The soil microbiome plays an important role in maintaining soil health, plant productivity, and soil ecosystem services. Molecular-based studies have shed light on the fact that the soil microbiome has been quantitatively underestimated. In addition to metagenomic studies, metaproteomics and metatranscriptomic studies that target the functional part of the microbiome are becoming more common. These are important for a better understanding of the functional role of the microbiome and for deciphering plant-microbe interactions. Free-living beneficial bacteria that promote plant growth by colonizing plant roots are called plant growth-promoting rhizobacteria (PGPRs). They exert their beneficial effects in different ways, either by facilitating the uptake of nutrients and synthesizing particular compounds for plants or by preventing and protecting plants from diseases.
- rhizosphere
- microbiome
- sustainable agriculture
1. Introduction
2. Soil Microbiome
2.1. Spatial Distribution of Soil Microbiome
Microorganisms are one of the most abundant living organisms on Earth, constituting approximately 17% of the global biomass [2]. Soil is the most complex habitat that contains a huge abundance of microbial life, which comprises approximately 4–5 × 1030 microbial cells [3]. The soil microbiome is mainly comprised of soil bacteria, archaea, fungi and viruses. Mendes et al. estimates that 108–109 bacteria, 107–108 viruses and 105–106 fungal cells are in one gram of soil [4]. Soil microbial communities provide ecosystem services as nutrient recycling, carbon sequestration, water retention, plant growth promotion and defense [5,6,7][5][6][7]. The focus on soil microbiota research has become noticeable due to its role in the global carbon cycle and climate change, as well as in sustainable agriculture. The diversity and abundance of microbes are affected by land-use patterns and soil compartments. Agricultural ecosystems are more homogeneous than natural environments due to lower plant diversity and frequent human disturbance. Two soil compartments can be distinguished based on the strength of their relationship with the plant roots. Bulk soil is defined as the part of the soil that is not or is loosely attached to the root, whereas the attached part of the soil is considered rhizosphere soil. Bacterial community composition also differs considerably between bulk and rhizosphere soils, decreasing diversity from the bulk soil to the roots [8]. In tobacco and Arabidopsis plants, the number of microbes present in the rhizosphere is approximately 10- to 100-fold higher than that in bulk soil [9]. The rhizosphere is considered a biological hotspot where plant–microbe, microbe–microbe, and microbe–plant interactions shape microbial community composition. Plant roots secrete organic compounds that support microbial activity [8]. Rhizosphere soil contains 108–1011 cultivable cells in one gram of soil, which corresponds to approximately 104 microbial species [10]. The spatial heterogeneity of the soil microbiome is determined on the one hand by environmental factors and on the other hand by populational processes [11]. Environmental factors can be both biotic and abiotic, and soil microbial colonization is influenced by plant root exudates in the rhizosphere and environmental parameters. The bulk soil microbial community is an important factor that shapes the rhizosphere microbiome, being the main reservoir from which soil microorganisms are attracted by chemotaxis to root exudates [9]. The same taxa are therefore present in bulk and rhizosphere soils but differ in their relative abundance [12]. Differences in the microbial community between bulk and rhizosphere soils were studied in maize fields by Li et al. [13]. They observed that the rhizosphere soil microbiota was enriched in Proteobacteria, Bacteroidetes, and Actinobacteria, accounting for 73–80% of total reads versus 46–56% in bulk soil. A decreased abundance was observed for Acidobacteria, Gemmatimonadetes, Chloroflexi, Firmicutes, and Nitrospira in the rhizosphere relative to that in the bulk soil [13]. Fan et al. studied the microbial community of wheat fields with an emphasis on three soil compartments, namely tightly and loosely bound soil and bulk soil [8]. They found that Proteobacteria, Actinobacteria, and Acidobacteria dominated all soils, whereas the abundance of Actinobacteria, Bacteroidetes, Alphaproteobacteria, and Verrucomicrobia was higher and the abundance of Gammaproteobacteria, Chloroflexi, and Deltaproteobacteria was lower in tightly bound soil than in the other soils. The greater relative abundance of Actinobacteria in tightly bound soil was explained by their antibiotic-producing potential, whereas the presence of Alphaproteobacteria was attributed to their fast growth characteristics.2.2. Soil Microbiome Taxonomic Diversity: Structure and Function
