One of the most consumed foods is milk and milk products, and guaranteeing the suitability of these products is one of the major concerns in our society. This has led to the development of numerous sensors to enhance quality controls in the food chain. However, this is not a simple task, because it is necessary to establish the parameters to be analyzed and often, not only one compound is responsible for food contamination or degradation. To attempt to address this problem, a multiplex analysis together with a non-directed (e.g., general parameters such as pH) analysis are the most relevant alternatives to identifying the safety of dairy food. In recent years, the use of new technologies in the development of devices/platforms with optical or electrochemical signals has accelerated and intensified the pursuit of systems that provide a simple, rapid, cost-effective, and/or multiparametric response to the presence of contaminants, markers of various diseases, and/or indicators of safety levels.
- multiplex detection
- food contaminants
- shelf life extension
- safety control
1. Introduction
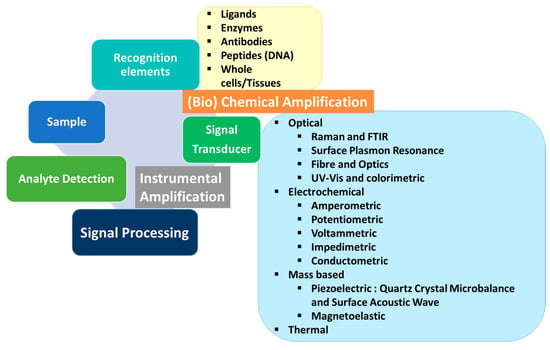
2. Optical Sensors
As for the use of optical multiplex sensors in the dairy industry, the uttermost efforts are directed towards the control and detection of microbial presence, either as microorganisms themselves or as their toxins. This is especially relevant in the detection of Staphylococcus aureus [4,5,6,7][1][2][3][4] and Salmonella typhimurium [5,7[2][4][5],8], although other microorganisms such as Escherichia coli [4,8][1][5] and Vibrio parahaemolyticus [5][2] are also studied, as well as bacterial toxins [9][6]. Furthermore, the second most relevant analytes in dairy products analyzed with multiplex optical sensors are antibiotics [10,11,12,13,14,15,16][7][8][9][10][11][12][13] used in cattle to prevent bacterial infections such as mastitis.2.1. Surface-Enhanced Raman Spectroscopy
A common approach in multiplex optical sensing for this application is the use of lateral flow systems, using surface-enhanced Raman spectroscopy (SERS), typically supported with gold nanoparticles, but also with fluorescence or even visual detection. In fact, nanoparticle-based SERS immunoprobes can be prepared easily with simple chemical reactions. As an example, Shi et al. [10,15][7][12] obtained gold nanoparticles 40 nm in diameter with the classical reduction method of HAuCl4 using trisodium citrate as the reducer. Then, these nanoparticles are labeled with a Raman reporter molecule, typically para-amino thiophenol or 5,5′-dithiobis (2-nitrobenzoic acid), by a direct reaction, and the final SERS probe is derivatized with specific antibodies by direct adsorption. BSA is used to block unspecific binding sites. A more complex SERS system, which is not based on lateral flow assays (LFAs), is proposed by Lu et al., for detecting melamine and dicyandiamide in dairy products [17][14], toxic compounds which are sometimes fraudulently added to dairy products to forge a high protein (nitrogen) content. In this case, they propose the use of a 3D hybrid SERS substrate utilizing polystyrene (PS) microspheres as the template matrix, silver as the active interlayer, and graphene oxide (GO) as the coating surface. Melamine is detected using its Raman peak at 685 cm−1, reaching a limit of detection of 2.8 × 10−10 M, while dicyandiamide is detected through its Raman absorption at 921 cm−1, with a limit of detection of 6.0 × 10−9 M. Using SERS methods, Zhang et al. [18][15] modified gold NPs for the simultaneous detection of Salmonella typhimurium and Staphylococcus aureus. This sensor has a high sensitivity for both pathogens, with a Calvaria limit of detection of 35 cfu mL−1 for S. aureus and 15 cfu mL−1 for S. typhimurium. A limit of detection of 35 cfu/mL, selectivity, and 3 h analysis time are the most remarkable characteristics of this sensor [18][15]. Multiplex SERS has also been used successfully to detect mycotoxins rather than microorganisms themselves. In line with this, Zhang et al. [19][16] proposed a multiplex sensor to quantify six different target mycotoxins: AFB1, zearalenone, fumonisin B1, deoxynivalenol, ochratoxin, and T-2 toxin. After synthesis and characterization, Au@AgNPs were used to prepare a SERS nanoprobe, and coupling these with the anti-mycotoxin antibody of each studied mycotoxin, 1332 and 1589 cm−1 peaks were selected as markers of mycotoxin presence. The results showed that the SERS intensity of the peaks at 1332 and 1589 cm−1 decreased in the presence of mycotoxins.2.2. Lateral Flow Assays (LFAs)
As stated before, many of the multiplex sensors for microorganisms or antibiotics are based on LFAs (Figure 3). Typically, these strips consist of four sections: a sample pad where the sample is poured and a conjugate pad containing antibodies against the analytes, labeled with an optically detectable mark (e.g., SERS probe or fluorescent tag). When the sample flows because of capillarity and reaches this zone, eventual antigens react with tagged antibodies and continue flowing as tagged antibody–antigen complexes. Then, there is a nitrocellulose membrane with immobilized anti-analyte antibodies which react with this labeled flowing complex, revealing a line. A second control line made of anti-IgG antibodies is usually incorporated, which will react with the labeled antibodies either if they reacted with their corresponding targets or not [9,20][6][17]. A similar although slightly different approach was used by Shi et al. in their multiplex sensor for neomycin and lincomycin in milk [10][7]. In this case, the test line was not prepared using anti-neomycin and anti-lincomycin antibodies but by immobilizing lincomycin and neomycin instead. In this way, if the sample does not contain the antibiotics, the antibodies present in the conjugate pad reach the test line and react with it, therefore creating a visible line. On the contrary, if the sample contains the antibiotics, they react with the antibodies so that they are not able to interact with the immobilized antibiotics on the test line, resulting in no visible line. In this case, the amount of antibiotics is inversely proportional to the intensity of the revealed test line.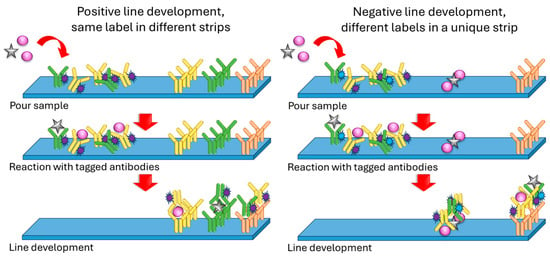
2.3. Immunoassays with Optical Detection
