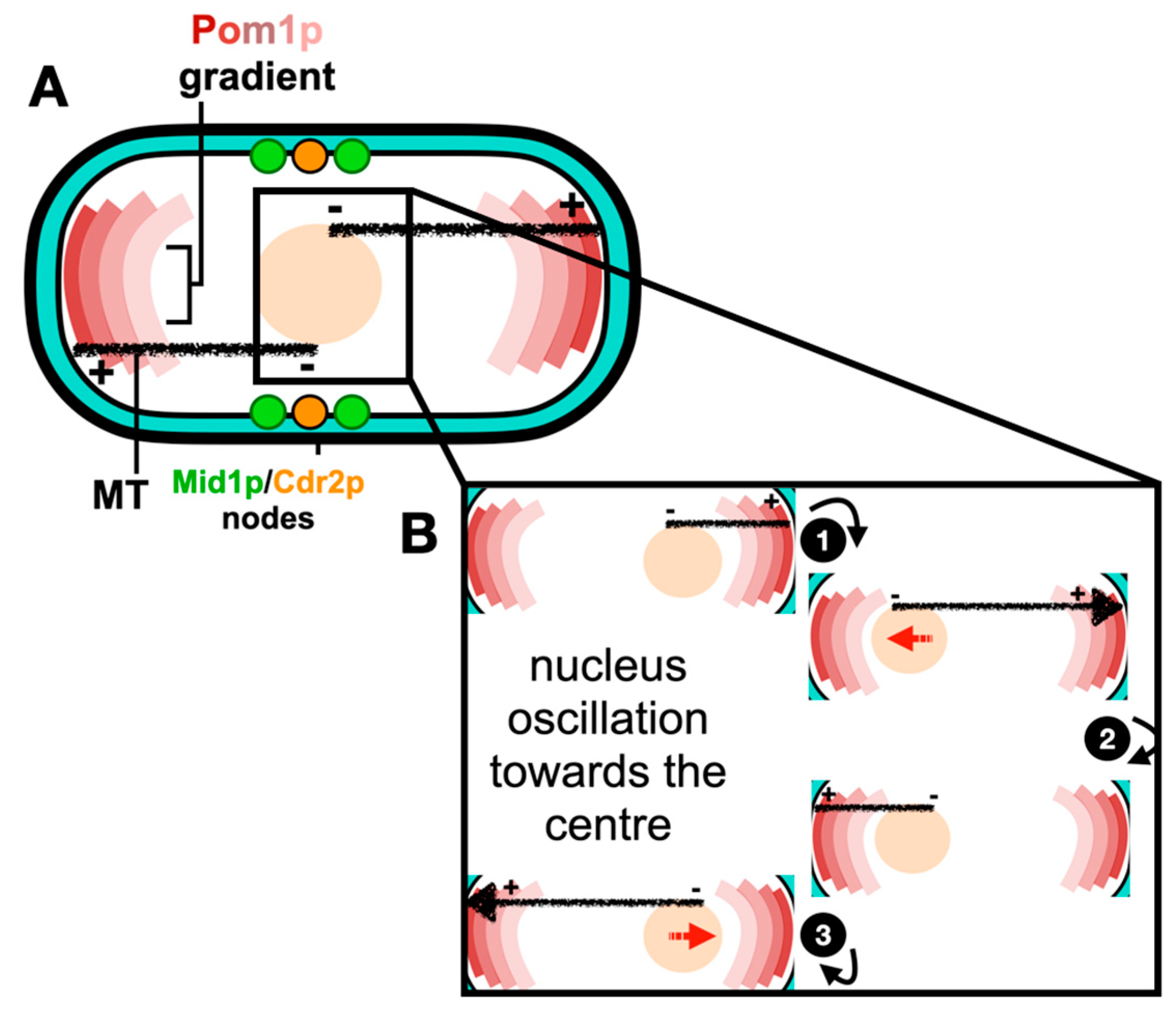
2.2. Molecular Organization of Nodes within the ACR
The current model for ACR assembly includes the formation of two types of interphase nodes: type 1 “stationary” nodes containing Mid1p, Cdr1p and Cdr2p; and type 2 “anchoring” nodes containing Blt1p, Klp8p, Gef2p and Nod1p [27,29][27][29]. Type 2 “anchoring” nodes diffuse into the cell cortex, and at mitotic onset, they are captured by type 1 “stationary” nodes to form cytokinesis nodes. Next, Mid1p recruits the myosin-II, Myo2p, Cdc15p, Rng2p and Cdc12p [30]. The cytokinesis nodes then merge into a ring-like structure, named the actin–myosin contractile ring (ACR) and, as its name implies, it is composed of actin filaments and myosin-II motors in addition to various classes of cytokinesis proteins [31]. Live cell imaging, high-speed fluorescence photo-activation localization microscopy (FPALM), and fluorescence resonance energy transfer (FRET) have been shown to be excellent methods to dissect ACR nodes. Recent findings have revealed that nodes are discrete units with stoichiometric ratios and a specific distribution of constituent proteins [28,30,31,32][28][30][31][32]. Furthermore, the localization of the ACR constituents is thought to be arranged in several layers relative to the plasma membrane, starting with the plasma-membrane-binding proteins and the tail of myosin-II, to the intermediate cytokinesis proteins, and farthest from the plasma membrane lies the myosin motor domains, F-actin and its cross-linkers [32]. Advances in laser scanning microscopy, such as Airyscanning using a very low laser power to acquire high-quality images, have increased the resolution and signal-to-noise ratio and enabled the detection and measurement of even faint individual cytokinesis nodes [33]. The coalescence of nodes leads to ACR assembly through the search, capture, pull