2.2.1. Taxonomic Diversity
The rhizosphere microbiome includes different groups of bacteria. They belong to the following phyla: Acidobacteria, Actinobacteria, Ascomycota, Bacteroidetes, Basidiomycota, Deinococcus-Thermus, Euryarchaeota, Firmicutes, and Proteobacteria [15][14]. The bacterial community diversity and composition differ between different crop plants, with higher differences reported between different plant types, such as legumes, forbs, and grasses (Figure 1).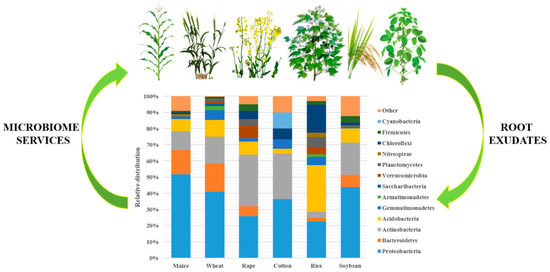
2.2.2. Factors Affecting Diversity
The main environmental factors affecting the community structure of the rhizosphere microbiome are pH, salinity, moisture, temperature, and nutrient content (C-N content and other nutrients); however, vegetation also has an important role [15,23][14][20]. Environmental factors create unique ecological niches that frame specific microbiomes. The spatial heterogeneity of the soil microbiome was studied in the case of switchgrass vegetation, where the bacterial community structure proved to be patchy, and the abundance of the dominant phyla (Verrucomicrobia) changed 2.5 fold in a 10 cm3 grid [5]. Anthropogenic effects that cause biotic and abiotic stresses and changing climatic conditions modify soil microbial and plant diversity [10,25][10][21]. Ullah et al. identified drought-tolerant bacteria in drought-treated cotton plants [20][17]. Thermophilic bacteria belonging to the phyla Chloroflexi and Gemmatimonadetes were found to be dominant in drought-affected environments [20][17]. The use of fertilizers, farming, and tillage methods in agroecosystems can induce changes in soil microbiomes. In the case of the switchgrass rhizosphere microbiome, a fertilizer-induced decrease in the relative abundance of the most abundant phylum (Verrucomicrobia) was observed [5]. The use of different soil amendments also reduced bulk soil microbiome diversity and influenced the rhizosphere community in Zea mays. Changes in community structure were caused by a lower abundance of Actinobacteria and Firmicutes and a higher abundance of Proteobacteria, Bacteroidetes, Verrucomicrobia, and Acidobacteria [12]. Fertilizer type and dose also contributed to changes in rhizobial community structure in Z. mays, and differences in the abundance of microbial groups were attributed to their different nutrient contents [26][22]. Tillage practices modified the root- and shoot-associated bacterial communities in rape plants, whereas farming practices affected the microbial community structure of rice [21][18].2.2.3. Ecological Function
In microbial communities, it is important to study species composition and existing ecological functional groups. Different microbes have different roles in the community structure, which might support soil health and plant productivity. Microbial communities are complex dynamic networks with various interactions between microbes, such as resource competition, metabolic dependencies (cross-feeding), spatial organization notably production of biofilms, signaling, horizontal gene transfer, coevolution, and viral looting [27][23]. Usually, a higher diversity of microbes increases the quantity of metabolites, secondary metabolites, phytohormones, biocontrol substances, and other beneficial substances, thereby contributing to soil structure and fertility, root system architecture and nutrient foraging, plant nutrition and hormonal balance, plant stress tolerance, agricultural productivity, and resilience to climate, land use, and agronomic practices [10].2.3. Beneficial Plant–Microbe Interactions
2.3.1. Biostimulant Microbes
Microorganisms and plants live in nature in association, but the microorganisms can be free-living, attached, or enter symbiosis with the host plants. There are different interactions such as commensalism, mutualism or parasitism. During evolution, plants interacted with a broad range of plant growth-promoting rhizobacteria (PGPR). Owing to recent advances in metagenomics, massive genome-sequencing strategies, and new identification techniques, bacterial rhizobiome mapping is rapidly progressing. These findings revealed novel bacterial species and their mechanisms involved in biocontrol and plant growth promotion [30][24]. Microorganisms living in soil can indirectly promote plant growth (Figure 2), especially by fixation of atmospheric N2, production of siderophores, plant growth hormones (cytokinins, auxins, and gibberellins), volatile compounds, and solubilization of nutrients and minerals (phosphorus, potassium, zinc, etc.).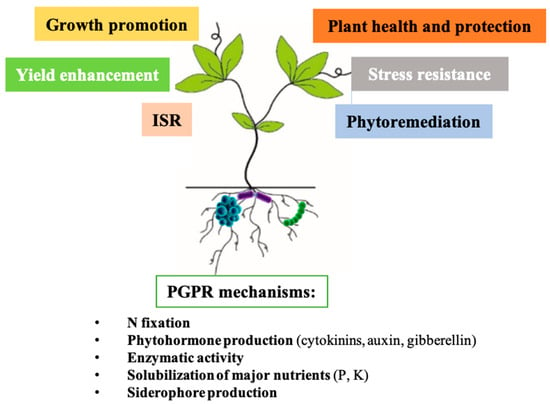
Biological Nitrogen Fixation
Phytohormone Production