Other approaches based on optical detection have also been exploited, in addition to lateral flow assays and immunoassays. As an example, Juronen et al. propose the use of bead injection analysis (BIA) together with the attachment of microorganisms onto microcolumns and the use of fluorescence-labeled antibodies [4][1]. The authors preconcentrated microorganisms into microcolumns, taking advantage of the affinity of Escherichia coli and Staphylococcus aureus for the human Fc antibody fragment. Therefore, Fc immobilized in Sephadex beads is used to concentrate microorganisms from milk samples, detected with a secondary antibody. The multiplex is possible by using different non-overlapping emitting fluorophores as labels in the secondary antibodies. A multiplex using fluorescence was also employed by Duan et al. to detect in a single analysis the presence of V. parahaemolyticus, S. aureus, and S. typhimurium [5][2]. In their case, detection relied on the Förster resonance energy transfer (FRET) phenomenon, using three different fluorescence tags as donors and carbon dots as acceptors. The authors prepared a different aptamer against each bacterium. Every aptamer was, likewise, tagged with a fluorescent molecule emitting at different wavelengths (colors). The labeled aptamers were brought into contact with carbon dots, where they got adsorbed via π–π stacking forces. The distance between the fluorescence label in the aptamer (donor) and the carbon dot (acceptor) was short enough to produce the FRET phenomenon and, therefore, fluorescence was quenched. When the proper bacterium was present, the aptamer bound to it, hence being released from the nanoparticle surface and ending the FRET quenching. Depending on the color of the fluorescence emission, the type of bacterium could be identified. Nevertheless, the use of matrices of immobilized antigens/antibodies and detection with a labeled secondary antibody is a quite frequent approach, with the detection technique depending on the label itself. Particularly interesting are those detections that require non-expensive or sophisticated instruments, such as a smartphone [13][10]. A similar system was used by Huang et al. for S. aureus enterotoxins [6][3]. In this case, the fluorescence donor was lanthanide-doped fluorescence nanoparticles (KGdF4:Ln3+) whereas the acceptor was graphene oxide. KGdF4:Ln3+ was activated with glutaraldehyde and decorated with avidin, which was then used to attach the biotinylated aptamer. The foundation of the sensor is the quenching of the probe fluorescence by the graphene oxide when the former is not bound to its analyte.2.4. Chemiluminescence
Chemiluminescence also produces good results in multiplex analysis. The use of horseradish peroxidase (HRP) and alkaline phosphatase (ALP) as labels with chemiluminescent substrates allowed the determination of 20 fluoroquinolones, 15 β-lactams, 15 sulfonamides, and chloramphenicol in milk [21,22][18][19]. Basically, the methodology of its use consists of a two-step procedure. The well surface is coated with norfloxacin–ovalbumin (for the recognition of quinolones), a penicillin binding protein (for the recognition of penicillins), 4-(4-aminophenylsulfonamido) benzoic acid–ovalbumin (for the recognition of sulfamides), and anti-chloramphenicol polyclonal antibodies (for the recognition of chloramphenicol). The well is then incubated with the sample and HRP-labeled ampicillin and single-chain variable fragment–alkaline phosphatase fusion protein, which binds to quinolones. A competitive procedure takes place, so the higher the concentration of β-lactams and quinolones, the lower the chemiluminescent signal coming from the HRP or ALP, respectively. In a second step, the sample is incubated with ALP-labeled anti-immunoglobulin, HRP-labeled chloramphenicol, and anti-sulfamethoxazole polyclonal antibodies, following again a competitive principle: the higher the amount of chloramphenicol or sulfamides, the lower the chemiluminescent signal rising from the HRP or ALP, respectively. Later on, the same authors propose a slight modification of the procedure using fluorescent quantum dots with different emission wavelengths to label anti-sulfonamides and anti-chloramphenicol antibiotics. Thus, penicillins and quinolones are detected chemiluminescently whereas chloramphenicol and sulfonamides are detected fluorescently [23][20].2.5. Label-Free Assays
Notwithstanding fluorescent or chemiluminescent labels, they are not mandatory in the development of multiplex optical detection systems for dairy products. Surface plasmon resonance imaging (SPRI), as well as light scattering, have been successfully exploited in the detection of antibiotic residues in milk and in the screening of Bacillus colonies. Although the proposed bacterial rapid detection using optical scattering technology (BARDOT) instrument for screening Bacillus colonies is barely a multiplex system, it confidently screens Bacillus from non-Bacillus microorganisms. This discrimination is based on the scattering pattern produced by the microorganism’s colony [24][21]. In the case of SPRI, different antibiotics were spotted on the chip surface in a known pattern. Then, anti-antibiotics and sample were added, following a competitive approach. Changes in spot mass deriving from antibody binding were detected by the system and that was related to the antibiotic content of the sample [25][22]. A similar lab-on-a-chip approach, but based on wavelength-interrogated optical sensors (WIOSs) was developed by Suarez et al. for detecting antibiotics in milk [26][23]. In these systems, the detection principle relies on grating waveguide resonant coupling which, briefly, produces a shift in a laser wavelength whose magnitude is proportional to the quantity of bio-analyte bound to the antibody-modified waveguide. Multiplexing, in Suarez et al.’s case for sulfonamides, fluoroquinolones, and tetracycline is achieved by coating different regions of the chip with the corresponding sensing element. Be that as it may, other detection approaches can be used in these label-free chip-based devices. As an example, Angelopoulou et al. used broadband Mach–Zehnder interferometers combined with an advanced microfluidic module to simultaneously detect bovine k-casein, peanut protein, soy protein, and gliadin in samples from a cleaning-in-place system of a dairy industry [27][24]. Microfluidics and origami paper-based sensing systems may also be used in cattle-related industries other than dairy ones, as proposed by Yang et al., to detect the presence of bovine herpes simplex virus and Brucella and Leptospira bacteria in bovine semen samples [28][25]. A paper-based analytical system was also used by Prasad et al. for multiple titrations in food and, particularly, to analyze lactose in dairy-based fruit drinks. The analysis was based on the addition of lactase, which decomposes lactose in glucose, which is further processed with glucose–oxidase-generating H2O2. This chemical is finally converted into water with the use of horseradish peroxidase, while potassium iodide is converted into iodine, thus developing a brown color related to the initial lactose [29][26].2.6. Dairy and Allergens