and release (SCPR) mechanism, whereby Cdc12p nucleates actin filaments as Myo2p pulls actin filaments, thus producing the force required to pull the individual nodes into the ACR [28,34,35][28][34][35]. Such an assembly of the ACR from node precursors is schematically described in Figure 2.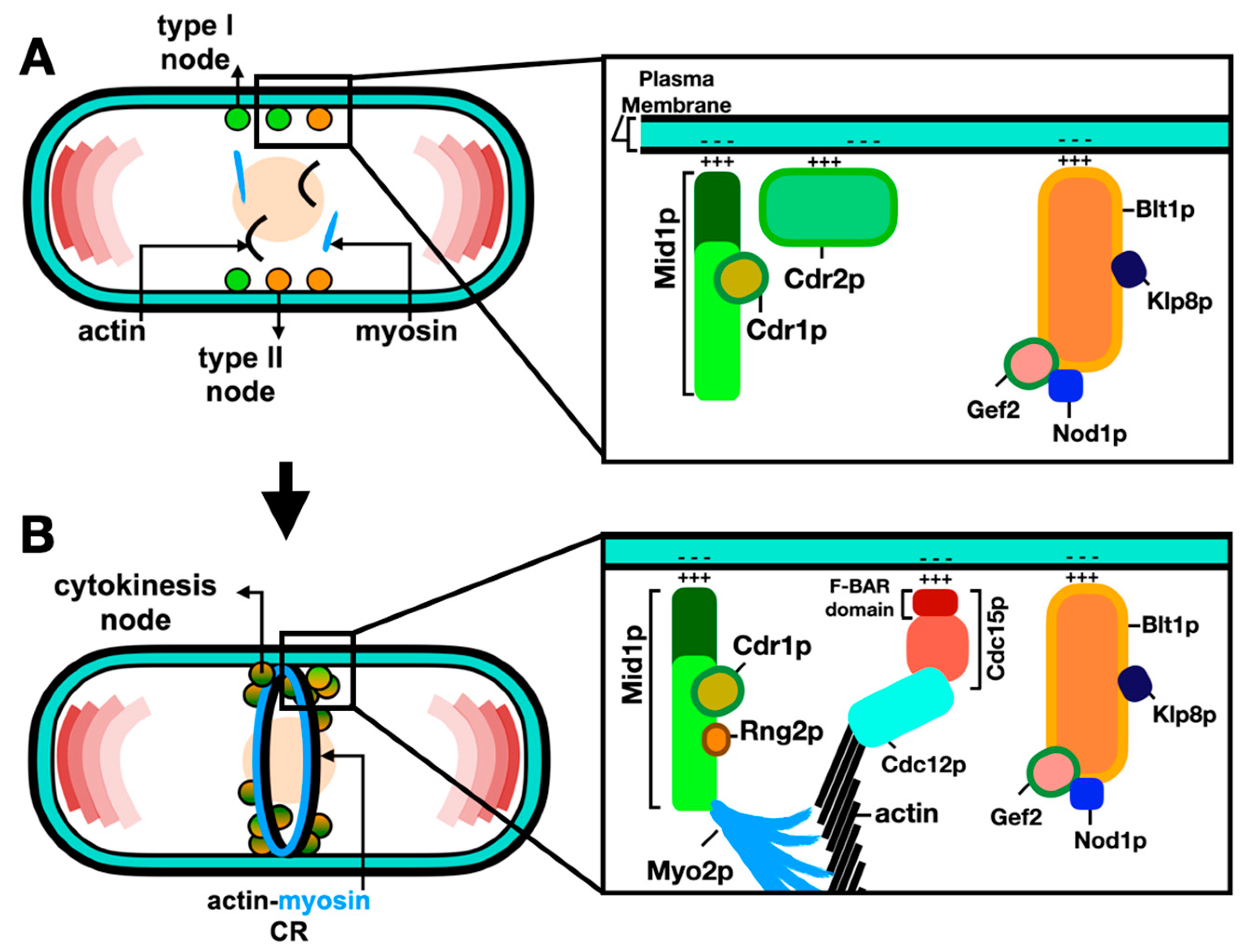
2.3. Anchorage of the ACR to the Plasma Membrane
In the assembled ACR, the Myo2p tails and Cdc15p anchor to the plasma membrane, with the Myo2p heads, Myp2p and the bundle of actin filaments localizing 60 nm away from the plasma membrane [36]. It is suggested that this organization connects the bundle of actin filaments to the plasma membrane [37]. Cdc15p next recruits Cdc12p to the ACR, and this interaction is thought to be essential for ACR organization and stability [38]. The phospho-status of Cdc15p influences its ability to bind the plasma membrane, with the phosphorylation of Cdc15p by Pom1p inhibiting its binding to the plasma membrane at the cell tips [39]. Additionally, the p21-activated protein kinase (Pak1p), another polarity kinase, was found to regulate the function of Mid1p and Cdc15p [40]. Cdc15p has three regulatory components: an N-terminal Fre/Cip4 homology Bin/Amphiphysin/Rvs domain (F-BAR), a medial intrinsically disordered region (IDR) and a C-terminal Src homology 3 domain (SH3). While the F-BAR domain enables protein oligomerization and concentration on the plasma membrane to scaffold protein assemblies resulting in membrane deformation [41], it was recently found that the phosphorylation of Cdc15p induces the separation of the Cdc15p IDR region resulting in an inhibition of Cdc15p phase separation, and the formation of condensate on the plasma membrane [42]. Moshtohry et al. [43] recently used laser ablation, a technique based on photodamage in which cellular structures could be degraded using a