Phytohormones are organic compounds that influence physiological processes in plants even at very low concentrations. The ability of soil microbiota to produce phytohormones is a potential source of phytohormones. Plant growth hormones, such as auxins (indole-3-acetic acid), gibberellins, abscisic acids, ethylene, and cytokinins, are biosynthesized by microorganisms. Phytohormones play a significant role in plant growth during cell division and enlargement, seed germination, root formation, and stem elongation. Phytohormones produced by bacteria are released into the plant body and have a positive effect on plant growth and development. Several reports have shown that bacteria can produce 60 times more plant growth regulator substances than plants can. All plant-associated microbes produce auxins, but not all PGP microbes have the ability to produce gibberellin. This capacity is related to root-associated microbes. Auxins, mostly indole-3-acetic acid (IAA), are synthesized by 80% of rhizosphere bacteria. Tryptophan is the main precursor for IAA biosynthesis in bacteria. Bacteria that promote IAA synthesis can take up tryptophan present in root exudates. There are five different tryptophan-dependent and tryptophan-independent pathways, as in Azospirillum brasilense, in which the biosynthetic intermediates are unknown [38][29]. Among bacterial phytohormones, IAA, which promotes root elongation and lateral root development, is the most studied. These plant hormones are highly effective under stressful conditions. Some plants are unable to produce enough auxins to cope with stress effects; therefore, bacterial auxins can alleviate stress conditions in plants [39][30]. Bacterial strains with IAA production capacity include Pseudomonas fluorescens, Pseudomonas syringae, Agrobacterium tumefaciens, Pantoea agglomerans, Azospirillum brasilense, Bacillus cereus, Bacillus amyloliquefaciens, Rhizobium sp., and Bradyrhizobium sp. [38][29]. Abscisic acid (ABA) is a stress-related hormone that plays a key role in photoperiodic induction of flowering, contributing to plant growth and development. It is involved in plant responses to different environmental stresses such as cold, salinity, and desiccation [39][30]. Several plant-associated bacteria can produce ABA, thereby increasing phytohormone levels in plants. ABA is an important factor in modulating plant defenses, so plant mutants with altered ABA biosynthesis or that are ABA-insensitive are more resistant to pathogens than wild-type plants [38][29]. ABA-producing endophytic bacteria include Achromobacter xylosoxidans, Brevibacterium halotolerans, Bacillus licheniformis, Bacillus pumilus, and Lysinibacillus fusiformis [42][31]. The gibberellin (GA) phytohormone plays a major role in leaf expansion and stem elongation in plants. When GA is applied exogenously, it can promote parthenocarpy in fruits, bolting plants, breaking tuber dormancy, and increasing fruit size and the number of buds. Several soil microorganisms have been reported to produce gibberellin, with positive or negative effects on nodulation and plant growth. These microorganisms can induce nodule organogenesis and inhibit nodulation during the infection stage [39][30]. The first described bacterium with GA production ability was Rhizobium phaseoli, which produces GA1 and GA4. Cytokinins (CK) play a role in many stages of plant development by stimulating plant cell division, root development, and root hair formation, activating dormant buds, and inducing the germination of plant seeds. These plant hormones affect apical dominance and regulate nodulation and nitrogen fixation. Some pathogenic and beneficial microbes produce cytokinin phytohormones. It has been reported that PGPRs from Pseudomonas and Bacillus genera produce cytokinin, especially zeatin [15,35][14][28]. Pseudomonas fluorescens and Rhizobium spp. are cytokinin-producing bacteria [43][32]. Ethylene (ET) is a plant stress hormone. Under stress conditions, higher amounts of ethylene can negatively affect plant growth. Ethylene production can be modulated by bacterial strains possessing 1-aminocyclopropane-1-carboxylic acid (ACC) deaminase activity. PGPRS acts as a sink for the ET precursor, 1-aminocyclopropane-1-carboxylic acid (ACC), consequently reducing ET levels in roots and simultaneously increasing root length and plant growth. ACC exuded by roots and seeds can be taken up by rhizobacteria, and due to activity of ACC deaminase (ACCd) is split into ammonia and α-ketobutyrate.Enzymatic Activity
1-aminocyclopropane-1-carboxylate deaminase (ACC-deaminase, ACCd) enzyme plays an important role in plant hormone and ethylene regulation. ACC deaminase is found in numerous microbial species, including Gram-negative and Gram-positive bacteria and fungi [46,47][33][34].Solubilization of Major Nutrients