The presence and detection of allergens in food is an important public health issue, which is responsible for the morbidity of 105 adults worldwide [30][27]. Focusing on milk and dairy products, the presence of allergens may be considered from two different perspectives. On the one hand, the presence of allergens in milk and dairy products must be taken into account but, on the other hand, some of the milk components may constitute an important health problem too, when dealing with people with lactose intolerance or allergy to β-lactoglobulin. Thus, methods for casein, β-lactoglobulin, and lactose analysis have become a subject of research. Different sensing strategies have been exploited to face this problem. There are commercially available kits for detecting milk allergens, which are mainly based on ELISA and, therefore, time-consuming and require a specialized person to be operated. For a list of them, the readers can check Table 3 in Ashley’s work [31][28]. There are also some commercial multiplex systems used for allergy diagnosis such as ALEX2 or ISAC [32[29][30][31][32],33,34,35], but they are oriented to serum or blood analysis rather than food, since these commercial tests seek to identify the IgE present in blood and they are, therefore, not adaptable for detecting allergens in food. The interferometric reflectance imaging sensor developed by Monroe et al. [36][33] works as a multiplex system to detect allergenic proteins, including β-lactoglobulin, in blood and plasma as well, but it detects the antigenic protein rather than the IgE, thus making the system adaptable for food matrices. Setting the focus on detecting the presence of milk allergens in food, there are some classic lateral flow multiplex devices available for the simultaneous detection of β-lactoglobulin and casein in food [37,38][34][35]. An interesting example of a multiplexed sensor for milk allergens is the microimmunoassay on a DVD published by Badran et al. which, briefly, consisted of immobilizing the allergen proteins (gliadin, casein, β-lactoglobulin, and ovalbumin) on a DVD surface and then performing a competitive microimmunoassay with gold-labeled antibodies and the sample. Special software was used to read the DVD surface and obtain a valid analytical signal [39][36]. Based on plasmon resonance images, Raz et al. [40][37] developed a sensor to identify 13 allergens in cookies and chocolate. Required selectivity was reached by spotting antibodies in a 4 × 7 array against each allergen. Macadamia, hazelnut, nut, almond, pistachio, egg, soy, or casein were some of the detected allergens in this multiplexed proposal. LODs were ranged between 0.2 mg kg−1 for eggs in the cookies and 4.6 mg kg−1 for hazelnut in the dark chocolate. Regarding the detection of allergens in milk and dairy, the same DVD approach was used by Tortajada-Genaro et al. to create a multiplex sensor for hazelnut, peanut, and soybean allergens based on detecting the presence of a specific genetic sequence in different food stuff, including milk [41][38]. The direct detection of antigens is a more common approach, as in the enzyme immunoassay by Blais et al., consisting of the immobilization of anti-antigen antibodies on a strip of polyester cloth, which is brought into contact with the sample. A biotinylated antibody and a further incubation with streptavidin–peroxidase allowed for optic detection by developing a blue color after reaction with tetramethylbenzidine [42][39]. Table 1 summarizes multiplex methodologies for analyzing allergens in milk/dairy or milk/allergens in food stuff.| Sample | Detection | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Food | Allergens | Technique | Sensing System | Total Assay Time | LOD | Ref. | |||||||
| Multiplex detection of allergens in milk, milk-containing products, and dairy products | |||||||||||||
| Milk, (*) cookies, ice cream | Gliadin, Ara h 1, Cor a1 (hazelnut), casein, ovalbumin | Amperometry | Magnet/SPE antibody-tagged immunomagnetic beads | <10 min | Ranging from 0.003 to 0.170 mg/kg | [43][40] | |||||||
| Powdered milk, (*) cookies, sponge cake | Hazelnut, peanut, soybean | Optical (laser) | Digoxin-labeled PCR products detected by hybridization on modified DVD surface | 5 h 20 min | 1 μg/g | [37][34] | |||||||
| MoniQA milk, NIST SRM 1549a milk, (*) Nutella hazelnut spread, 2% milk | Ana o 3 (cashew), Ara h 3/Ara h 6 (peanut), Cor a 9 (hazelnut), Gal d 1/Gal d 2 (egg), Gly m 5 (soy), Bos d 5 (milk), tropomyosin (shrimp) | Fluorescent multiplex array | Monoclonal or polyclonal antibodies covalently coupled to Luminex xMAP® system | 30 min | Ranging from 0.02 to 1.95 ng/mL | [44][41] | |||||||
| (*) Cookies | Casein, soy protein, gluten | Flow cytometry | Fluorescent microsphere-based immunoassay | 1 h 10 min | 0.4 ppm | [45][42] | |||||||
| (*) Oatmeal cookies, milk chocolate, chocolate ice cream | Hazelnut, Brazil nut, peanut | Colorimetry | Enzyme immunoassay system with chromogenic substrate | 4 h 10 min | Ranging from 0.1 to 1.0 μg/g | [38][35] | |||||||
| (*) Cookies | Hazelnut, peanut | Colorimetry | Lab-on-chip/Carbon dot label for lateral flow immunoassay | 15 min | 0.1 ppm | [46][43] | |||||||
| Multiplex detection of milk allergens in food stuff | |||||||||||||
| 53 | ] | Allergen-free probiotics | Gliadin, β-lactoglobulin, hazelnut, almond, peanut, soy | Colorimetry | DVD functionalized with the capture bioreceptors in microarray format | 20 samples in 70 min | Ranging from 0.1 to 143.4 ng/mL | [47][44 | |||||
| Pesticide | Malathion (MAL), chlorpyrifos (CLO) |
GCE/Au nanoparticles, cDNA/Ce(III)–Ce(V)–MOF | ] | ||||||||||
| SWV | 1 | −1 | pM–1 μM | 0.045 pM, | 0.038 pM |
[57][54] | A panel of 38 food commodities based on AOAC recommendations and milk from six different animal sources | Casein, β-lactoglobulin | Colorimetry/Visual | Antibodies coupled to red and blue Carboxyl-dyed, antibody-modified latex beads | 10 min | 0.5 ppm β-lactoglobulin, 2 ppm for caseins | [34][31] |
| Infant jar food, apple juice | Gliadin, casein, β-lactoglobulin, ovalbumin | Optical (laser) | Immunoassay developed on DVD surface | 1 h 25 min | 31 μg/L (casein), 120 μg/L (β-lactoglobulin) | [36][33] | |||||||
3. Electrochemical Devices/Platforms