focused pulsed laser, to investigate the mechanical role of Cdc15p during S. pombe cytokinesis and found that the ACR recoils after being severed. However, this recoil profile was greater and slower in the ablated ACR of Cdc15-depleted cells, suggesting that the loss of Cdc15p decreases the stiffness of the ACR material. Furthermore, another F-BAR protein, Imp2p, was found to contribute to the stiffness of the ACR [37].References
- Pollard, T.D.; Wu, J.-Q. Understanding cytokinesis: Lessons from fission yeast. Nat. Rev. Mol. Cell Biol. 2010, 11, 149–155.
- Hartwell, L.H.; Culotti, J.; Reid, B. Genetic Control of the Cell-Division Cycle in Yeast, I. Detection of Mutants. Proc. Natl. Acad. Sci. USA 1970, 66, 352–359.
- Hartwell, L.H.; Mortimer, R.K.; Culotti, J.; Culotti, M. Genetic control of the cell division cycle in yeast: V. genetic analysis of cdc mutants. Genetics 1973, 74, 267–286.
- Nurse, P. Genetic control of cell size at cell division in yeast. Nature 1975, 256, 547–551.
- Nurse, P.; Thuriaux, P.; Nasmyth, K. Genetic control of the cell division cycle in the fission yeast Schizosaccharomyces pombe. Mol. Gen. Genet. 1976, 146, 167–178.
- Chang, F.; Nurse, P. Finishing the cell cycle: Control of mitosis and cytokinesis in fission yeast. Trends Genet. 1993, 9, 333–335.
- Chang, F.; Nurse, P. How fission yeast fission in the middle. Cell 1996, 84, 191–194.
- Nurse, P. Fission yeast cell cycle mutants and the logic of eukaryotic cell cycle control. Mol. Biol. Cell 2020, 31, 2871–2873.
- Chang, F. Forces that shape fission yeast cells. Mol. Biol. Cell 2017, 28, 1819–1824.
- Piel, M.; Tran, P.T. Cell shape and cell division in fission yeast. Curr. Biol. 2009, 19, R823–R827.
- Tran, P.T.; Marsh, L.; Doye, V.; Inoue, S.; Chang, F. A mechanism for nuclear positioning in fission yeast based upon microtubule pushing. J. Cell Biol. 2001, 153, 397–411.