Soil microorganisms play a major role in nutrient cycling. The crop residues incorporated in the soil represent the carbon, energy, and nutrient sources of microorganisms. Rhizobia can solubilize nutrients such as phosphorus, iron, potassium, and zinc, thus increasing their availability to plants [46][33]. Among the macronutrients, phosphorus (P) is essential for plant growth and development. P is abundantly available in the soil in organic (phytin) and inorganic forms (P minerals such as apatite and secondary P minerals such as Al, Fe, and Ca phosphates). P is a major growth-limiting nutrient despite being present in soil in abundancy in insoluble form. The soluble level of P in soil determines the P accessibility to plants [38,44][29][35]. PGP microbes are a biological rescue system because they are capable of solubilizing insoluble inorganic P in soil, increasing its availability to plants in the form of orthophosphate. The major mechanism of P solubilization involves the production of organic acids. As a result, insoluble P is transformed into its soluble form. The produced organic acids decrease the soil pH or chelate mineral ions, resulting in phosphate solubilization [38][29]. The organic acids most frequently produced by Gram-negative PGPRs are gluconic acid and 2-ketogluconic acid. PGPRs can release from soil organic and inorganic phosphorus by producing several enzymes, such as phytases, phosphatases, phosphanatases and lyases [38][29]. In this process, the microbes also produce organic acids (gluconate, acetate, ketogluconate, oxalate, lactate, tartarate, succinate, citrate, and glycolate), but this depends on the type of carbon source used as substrate. The highest amount of solubilized P was observed in in vitro conditions when glucose, sucrose or galactose was used as the sole carbon source [15][14]. Potassium (K) is considered the third major macronutrient for plant growth and crop yields. More than 90% of the potassium that exists in soil is in the form of insoluble rocks and silicate minerals. The soluble form of K is present in soil in low concentration. One of the major constraints in crop production is the potassium deficiency due to imbalanced fertilizer application. Lack of potassium causes poorly developed roots, small seeds, lower crop yields and slow growth. An alternative indigenous source of potassium for plants is the potassium solubilized by soil microorganisms.Solubilization of Iron with Siderophore Production
The bioavailability of iron as an essential micronutrient is limited by the soil. Siderophores produced by soil bacteria play a key role in plant iron nutrition. These compounds are low-molecular-weight chelators with a high affinity for iron (III), the most common form of iron in nature. The iron solubilization mechanism relies on the formation of a stable siderophore-Fe+3 complex that can be absorbed by plants [38,48][29][36]. To date, more than 500 siderophores have been identified. Plant growth-promoting Pseudomonas fluorescens produces pyoverdine among other siderophores. Microorganisms can produce other siderophoric compounds such as catechol, hydroxamate, carboxylate, and phenolate, which contribute to plant protection against pathogens. Bacterial strains with iron chelation properties belong to Azotobacter, Bacillus, Enterobacter, Arthrobacter, Nocardia, and Streptomyces [46][33]. The direct beneficial effect of siderophores is the improvement in the iron nutritional status of the plant, contributing to plant growth promotion. It has been hypothesized that bacterial siderophores chelate Fe+3 from the soil, making it accessible to phytosiderophores, but the exact mechanism is unknown. It has been shown that plants can incorporate Fe+3-pyoverdine complexes resulting in an increase in the iron content of plant tissues. The indirect beneficial role of bacterial siderophores in plant growth promotion is their capacity to reduce the availability of iron to phytopathogens [38][29]. Siderophore synthesis is influenced by several environmental factors such as pH, the level of iron, the presence of other trace elements, and an adequate supply of carbon, phosphorus, and nitrogen sources [48][36]. Siderophores transport iron into bacterial cells by means of a system involving ferric-specific ligands (siderophores) and their corresponding membrane receptors, which are chelating agents in bacteria.2.3.2. Biocontrol Activity of Microbes
The overuse of chemicals in agriculture, such as pesticides, insecticides, herbicides, and fertilizers, negatively affects consumer health, biomagnification of chemicals, and economic loss [49,50][37][38]. Biological control organisms are defined as living organisms other than disease-resistant host plants that suppress the activity of plant pathogens in the soil environment [51][39]. Microbial control agents can exert their plant-protecting characteristics based on their mode of interaction with pathogens through direct and indirect mechanisms (Figure 3).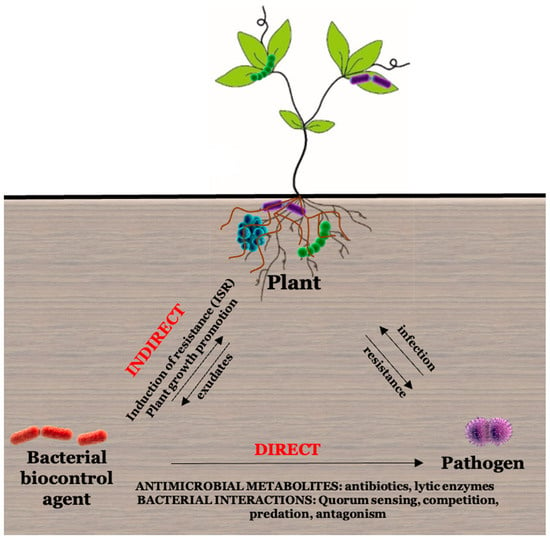