In recent years, the use of new technologies in the development of devices/platforms with electrochemical transduction has accelerated and intensified the pursuit of systems that provide a simple, rapid, cost-effective, and multiparametric response to the presence of contaminants, markers of various diseases, and/or indicators of safety levels in milk and its derivatives. However, achieving the simultaneous determination of two or more analytes in situ, in a single measurement, and in real time, using only one working electrode, a ‘real sensor’, remains one of the most daunting challenges, primarily due to the intricate nature of the sample matrix. To address these requirements, different approaches have been explored, predominantly employing aptamers and antibodies as specific recognition elements. Electrochemical aptasensors combine the high sensitivity and versatility of electrochemical detection systems on the one hand, and the specificity of aptamers on the other, making them highly capable and effective probes [48,49][45][46] which undoubtedly offer a wide range of possibilities for the simultaneous detection of multiple analytes. They constitute a rapidly developing area of research, particularly in the development of labels that allow for a one-to-one relationship with the analyte and signal amplification. In this regard, one of the areas that has seen intensified research is the field of new materials, especially those at the nanoscale. The use of nanomaterials as multiple labels represents a recent approach, with notable examples including metallic quantum dots, metal–organic frameworks (MOFs), and silica particles, which exhibit high porosity, providing them with a large specific surface area and a high density of molecular loading. These are complemented by more traditional labels such as redox and enzymatic labels. Despite intensive research efforts in the development of such multisensing platforms in recent years, their application, particularly in dairy samples, is scarce and limited to a maximum of three analytes.3.1. Multiplex Electrochemical Platform for Antibiotics
The first platform for the multiplex determination of antibiotic residues, sulfapyridine (SPY) and tetracycline (TC), in spiked and certified milk samples, was reported in 2013 by Conzuelo et al. [50][47]. The platform features a dual-modified screen-printed carbon electrode (SPCE) with a protein G modification, which enhances sensitivity. The SPCE is specifically designed with two distinct regions, each immobilized with capture antibodies for SPY or TC. This dual modification allows for the simultaneous detection of both analytes in a single measurement. The immunoassay is conducted using horseradish peroxidase (HRP)-labeled tracers, and the analytical signal is recorded following the addition of H2O2 in the presence of hydroquinone as a mediator. The platform achieves good recoveries within a 30-min analysis time, enabling the detection of trace amounts of SPY and TC residues below the regulatory levels set by the EU and FDA. The first studies based on the use of aptamers for the multiplex electrochemical analysis of dairy samples appeared in the literature in 2016. These studies present innovative technological platforms, but they cannot be formally considered as aptasensors, particularly due to the need for sample pretreatments or because the analysis requires multiple non-integrated stages with long durations (e.g., hybridizations, digestions). Xue et al. [51][48] developed a platform that enabled the simultaneous detection of streptomycin (STR), chloramphenicol (CHL), and TC residues in milk samples, based on the use of specific aptamers for each antibiotic and quantum dots (QDs) as labels. Complementary DNA (cDNA) sequences were designed for each antibiotic, targeting not only the corresponding aptamers but also part of the respective capture DNA (Cap-DNA) and oligonucleotides labeled with different types of QDs. The process started with hybridization between the cDNA and aptamers to form DNA duplexes. However, in the presence of STR, CHL, and TC, these antibiotics specifically bind to their aptamers, leading to the release of the corresponding cDNA. The released cDNA hybridizes with the Cap-DNA immobilized on the surface of a gold electrode through self-assembly. Finally, oligonucleotides labeled with PbS, CdS, and ZnS QDs are added, which hybridize to the free end of the corresponding cDNA. The captured QDs, acting as signal amplifiers, are dissolved with nitric acid and measured using square wave anodic stripping voltammetry (SWASV), producing distinct electrochemical signals for each antibiotic proportional to their concentration. Furthermore, Chen et al. [52,53][49][50] pursued two approaches to develop electrochemical multiplex analysis systems for the detection of ultratrace levels of antibiotics, utilizing amine-functionalized nanoscale metal–organic framework materials (NMOF, UiO-66-NH2) as distinctive markers. Leveraging the porous nature of NMOF with its high surface area, they effectively encapsulated numerous metal ions (Cd2+ or Pb2+) within it, serving as tracers and enhancers of the analytical signal (Figure 4a). The initial system relied on specific aptamers labeled with NMOF, combined with an exonuclease amplification strategy, enabling the simultaneous determination of oxytetracycline (OTC) and kanamycin (KAN) in deproteinized milk samples [5][2]. Commercial magnetic beads were utilized to immobilize oligonucleotides, which were selectively hybridized with the aptamers labeled for each antibiotic. Upon exposure to OTC and KAN, the preferential binding of the antibiotics to the labeled aptamer caused its release and subsequent digestion by the exonuclease (Figure 4b). This resulted in the antibiotic molecules being free to interact with fresh aptamers on the electrode surface, initiating a new cycle. The supernatant, containing NMOF loaded with metal ions, was quantified using square wave voltammetry (SWV) to determine the concentrations of Cd2+ or Pb2+. Through these amplification steps, the signal was enhanced approximately 10-fold compared to a system without exonuclease. However, it is important to note that this platform has shown functionality solely in pretreated milk samples (deproteinized, dried, and reconstituted in PBS), which limits its adaptability as an aptasensor. In an alternative approach, NMOF loaded with Cd2+ or Pb2+ was employed as a marker for complementary oligonucleotides specific to KAN and CHL antibiotics [49][46]. The labeled complementary DNA strands were conjugated with their respective aptamers, which were previously immobilized on commercial magnetic beads (Figure 4b). Upon the presence of KAN and CHL, the release of the labeled DNA strands was detected via SWV. The results exhibited remarkable sensitivity with limits of detection (LODs) of 0.16 pM and 0.19 pM (S/N = 3) for KAN and CHL, respectively, representing an improvement of approximately three to four orders of magnitude compared to commercial enzyme-linked immunosorbent assay (ELISA).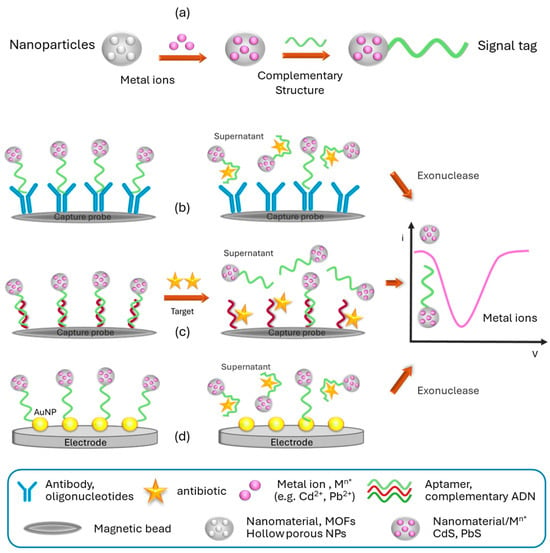
3.2. Multiplex Devices for Bacterial Recognition