- Daga, R.R.; Chang, F. Dynamic positioning of the fission yeast cell division plane. Proc. Natl. Acad. Sci. USA 2005, 102, 8228–8232.
- Daga, R.R.; Yonetani, A.; Chang, F. Asymmetric Microtubule Pushing Forces in Nuclear Centering. Curr. Biol. 2006, 16, 1544–1550.
- Gallardo, P.; Barrales, R.R.; Daga, R.R.; Salas-Pino, S. Nuclear Mechanics in the Fission Yeast. Cells 2019, 8, 1285.
- Martin, S.G.; McDonald, W.H.; Yates, J.R., 3rd; Chang, F. Tea4p links microtubule plus ends with the formin for3p in the establishment of cell polarity. Dev. Cell. 2005, 8, 479–491.
- Hersch, M.; Hachet, O.; Dalessi, S.; Ullal, P.; Bhatia, P.; Bergmann, S.; Martin, S.G. Pom1 gradient buffering through intermolecular auto-phosphorylation. Mol. Syst. Biol. 2015, 11, 818.
- Moseley, J.B.; Mayeux, A.; Paoletti, A.; Nurse, P. A spatial gradient coordinates cell size and mitotic entry in fission yeast. Nature. 2009, 7248, 857–860.
- Gerganova, V.; Floderer, C.; Archetti, A.; Michon, L.; Carlini, L.; Reichler, T.; Manley, S.; Martin, S.G. Author response: Multi-phosphorylation reaction and clustering tune Pom1 gradient mid-cell levels according to cell size. Elife 2019, 3, e45983.
- Pan, K.Z.; Saunders, T.E.; Flor-Parra, I.; Howard, M.; Chang, F. Cortical regulation of cell size by a sizer cdr2p. Elife 2014, 3, e02040.
- Almonacid, M.; Celton-Morizur, S.; Jakubowski, J.L.; Dingli, F.; Loew, D.; Mayeux, A.; Chen, J.-S.; Gould, K.L.; Clifford, D.M.; Paoletti, A. Temporal control of contractile ring assembly by Plo1 regulation of Myosin II recruitment by Mid1/Anillin. Curr. Biol. 2011, 21, 473–479.
- Paoletti, A.; Chang, F. Analysis of mid1p, a protein required for placement of the cell division site, reveals a link between the nucleus and the cell surface in fission Yeast. Mol. Biol. Cell 2000, 11, 2757–2773.
- Rezig, I.M.; Yaduma, W.G.; Gould, G.W.; McInerny, C.J. Anillin/Mid1p interacts with the ESCRT-associated protein Vps4p and mitotic kinases to regulate cytokinesis in fission yeast. Cell Cycle 2021, 20, 1845–1860.
- Saha, S.; Pollard, T.D.; Akamatsu, M.; Lin, Y.; Bewersdorf, J.; Lidke, M.E.D.; Sherlekar, A.; Rikhy, R.; Montell, M.E.D.; Goss, J.W.; et al. Anillin-related protein Mid1p coordinates the assembly of the cytokinetic contractile ring in fission yeast. Mol. Biol. Cell 2012, 23, 3982–3992.
- Chatterjee, M.; Pollard, T.D. The functionally important N-terminal half of fission yeast mid1p anillin is intrinsically disordered and undergoes phase separation. Biochemistry 2019, 58, 3031–3041.
- Steever, A.; Pringle, J.; Bähler, J.; Wang, Y.-L.; Gould, K.; McCollum, D.; Wheatley, S. Role of polo kinase and mid1p in determining the site of cell division in fission yeast. J. Cell Biol. 1998, 143, 1603–1616.
- Rezig, I.M.; Yaduma, W.G.; Gould, G.W.; McInerny, C.J. The role of anillin/Mid1p during medial division and cytokinesis: From fission yeast to cancer cells. Cell Cycle 2022, 22, 633–644.
- Akamatsu, M.; Berro, J.; Pu, K.-M.; Tebbs, I.R.; Pollard, T.D. Cytokinetic nodes in fission yeast arise from two distinct types of nodes that merge during interphase. J. Cell Biol. 2014, 204, 977–988.
- Laplante, C.; Huang, F.; Tebbs, I.R.; Bewersdorf, J.; Pollard, T.D. Molecular organization of cytokinesis nodes and contractile rings by super-resolution fluorescence microscopy of live fission yeast. Proc. Natl. Acad. Sci. USA 2016, 113, E5876–E5885.