Antibiotics
Interference of Quorum Sensing with Virulence
Quorum sensing is a cell–cell communication process in bacteria that involves extracellular signaling molecules (autoinducers) and serves to share information about cell density. Because many processes are advantageous only in this group, when the bacterial population increases, gene expression is altered. Processes such as biofilm formation, antibiotic production, and virulence factor secretion are controlled by QS. Quorum sensing (QS) is important for expressing bacterial pathogenicity in plants. QS is required for the colonization and expression of virulence factors in plant pathogenic strains, such as Pseudomonas syringae, Pantoea stewartii, Erwinia chrysanthemi, and Burkholderia glumae [66,67,68,69][42][43][44][45].Lytic Enzymes
A potential mechanism of action against pathogens is the production of lytic enzymes. PGPRs inhibit the growth of fungal pathogens (Fusarium oxysporum, Sclerotinia sclerotiorum, and Botrytis cinerea) and other soil-borne pathogens through the excretion of enzymes such as chitinases, hydrolases, proteases, and glucanases [46,48][33][36].Induced Systemic Resistance (ISR)
Disease control by various beneficial bacterial strains involves the induction of systemic resistance. Different microbial metabolites and biocontrol agents can generate an immune response in the host plant, resulting in systemic disease resistance [46][33]. Plants recognize microbial compounds (flagellin, lipopolysaccharide, exopolysaccharide, and chitin oligosaccharides) produced by beneficial microorganisms. Different bacterial species are effective against fungal, bacterial, and viral infections through ISR, including Bacillus amyloliquefaciens, B. atrophaeus, B. cereus, B. megaterium, B. subtilis, Paenibacillus alvei, Pseudomonas fluorescens, Pseudomonas aeruginosa, and Streptomyces pactum [82][46].2.4. Plant-Beneficial Function Encoding Gene Clusters and Mobile Genetic Elements
Horizontal gene transfer (HGT) is an event in which bacteria incorporate advantageous genes into their genomes. Horizontally transmitted genes are crucial for bacterial adaptation to changing environments or to plant-microbe interactions. They are often grouped into genomic islands and gene clusters [83][47]. Up to 20% of the bacterial genome is disseminated during horizontal gene transfer events [84][48]. The rhizosphere is considered to be one of the hotspots of microbial gene transfer whereas the microbiome is a rich reservoir of genetic functionality [85,86][49][50]. Many function genes in soil bacteria are encoded by plasmids that act as mobile genetic elements (MGEs). Plasmids are most commonly considered antibiotic resistance gene carriers; however, they are also important carriers of heavy metal detoxification genes, N fixation genes, and other plant growth stimulation genes. The pSym plasmid found in Rhizobium sp., in addition to nodulation and atmospheric nitrogen fixation genes, is involved in phytohormone synthesis and transport of root exudate compounds. This conjugative plasmid is commonly transferred to the soil or rhizosphere community, mainly after sensing certain plant compounds such as flavonoids [86,87][50][51]. An approximately 150 kb plasmid was observed in an endophytic plant growth-promoting Enterobacter sp. (pENT638-1), which has a role in host colonization [85][49]. HGT is a common strategy for changing adaptation-related genes, such as those related to antibiotic resistance and heavy metal resistance among soil bacteria. Plant growth-promoting rhizobacteria possess more than one beneficial function as a result of gene accumulation in the rhizosphere and soil environment governed by selection mechanisms. The major function genes related to plant-beneficial function are as follows: (i). nitrogen fixation-contributing nifHDK genes (encoding nitrogenase), (ii). Mineral phosphate solubilization pqqBCDEFG genes (encoding pyrroloquinoline quinone), (iii). inhibition of ethylene biosynthesis acdS gene (encoding 1-aminocyclopropane-1-carboxylate), (iv). IAA-producing ipdC/ppdC genes (encoding indole-3-pyruvate decarboxylase/phenylpyruvate decarboxylase), (v). antimicrobial compound synthesis hcnABC/phlACBD (hydrogen cyanide/2,4-diacetylphloroglucinol) genes, and (vi). induced systemic resistance conferring budAB/budC (acetoine/2,3-butanediol) genes.2.5. Synergistic Microbial Processes