Bacterial foodborne intoxication is an ever-present threat that can be prevented through the proper handling and manipulation of food products. However, it is of interest to control the presence or absence of the most common microorganisms in milk samples during processing, transportation, and storage, particularly if it can be achieved through a multianalyte analysis. In this direction, sandwich-type immunoassays, among other approaches, have been widely employed in the development of electrochemical biosensing devices for bacterial recognition. In this regard, a decade ago, Viswanathan et al. [57][54] designed a simple and disposable electrochemical immunosensor for the simultaneous measurement of three common foodborne pathogenic bacteria, namely Escherichia coli, Campylobacter, and Salmonella, in spiked milk. They utilized a previously modified SPCE with carbon nanotubes to enhance the electrochemical reaction performance, substrate interaction, and sensitivity, along with polyallylamine. The conjugated antibodies were labeled with QDs (CdS, PbS, and CuS for Escherichia coli, Campylobacter, and Salmonella, respectively). After the completion of the immunoassay, the metal ions released from the QDs, using a nitric acid solution, were analyzed by SWASV, generating independent signals for each pathogen. In another study, Eissa and Zourob [58][55] detected Listeria monocytogenes and Staphylococcus aureus using an original device consisting of a matrix of gold nanoparticle-modified SPCEs, with streptavidin immobilized on the surface of the nanoparticles. The modified SPCE surfaces were made to interact with specific biotinylated peptides, which in turn were linked to magnetic nanoparticles. Simultaneous detection was achieved by harnessing the specific proteolytic activities of proteases produced by each bacterium, acting on the respective peptides on each electrode. The cleaved magnetic nanoparticles were separated from the surface, and changes in the maximum reduction current of square wave voltammetry for the ferro/ferricyanide redox couple were recorded for each case. The multiplexed biosensor exhibited high sensitivity and selectivity, particularly against other non-specific bacterial proteases commonly found in food samples, with a response time of the order of 1 min. The use of modified GCEs has also been explored for designing platforms used in pathogen detection. For instance, Viswanath et al. [59][56] modified GCE with a zeolitic imidazolate framework (ZIF-8) and gold nanoparticles for the simultaneous detection of Aeromonas hydrophila (Ah) and Pseudomonas aeruginosa (Ps) using traditional labels such as thionine and ferrocene-conjugated antibodies, respectively. The platform was employed not only for pathogen detection in milk but also in fish tissues and juice samples. Meanwhile, Viswanath et al. [60][57] employed the same labels for the detection of Listeria monocytogenes (Lm) and Enterobacter cloacae (Ec) in milk and juice samples, coating the electrode with sandwich-like structures consisting of gold nanoparticles, carbon nanotubes, bovine serum albumin, and anti-Lm or anti-Ec.3.3. Platforms for Other Targets of Interest
Efforts in the development of devices for multiplexed electrochemical detection in milk samples have been mainly focused on monitoring antibiotics and pathogens, although occasionally, the detection of pesticides, immunoglobulins, and microRNAs has also been addressed. Recently, Ma et al. [61][58] reported a multiplexed electrochemical aptasensor based on mixed-valence Ce–MOF for the simultaneous determination of malathion (MAL) and chlorpyrifos (CLO), two organophosphate pesticides. The device was fabricated by layer-by-layer assembly on a modified GCE with gold nanoparticles, where a mixture of complementary oligonucleotides for CLO and MAL was immobilized on the nanoparticles. On the other hand, specific aptamers for the pesticides were labeled with thionine or ferrocene and hybridized with the respective immobilized complementary strands on the electrode. The presence of the pesticides released the aptamers, modifying the square wave voltametric oxidation peaks. In addition to adulterants such as melamine and other compounds [62][59], another aspect of interest regarding the quality of dairy products is the adulteration of declared milk with colostrum or milk from other animals. This aspect has been addressed by Kokkinos et al. [63][60], who designed a foldable lab-on-chip device screen-printed on a flexible membrane for the detection of bovine casein and immunoglobulin G (IgG). This determination is based on two spatially separated competitive immunoassays using biotinylated antibodies labeled with Pb or Cd QDs, conjugated with streptavidin. After completing the bioassays, the QDs are dissolved, and the device is folded, with both zones in contact with the electrochemical cell, allowing simultaneous ASV detection of the metal ions now present in the nanostructured bismuth layer formed by reduction during the preconcentration process. The device has mechanical stability, sample volumes of one drop, portability, operational and manufacturing simplicity, suitability for on-site analysis, and can be disposable. Extending this concept, Ruiz-Valdepeñas et al. [64][61] have developed a multiplexed electrochemical bioplatform that allows the detection of milk adulteration with milk or colostrum from other animals by identifying cows, sheep, or goats’ IgG in just 30 min. Its operation is based on the use of magnetic microspheres as solid support for the implementation of sandwich-type immunoassays using peroxidase conjugates for sensitive and selective IgG detection. The scope of recognition was controlled by amperometric measurements of the hydrogen peroxide/hydroquinone system using disposable electrodes. The device is capable of providing information on the animal origin of the milk, providing total and/or individual levels of IgG, the industrial heat treatment it has undergone, and whether it has been adulterated with milk or colostrum. Finally, it is also important in the dairy industry to control the health status of animals, since certain diseases can have a significant impact on production, causing significant economic and animal losses. Chand et al. [65][62] used an ECS matrix coupled with a microfluidic system with electrochemical detection for the multiplexed biodetection of specific miRNAs of paratuberculosis, a bacterial disease that affects the intestinal tract of dairy cattle. The ECSs were modified with MoS2 nanolayers decorated with copper ferrite nanoparticles that produced an amplification of the analytical signal. Meanwhile, the carrier contained MoS2 nanosheets where thiolated probes were labeled with biotin specific to miRNA and ferrocene thiol. The presence of the target miRNA triggers the opening of the molecular probe present in the nanocarriers and an increase in the ferrocene SWV signal, which was used for the determination of miRNAs in enriched serum and positive clinical samples. It is evident that the described studies present novel technological platforms for multiplexed analysis with electrochemical detection for the identification and/or quantification of target analytes in dairy samples. In Table 2, summarizes analytes are summarized along with characteristics of multiplexed sensors with electrochemical detection. These devices, primarily based on the use of biomaterials for specific recognition, possess high intrinsic potential for use in the quality control and food safety of dairy products. However, their long response times and the lack of in-depth investigation regarding their application in real samples, including derivatives, are possibly the main factors underlying their limited diffusion and application in the food industry.| Target | Analyte | Electrode/Modification/Label | Analytical Signal | Linear Range | LOD | Reference |