- Zhu, Y.-H.; Ye, Y.; Wu, Z.; Wu, J.-Q. Cooperation between Rho-GEF Gef2 and its binding partner Nod1 in the regulation of fission yeast cytokinesis. Mol. Biol. Cell 2013, 24, 3187–3204.
- Akamatsu, M.; Lin, Y.; Bewersdorf, J.; Pollard, T.D.; Saha, S.; Wang, M.E.Y.-L.; Wills, R.C.; Goulden, B.D.; Hammond, G.R.V.; Kozminski, M.E.K.G.; et al. Analysis of interphase node proteins in fission yeast by quantitative and superresolution fluorescence microscopy. Mol. Biol. Cell 2017, 28, 3203–3214.
- Malla, M.; Pollard, T.D.; Chen, Q. Counting actin in contractile rings reveals novel contributions of cofilin and type II myosins to fission yeast cytokinesis. Mol. Biol. Cell 2022, 33, ar51.
- McDonald, N.A.; Lind, A.L.; Smith, S.E.; Li, R.; Gould, K.L. Nanoscale architecture of the Schizosaccharomyces pombe contractile ring. Elife 2017, 15, e28865.
- Sayyad, W.A.; Pollard, T.D. The number of cytokinesis nodes in mitotic fission yeast scales with cell size. Elife 2022, 12, e76249.
- Vavylonis, D.; Wu, J.-Q.; Hao, S.; O’Shaughnessy, B.; Pollard, T.D. Assembly mechanism of the contractile ring for cytokinesis by fission yeast. Science 2008, 319, 97–100.
- Zimmermann, D.; Homa, K.E.; Hocky, G.M.; Pollard, L.W.; De La Cruz, E.M.; Voth, G.A.; Trybus, K.M.; Kovar, D.R. Mechanoregulated inhibition of formin facilitates contractile actomyosin ring assembly. Nat. Commun. 2017, 8, 703.
- Swulius, M.T.; Nguyen, L.T.; Ladinsky, M.S.; Ortega, D.R.; Aich, S.; Mishra, M.; Jensen, G.J. Structure of the fission yeast actomyosin ring during constriction. Proc. Natl. Acad. Sci. USA 2018, 115, E1455–E1464.
- Bellingham-Johnstun, K.; Anders, E.C.; Ravi, J.; Bruinsma, C.; Laplante, C. Molecular organization of cytokinesis node predicts the constriction rate of the contractile ring. J. Cell Biol. 2021, 220, e202008032.
- Snider, C.E.; Chandra, M.; McDonald, N.A.; Willet, A.H.; Collier, S.E.; Ohi, M.D.; Jackson, L.P.; Gould, K.L. Opposite surfaces of the Cdc15 F-BAR domain create a membrane platform that coordinates cytoskeletal and signaling components for cytokinesis. Cell Rep. 2020, 33, 108526.
- Bhattacharjee, R.; Mangione, M.C.; Wos, M.; Chen, J.-S.; Snider, C.E.; Roberts-Galbraith, R.H.; McDonald, N.A.; Presti, L.L.; Martin, S.G.; Gould, K.L. DYRK kinase Pom1 drives F-BAR protein Cdc15 from the membrane to promote medial division. Mol. Biol. Cell 2020, 31, 917–929.
- Magliozzi, J.O.; Sears, J.; Cressey, L.; Brady, M.; Opalko, H.E.; Kettenbach, A.N.; Moseley, J.B. Fission yeast Pak1 phosphorylates anillin-like Mid1 for spatial control of cytokinesis. J. Cell Biol. 2020, 219, e201908017.
- Snider, C.E.; Noor, W.N.I.W.M.; Nguyen, N.T.H.; Gould, K.L.; Suetsugu, S. The state of F-BAR domains as membrane-bound oligomeric platforms. Trends Cell Biol. 2021, 31, 644–655.
- Bhattacharjee, R.; Hall, A.R.; Mangione, M.C.; Igarashi, M.G.; Roberts-Galbraith, R.H.; Chen, J.-S.; Vavylonis, D.; Gould, K.L. Author response: Multiple polarity kinases inhibit phase separation of F-BAR protein Cdc15 and antagonize cytokinetic ring assembly in fission yeast. Elife 2023, 7, e83062.
- Moshtohry, M.; Bellingham-Johnstun, K.; Elting, M.W.; Laplante, C. Laser ablation reveals the impact of Cdc15p on the stiffness of the contractile ring. Mol. Biol. Cell 2022, 33, br9.