In many cases, plant inoculation with bacterial consortia proved to be more efficient than inoculation with a single bacterial strain. Synergistic processes between ACC deaminase and IAA production and N2 fixation [89[52][53][54][55][56][57][58][59],90,91,92,93,94,95,96], ACC deaminase and IAA production and stress tolerance [93[56][60],97], and phosphate solubilization and ACC deaminase [98][61] were reported. The role of ACC deaminase and IAA in BNF fixation process is complex; they can enhance nodule formation, improve the competitiveness of rhizobia for nodulation, suppress nodule senescence, and upregulate genes associated with legume–rhizobia symbiosis [96][59]. The role of ACC deaminase in nodule formation was studied using either knockout or overproducing strains for the ACC deaminase-producing gene [96][59]. When Mesorhizobium loti ACC deaminase-overproducing mutant strain was tested for the efficiency of plant colonization and nodulation, it was found to be more efficient than the wild type [89][52]. This relationship is relatively complex, whereas in Mesorhizobium loti, the acdS gene was found in the symbiotic island and its expression was regulated by the N2 fixation regulator NifA2 [99][62]. In senescent nodules, increased gene expression of PsACS2 encoding ACC synthase (an enzyme involved in ethylene synthesis) and increased transcription have been observed [90][53]. Therefore, ACC deaminase-producing PGP bacteria can enhance N2 fixation by extending the lifespan of functional nodules.2.6. Innovations in Carrier Materials for Bioinoculants
Carrier materials for bioinoculants must be chemically stable, nontoxic, low-cost, and able to provide a protective niche for microorganisms to ensure the viability of cells during storage and controlled release [101,102][63][64]. Many types of bioinoculant carriers have been studied in the recent decades. They can be classified as solid, liquid, organic, or inorganic. Additives that nutritionally support microorganisms are used in these bioformulations [101,102][63][64]. Peat, biochar, bagasse, cork compost, attapulgite, sepiolite, perlite, and amorphous silica were used as media for the solid bioformulations. They provide support for beneficial microbes, in contrast with liquid bioformulations that are more sensitive to prolonged storage. Immobilized formulations or encapsulation is an emerging technology with significant advantages over the above-mentioned formulations [101,102][63][64]. Microbial cells are immobilized by adhesion/biofilm formation on solid supports or entrapment, thereby conferring a protective environment for bacterial cells [103][65]. The use of environmentally friendly biopolymer matrices is well suited to sustainable agriculture. Microbial cells are encapsulated using various techniques such as ionic gelation, emulsification, and spray drying [102][64]. Additives are used to improve the stability, encapsulation efficiency, and mechanical properties of the carrier polymer, as well as fillers to improve microbial survival [106,108][66][67]. Alginates are the most widely studied microbial carriers, mainly for Azospirillum sp. and Pseudomonas sp. [102][64]. Alginate bead-entrapped A. brasilense showed better viability during prolonged storage [104][68]. Calcium alginate microspheres have been used for Trichoderma viride spore encapsulation and provided a supportive environment for growth [109][69].2.7. Engineering Microbiome
Many plant growth-promoting microbes and microbial consortia have been studied and proposed as potential bioinoculants. Various carriers have been tested to maximize their colonization and persistence in harsh soil environments. Nevertheless, limitations of natural bioinoculant use have been reported due to the complexity of soil–microbe–plant interconnectedness. A better understanding of the rhizosphere biochemical and molecular specificity that governs plant–microbe interactions is required to be used in rhizosphere microbiome engineering [112][70]. Rhizosphere microbiome engineering has gained much attention in advanced agricultural research [113,114][71][72]. Microbiome engineering uses a microbe-focused approach that is based on constructing synthetic communities called SynComs. These communities can be constructed using bottom-up strategies. The bottom-up approach involves the identification of keystone microbial taxa (e.g., Agrobacterium, Pseudomonas, Enterobacter) and the use of a combination of microbial isolates. SynComs complexity is important in terms of their effectiveness and stability in a changing environment, and functional species can be substituted because of their stable metabolic network [113][71].References
- De Corato, U. Towards New Soil Management Strategies for Improving Soil Quality and Ecosystem Services in Sustainable Agriculture: Editorial Overview. Sustainability 2020, 12, 9398.