|---|---|---|---|---|---|---|
| Antibiotics | Sulphapyridine (SPY), Oxytetracycline (OTC) |
SPCE/protein G/– | Amperometric | 0.39, 1.93 nM |
1.92–454 nM, – |
[46][43] |
| Streptomycin (STR), Chloramphenicol (CHL), Tetracycline (TC) |
Gold electrode | SWV | – – |
10 nM, 5 nM, 20 nM |
[47][44] | |
| Kanamycin (KAN), Streptomycin (STR) |
SPCE/carbon nanofibers, carbon–gold nanoparticles/CdS, PbS | DPV | 10−1–103 nM | 87.3 pM, 45.0 pM |
[51][48] | |
| Kanamycin (KAN), Tobramycin (TOB) |
Au electrode/gold nanoshells/SCd, SPb | DPV | 1–4 × 102 nM, 1–1 × 104 nM |
0.12 nM, 0.49 nM |
[52][49] | |
| Pathogens | Escherichia coli, Campylobacter, Salmonella |
SPCE/multiwall carbon nanotube–polyallylamine/CdS, PbS, CuS QDs | SWASV | 103–5 × 105 cells/mL | 400 cells/mL, 400 cells/mL, 800 cells/mL |
[53][50] |
| Listeria monocytogenes, Staphylococcus aureus |
SPCE/gold nanoparticle–streptavidin/magnetic nanoparticles | SWV | 10–107 CFU/mL, 10–107 CFU/mL |
9 CFU/mL, 3 CFU/mL |
[54][51] | |
| Aeromonas hydrophile (Ah), Pseudomonas aeruginosa (Ps) |
GCE/ZIF-8-gold nanoparticles/thionine, ferrocene | SWV | 101–103 CFU/mL, 101–105 CFU/mL |
3.60 CFU/mL, 8095 CFU/mL |
[55][52] | |
| Listeria monocytogenes (Lm), Enterobacter cloacae (Ec) |
GCE electrodes/carbon nanotubes, Au nanoparticles anti-Lm, anti-Ec/thionine ferrocene |
SWV | 101–107 CFU/mL, 101–106 CFU/mL |
3.22 CFU/mL, 4.17 CFU/mL |
[56][ | |
| Immunoglobulins | Bovine casein, bovine immunoglobulin G |
Graphite/bismuth layer/CdS, PbS QDs | ASV | 1–102% v/v | 0.04 μg/mL, 0.02 μg/mL |
[59][56] |
| Bovine immunoglobulin G, bovine immunoglobulin G, caprine immunoglobulin G |
SPCE/–/HRP, hydrogen peroxide, hydroquinone | Amperometric | 2.6–250 ng/mL, 2.7–250 ng/mL, 2.2–250 ng/mL |
0.74 ng/mL, 0.82 ng/mL, 0.66 ng/mL |
[60][57] | |
| mi-RNA | mi-RNAs | SPCE/MoS2 nanosheets, CuFe2O4/MoS2 nanosheets, ferrocene | SWV | 1 pM to 1.5 nM | 0.48 pM | [61][58] |
| Others | Glucose, galactose, lactose, urea | Gold thin-film interdigitated sensors/AgNPs/enzymes | Impedance spectroscopy | PCA discrimination of milks with different nutritional characteristics | [66][63] | |
| Enrofloxacin Melamine |
SPCE into fluidic microarray/Au@PtNPs | Impedance spectroscopy/Cyclic voltammetry | 0.1–1000 ng/mL 0.1–500 ng/mL |
18.97 pg/mL 26.80 pg/mL |
[58][55] | |
References
- Juronen, D.; Kuusk, A.; Kivirand, K.; Rinken, A.; Rinken, T. Immunosensing system for rapid multiplex detection of mastitis-causing pathogens in milk. Talanta 2018, 178, 949–954.
- Duan, N.; Gong, W.; Wang, Z.; Wu, S. An aptasensor based on fluorescence resonance energy transfer for multiplexed pathogenic bacteria determination. Anal. Methods 2016, 8, 1390–1395.
- Huang, Y.; Zhang, H.; Chen, X.; Wang, X.; Duan, N.; Wu, S.; Xu, B.; Wang, Z. A multicolor time-resolved fluorescence aptasensor for the simultaneous detection of multiplex Staphylococcus aureus enterotoxins in the milk. Biosens. Bioelectron. 2015, 74, 170–176.
- Shriver-Lake, L.C.; Erickson, J.S.; Sapsford, K.E.; Ngundi, M.M.; Shaffer, K.M.; Kulagina, N.V.; Hu, J.E.; Gray III, S.A.; Golden, J.P.; Ligler, F.S.; et al. Blind laboratory trials for multiple pathogens in spiked food matrices. Anal. Lett. 2007, 40, 3219–3231.
- Cho, I.H.; Mauer, L.; Irudayaraj, J. In-situ fluorescent immunomagnetic multiplex detection of foodborne pathogens in very low numbers. Biosens. Bioelectron. 2014, 57, 143–148.
- Wang, C.; Xiao, R.; Wang, S.; Yang, X.; Bai, Z.; Li, X.; Rong, Z.; Shen, B.; Wang, S. Magnetic quantum dot based lateral flow assay biosensor for multiplex and sensitive detection of protein toxins in food samples. Biosens. Bioelectron. 2019, 146, 111754.
- Shi, Q.; Tao, C.; Kong, D. Multiplex SERS-based lateral flow assay for one-step simultaneous detection of neomycin and lincomycin in milk. Eur. Food Res. Technol. 2022, 248, 2157–2165.
- Li, X.; Wang, X.; Wang, L.; Yang, T.; Wang, D. Duplex Detection of Antibiotics in Milk Powder Using Lateral-Flow Assay Based on Surface-Enhanced Raman Spectroscopy. Food Anal. Methods 2021, 14, 165–171.
- Li, Y.; Jin, G.; Liu, L.; Kuang, H.; Jing, X.; Chuanlai, X. A portable fluorescent microsphere-based lateral flow immunosensor for the simultaneous detection of colistin and bacitracin in milk. Analyst 2020, 145, 7884–7892.
- Li, Z.; Li, Z.; Zhao, D.; Wen, F.; Jiang, J.; Xu, D. Smartphone-based visualized microarray detection for multiplexed harmful substances in milk. Biosens. Bioelectron. 2017, 87, 874–880.
- Zhou, C.; Huang, C.; Zhang, H.; Yang, W.; Jiang, F.; Chen, G.; Liu, S.; Chen, Y. Machine-Learning-driven optical immunosensor based on microespheres-encoded signal transduction for the rapid and multiplexed detection of antibiotics in milk. Food Chem. 2024, 437, 137740.
- Shi, Q.; Huang, J.; Sun, Y.; Yin, M.; Hu, M.; Hu, X.; Zhang, Z.; Zhang, G. Utilization of a lateral flow colloidal gold immunoassay strip based on surface-enhanced Raman spectroscopy for ultrasensitive detection of antibiotics in milk. Spectrochim. Acta A Mol. Biomol. Spectrosc. 2018, 197, 107–113.
- Jiang, R.; Lin, D.; Zhang, Q.; Li, L.; Yang, L. Multiplex chroma-response based fluorescent smartphone sensing platform for rapid and visual quantitaive determination of antibiotic. Sens. Actuators B. Chem. 2022, 350, 130902.
- Lu, L.; Xu, L.; Zhang, Y.; Jiang, T. Multiplexed surface-enhanced Raman scattering detection of melamine and dicyandiamide in dairy food enabled by three-dimensional polystyrene@silver@graphene oxide hybrid substrate. Appl. Surf. Sci. 2022, 603, 154419.
- Zhang, H.; Ma, X.; Liu, Y.; Duan, N.; Wu, S.; Wang, Z.; Xu, B. Gold nanoparticles enhanced SERS aptasensor for the simultaneous detection of Salmonella typhimurium and Staphylococcus aureus. Biosens. Bioelectron. 2015, 74, 872–877.
- Zhang, Z.; Tang, S.; Jin, Y.; Yang, C.; He, L.; Wang, J.; Chen, Y. Multiplex SERS-based lateral flow immunosensor for the detection of major mycotoxins in maize utilizing dual Raman labels and triple test lines. J. Hazard. Mater. 2020, 393, 122348.
- Yahaya, M.L.; Zakaria, N.D.; Noordin, R.; Abdul Razak, K. Development of rapid gold nanoparticles based lateral flow assays for simultaneous detection of Shigella and Salmonella genera. Biotechnol. Appl. Biochem. 2021, 68, 1095–1106.