- Bar-On, Y.M.; Phillips, R.; Milo, R. The biomass distribution on Earth. Proc. Natl. Acad. Sci. USA 2018, 115, 6506–6511.
- Dubey, D.-A.; Malla, M.; Khan, F.; Chowdhary, K.; Yadav, S.; Kumar, A.; Sharma, S.; Khare, P.K.; Khan, M.; Khan, M. Soil microbiome: A key player for conservation of soil health under changing climate. Biodivers. Conserv. 2019, 28, 2405–2429.
- Mendes, R.; Garbeva, P.; Raaijmakers, J.M. The rhizosphere microbiome: Significance of plant beneficial, plant pathogenic, and human pathogenic microorganisms. FEMS Microbiol. Rev. 2013, 37, 634–663.
- O’Brien, S.L.; Gibbons, S.M.; Owens, S.M.; Hampton-Marcell, J.; Johnston, E.R.; Jastrow, J.D.; Gilbert, J.A.; Meyer, F.; Antonopoulos, D.A. Spatial scale drives patterns in soil bacterial diversity. Environ. Microbiol. 2016, 18, 2039–2051.
- Jansson, J.K.; Hofmockel, K.S. Soil microbiomes and climate change. Nat. Rev. Microbiol. 2020, 18, 35–46.
- Thakur, M.P.; Geisen, S. Trophic Regulations of the Soil Microbiome. Trends Microbiol. 2019, 27, 771–780.
- Fan, K.; Cardona, C.; Li, Y.; Shi, Y.; Xiang, X.; Shen, C.; Wang, H.; Jack, G.; Chu, H. Rhizosphere-associated bacterial network structure and spatial distribution differ significantly from bulk soil in wheat crop fields. Soil Biol. Biochem. 2017, 113, 275–284.
- Bakker, P.; Berendsen, R.; Doornbos, R.; Wintermans, P.; Pieterse, C. The rhizosphere revisited: Root microbiomics. Front. Plant Sci. 2013, 4, 165.
- Saleem, M.; Hu, J.; Jousset, A. More than the Sum of Its Parts: Microbiome Biodiversity as a Driver of Plant Growth and Soil Health. Annu. Rev. Ecol. Evol. Syst. 2019, 50, 145–168.
- Ettema, C.H.; Wardle, D.A. Spatial soil ecology. Trends Ecol. Evol. 2002, 17, 177–183.
- Bakker, M.G.; Chaparro, J.M.; Manter, D.K.; Vivanco, J.M. Impacts of bulk soil microbial community structure on rhizosphere microbiomes of Zea mays. Plant Soil 2015, 392, 115–126.
- Li, X.; Rui, J.; Mao, Y.; Yannarell, A.; Mackie, R. Dynamics of the bacterial community structure in the rhizosphere of a maize cultivar. Soil Biol. Biochem. 2014, 68, 392–401.
- Yadav, A.N.; Kumar, V.; Dhaliwal, H.; Prasad, R.; Saxena, A. Microbiome in Crops: Diversity, Distribution, and Potential Role in Crop Improvement. In New and Future Developments in Microbial Biotechnology and Bioengineering: Crop Improvement through Microbial Biotechnology; Elsevier: Amsterdam, The Netherlands, 2018; pp. 305–332. ISBN 978-0-444-63987-5.