- Tao, X.; Wang, J.; Xie, Y.; Zuo, X.; Mo, F.; Zhou, S.; Li, H. Dual-Label Chemiluminescence Strategy for Multiplexed Immunoassay of 20 Fluoroquinolones, 15 β-Lactams. 15 Sulfonamides, and CAP in Milk. Food Anal. Methods 2017, 10, 3009–3022.
- Yu, X.; Tao, X.; Shen, J.; Zhang, S.; Cao, X.; Chen, M.; Wang, W.; Wang, Z.; Wen, K. A one-step chemiluminescence immunoassay for 20 fluoroquinolone residues in fish and shrimp based on a single chain Fv–alkaline phosphatase fusion protein. Anal. Methods 2015, 7, 9032–9039.
- Zhang, Y.; Chang, X.; Wang, X.; Tao, X. A quadruple-labeling luminescence strategy for multiplexed immunoassay of 51 drugs in milk with an automated pretreatment system. Anal. Methods 2019, 11, 5055–5063.
- Singh, A.K.; Sun, X.; Bai, X.; Kim, H.; Abdalhaseib, M.U.; Bae, E.; Bhunia, A.K. Label-free, non-invasive light scattering sensor for rapid screening of Bacillus colonies. J. Microbiol. Methods 2015, 109, 56–66.
- Raz, S.R.; Bremer, M.G.E.G.; Haasnoot, W.; Norde, W. Label-Free and Multiplex Detection of Antibiotic Residues in Milk Using Imaging Surface Plasmon Resonance-Based Immunosensor. Anal. Chem. 2009, 81, 7743–7749.
- Suarez, G.; Jin, Y.H.; Auerswald, J.; Berchtold, S.; Knapp, H.F.; Diserens, J.M.; Leterrier, Y.; Manson, J.A.E.; Voirin, G. Lab-on-a-chip for multiplexed biosensing of residual antibiotics in milk. Lab Chip 2009, 9, 1625–1630.
- Angelopoulou, M.; Petrou, P.S.; Makarona, E.; Haasnoot, W.; Moser, I.; Jobst, G.; Goustouridis, D.; Lees, M.; Kalatzi, K.; Raptis, I.; et al. Ultrafast Multiplexed-Allergen Detection through Advanced Fluidic Design and Monolithic Interferometric Silicon Chips. Anal. Chem. 2018, 90, 9559–9567.
- Yang, Z.; Xu, G.; Reboud, J.; Ali, S.A.; Kaur, G.; McGiven, J.; Boby, N.; Gupta, P.K.; Chaudhuri, P.; Cooper, J.M. Rapid Veterinary Diagnosis of Bovine Reproductive Infectious Diseases from Semen Using Paper-Origami DNA Microfluidics. ACS Sens. 2018, 3, 403–409.
- Prasad, A.; Tran, T.; Gartia, M.R. Multiplexed Paper Microfluidics for Titration and Detection of Ingredients in Beverages. Sensors 2019, 19, 1286.
- Chen, X.; Yao, C.; Li, Z. Microarray-based chemical sensors and biosensors: Fundamentals and food safety applications, TrAC. Trends Anal. Chem. 2023, 158, 116785.
- Ashley, J.; D’Aurelio, R.; Piekarska, M.; Temblay, J.; Pleasants, M.; Trinh, L.; Rodgers, T.L.; Tothill, I.E. Development of a β-Lactoglobulin Sensor Based on SPR for Milk Allergens Detection. Biosensors 2018, 8, 32.
- Bojcukova, J.; Vlas, T.; Forstenlechner, P.; Panzner, P. Comparison of two multiplex arrays in the diagnostics of allergy. Clin. Transl. Allergy 2019, 9, 31.
- Scala, E.; Caprini, E.; Abeni, D.; Meneguzzi, G.; Buzzulini, F.; Cecchi, L.; Villalta, D.; Asero, R. A qualitative and quantitative comparison of IgE antibody profiles with two multiplex platforms for component-resolved diagnostics in allergic patients. Clin. Exp. Allergy 2021, 51, 1603–1612.
- ImmunoCAP ISAC Test. Available online: https://www.thermofisher.com/phadia/wo/en/our-solutions/immunocap-allergy-solutions/specific-ige-multiplex.html (accessed on 14 February 2024).
- ELISA Based In-Vitro Multiplex Allergy Test. Available online: https://www.macroarraydx.com/products/alex (accessed on 14 February 2024).
- Monroe, M.R.; Daaboul, G.G.; Tuysuzoglu, A.; López, C.A.; Little, F.F.; Ünlü, M.S. Single Nanoparticle Detection for Multiplexed Protein Diagnostics with Attomolar Sensitivity in Serum and Unprocessed Whole Blood. Anal. Chem. 2013, 85, 3698–3706.
- Galan-Malo, P.; Pellicer, S.; Pérez, M.D.; Sánchez, L.; Razquin, P.; Mata, L. Development of a novel duplex lateral flow test for simultaneous detection of casein and β-lactoglobulin in food. Food Chem. 2019, 293, 41–48.
- Masiri, J.; Barrios-López, B.; Benoit, L.; Tamayo, J.; Day, J.; Nadala, C.; Sung, S.L.; Samadpour, M. Development and Validation of a Lateral Flow Immunoassay Test Kit for Dual Detection of Casein and β-Lactoglobulin Residues. J. Food Prot. 2016, 79, 477–483.
- Badran, A.A.; Morais, S.; Maquieira, Á. Simultaneous determination of four food allergens using compact disc immunoassaying technology. Anal. Bioanal. Chem. 2017, 409, 2261–2268.
- Raz, S.R.; Liu, H.; Norde, W.; Bremer, M.G.E.G. Food Allergens Profiling with an Imaging Surface Plasmon Resonance-Based Biosensor. Anal. Chem. 2010, 82, 8485–8491.
- Tortajada-Genaro, L.A.; Santiago-Felipe, S.; Morais, S.; Gabaldón, J.A.; Puchades, R.; Maquieira, Á. Multiplex DNA Detection of Food Allergens on a Digital Versatile Disk. J. Agric. Food Chem. 2012, 60, 36–43.
- Blais, B.W.; Gaudreault, M.; Phillippe, L.M. Multiplex enzyme immunoassay system for the simultaneous detection of multiple allergens in foods. Food Control 2003, 14, 43–47.
- Lin, H.Y.; Huang, C.H.; Park, J.; Pathania, D.; Castro, C.M.; Fasano, A.; Weissleder, R.; Lee, H. Integrated Magneto-Chemical Sensor for On-Site Food Allergen Detection. ACS Nano 2017, 11, 10062–10069.
- Filep, S.C.; Black, K.R.; Smith, B.R.E.; Block, D.S.; Kuklinska-Pijanka, A.; Bermingham, M.; Oliver, M.A.; Thorpe, C.M.; Schuhmacher, Z.P.; Agah, S.; et al. Simultaneous quantification of specific food allergen proteins using a fluorescent multiplex array. Food Chem. 2022, 389, 132986.
- Gomaa, A.; Boye, J. Simultaneous detection of multi-allergens in an incurred food matrix using ELISA, multiplex flow cytometry and liquid chromatography mass spectrometry (LC–MS). Food Chem. 2015, 175, 585–592.
- Ross, G.M.S.; Filippini, D.; Nielen, M.W.F.; Salentijn, G.I.J. Interconnectable solid-liquid protein extraction unit and chip-based dilution for multiplexed consumer immunodiagnostics. Anal. Chim. Acta 2020, 1140, 190–198.
- Sena-Torralba, A.; Smits, N.G.E.; Blázquez, D.; Albero-Pérez, C.; Pallás-Tamarit, Y.; Maquieira, Á.; Morais, S. Simultaneous quantification of six major allergens in commercial foods for children using a multiplex array on a digital versatile disc. Food Chem. 2023, 404, 134570.
- Grabowska, I.; Hepel, M.; Kurzątkowska-Adaszyńska, K. Advances in Design Strategies of Multiplex Electrochemical Aptasensors. Sensors 2022, 22, 161.
- Rapini, R.; Marrazza, G. Electrochemical aptasensors for contaminants detection in food and environment: Recent advances. Bioelectrochemistry 2017, 118, 47–61.
- Conzuelo, F.; Campuzano, S.; Gamella, M.; Pinacho, D.G.; Reviejo, A.J.; Marco, M.P.; Pingarrón, J.M. Integrated disposable electrochemical immunosensors for the simultaneous determination of sulfonamide and tetracycline antibiotics residues in milk. Biosens. Bioelectron. 2013, 50, 100–105.
- Xue, J.; Liu, J.; Wang, C.; Tiana, Y.; Zhou, N. Simultaneous electrochemical detection of multiple antibiotic residues in milk based on aptamers and quantum dots. Anal. Methods 2016, 8, 1981–1988.
- Chen, M.; Gan, N.; Zhou, Y.; Li, T.; Xua, Q.; Cao, Y.; Chen, Y. An electrochemical aptasensor for multiplex antibiotics detection based on metal ions doped nanoscale MOFs as signal tracers and RecJf exonuclease assisted targets recycling amplification. Talanta 2016, 161, 867–874.
- Chen, M.; Gan, N.; Zhou, Y.; Li, T.; Xua, Q.; Cao, Y.; Chen, Y. A novel aptamer- metal ions- nanoscale MOF based electrochemicalbiocodes for multiple antibiotics detection and signal amplification. Sens. Actuators B Chem. 2017, 242, 1201–1209.
- Yan, Z.; Gan, N.; Li, T.; Cao, Y.; Chen, Y. A sensitive electrochemical aptasensor for multiplex antibiotics detection based on high-capacity magnetic hollow porous nanotracers coupling exonuclease-assisted cascade target recycling. Biosens. Bioelectron. 2016, 78, 51–57.
- Li, F.; Wang, X.; Sun, X.; Guo, Y. Multiplex electrochemical aptasensor for detecting multipleantibiotics residues based on carbon fiber and mesoporouscarbon-gold nanoparticles. Sens. Actuators B Chem. 2018, 265, 217–226.
- Li, F.; Wu, Y.; Chen, D.; Guo, Y.; Wang, X.; Sun, X. Sensitive dual-labeled electrochemical aptasensor for simultaneous detection of multi-antibiotics in milk. Int. J. Hydrogen Energy 2021, 46, 23301–23309.
- Viswanathan, S.; Rani, C.; Ho, L.A. Electrochemical immunosensor for multiplexed detection of food-borne pathogens using nanocrystal bioconjugates and MWCNT screen-printed electrode. Talanta 2012, 94, 315–319.
- Eissa, S.; Zourob, M. Ultrasensitive peptide-based multiplexed electrochemical biosensor for the simultaneous detection of Listeria monocytogenes and Staphylococcus aureus. Mikrochim. Acta 2020, 187, 486.
- Viswanath, K.B.; Krithiga, N.; Jayachitra, A.; Mideen, A.K.S.; Amali, A.J.; Vasantha, V.S. Enzyme-Free Multiplex Detection of Pseudomonas aeruginosa and Aeromonas hydrophila with Ferrocene and Thionine-Labeled Antibodies Using ZIF-8/Au NPs as a Platform. ACS Omega 2018, 3, 17010–17022.
- Viswanath, K.B.; Suganya, K.; Krishnamoorthy, G.; Marudhamuthu, M.; Selvan, S.T.; Vasantha, V.S. Enzyme-Free Multiplex Detection of Foodborne Pathogens Using Au Nanoparticles-Decorated Multiwalled Carbon Nanotubes. ACS Food Sci. Technol. 2021, 1, 1236–1246.
- Ma, D.; Liu, J.; Liu, H.; Yi, J.; Xia, F.; Tian, D.; Zhou, C. Multiplexed electrochemical aptasensor based on mixed valence Ce (III, IV)-MOF for simultaneous determination of malathion and chlorpyrifos. Anal. Chim. Acta 2022, 1230, 340364.
- Gu, Y.; Wang, J.; Pan, M.; Yun, Y.; Wen, W.; Fang, G.; Wang, S. On-chip multiplex electrochemical immunosensor based on disposable 24-site fluidic micro-array screen printing analytical device for multi-component quantitative analysis. Sens. Actuat. B Chem. 2018, 260, 499–507.
- Kokkinos, C.; Angelopoulou, M.; Economou, A.; Prodromidis, M.; Florou, A.; Haasnoot, W.; Petrou, P.; Kakabakos, S. Lab-on-a-Membrane Foldable Devices for Duplex Drop-Volume Electrochemical Biosensing Using Quantum Dot Tags. Anal. Chem. 2016, 88, 6897–6904.
- Ruiz-Valdepeñas Montiel, V.; Povedano, E.; Benedé, S.; Mata, L.; Galán-Malo, P.; Gamella, M.; Reviejo, A.J.; Campuzano, S.; Pingarrón, J.M. Disposable Amperometric Immunosensor for the Detection of Adulteration in Milk through Single or Multiplexed Determination of Bovine, Ovine, or Caprine Immunoglobulins G. Anal. Chem. 2019, 91, 11266–11274.
- Chand, R.; Ramalingam, S.; Neethirajan, S. A 2D transition-metal dichalcogenide MoS2 based novel nanocomposite and nanocarrier for multiplex miRNA detection. Nanoscale 2018, 10, 8217–8225.
- Perez-Gonzalez, C.; Salvo-Comino, C.; Martin-Pedrosa, F.; Rodriguez-Mendez, M.L.; García-Cabezón, C. A new data analysis approach for an AgNPs-modified impedimetric bioelectronic tongue for dairy analysis. Food Control 2024, 156, 110136.