- Mahoney, A.K.; Yin, C.; Hulbert, S.H. Community Structure, Species Variation, and Potential Functions of Rhizosphere-Associated Bacteria of Different Winter Wheat (Triticum aestivum) Cultivars. Front. Plant Sci. 2017, 8, 132.
- Rathore, R.; Dowling, D.N.; Forristal, P.D.; Spink, J.; Cotter, P.D.; Bulgarelli, D.; Germaine, K.J. Crop Establishment Practices Are a Driver of the Plant Microbiota in Winter Oilseed Rape (Brassica napus). Front. Microbiol. 2017, 8, 1489.
- Ullah, A.; Akbar, A.; Luo, Q.; Khan, A.H.; Manghwar, H.; Shaban, M.; Yang, X. Microbiome Diversity in Cotton Rhizosphere Under Normal and Drought Conditions. Microb. Ecol. 2019, 77, 429–439.
- Edwards, J.; Johnson, C.; Santos-Medellín, C.; Lurie, E.; Podishetty, N.K.; Bhatnagar, S.; Eisen, J.A.; Sundaresan, V. Structure, variation, and assembly of the root-associated microbiomes of rice. Proc. Natl. Acad. Sci. USA 2015, 112, E911–E920.
- Sugiyama, A.; Ueda, Y.; Zushi, T.; Takase, H.; Yazaki, K. Changes in the Bacterial Community of Soybean Rhizospheres during Growth in the Field. PLoS ONE 2014, 9, e100709.
- Praeg, N.; Pauli, H.; Illmer, P. Microbial Diversity in Bulk and Rhizosphere Soil of Ranunculus glacialis Along a High-Alpine Altitudinal Gradient. Front. Microbiol. 2019, 10, 1429. Available online: https://www.frontiersin.org/articles/10.3389/fmicb.2019.01429 (accessed on 24 August 2023).
- Lakshmanan, V.; Selvaraj, G.; Bais, H.P. Functional soil microbiome: Belowground solutions to an aboveground problem. Plant Physiol. 2014, 166, 689–700.
- Aira, M.; Gómez-Brandón, M.; Lazcano, C.; Bååth, E. Plant genotype strongly modifies the structure and growth of maize rhizosphere microbial communities. Soil Biol. Biochem. 2010, 42, 2276–2281.
- Geller, A.M.; Levy, A. “What I cannot create, I do not understand”: Elucidating microbe-microbe interactions to facilitate plant microbiome engineering. Curr. Opin. Microbiol. 2023, 72, 102283.
- Khan, A.A.H. Plant-Bacterial Association and Their Role as Growth Promoters and Biocontrol Agents. In Plant Growth Promoting Rhizobacteria for Sustainable Stress Management; Springer: Cham, Switzerland, 2019; pp. 389–419. ISBN 9789811369858.
- Eva, L.; Gyongyver, M. Is PGPR an Alternative for NPK Fertilizers in Sustainable Agriculture? In Microbial Interventions in Agriculture and Environment; Springer: Cham, Switzerland, 2019; pp. 51–62. ISBN 9789811383908.
- Turan, M.; Topcuoğlu, B.; Kıtır, N.; Alkaya, Ü.; Erçelik, F.; Nikerel, E.; Güneş, A.; Turan, M.; Topcuoğlu, B.; Kıtır, N.; et al. Plant Growth Promoting Rhizobacteria’s (PGPRS) Enzyme Dynamics in Soil Remediation. In Soil Contamination—Current Consequences and Further Solutions; IntechOpen: London, UK, 2016; ISBN 978-953-51-2816-8.
- Fowler, D.; Steadman, C.E.; Stevenson, D.; Coyle, M.; Rees, R.M.; Skiba, U.M.; Sutton, M.A.; Cape, J.N.; Dore, A.J.; Vieno, M.; et al. Effects of global change during the 21st century on the nitrogen cycle. Atmos. Chem. Phys. 2015, 15, 13849–13893.
- Ladha, J.K.; Peoples, M.B.; Reddy, P.M.; Biswas, J.C.; Bennett, A.; Jat, M.L.; Krupnik, T.J. Biological nitrogen fixation and prospects for ecological intensification in cereal-based cropping systems. Field Crops Res. 2022, 283, 108541.
- Matilla, M.; Krell, T. Plant Growth Promotion and Biocontrol Mediated by Plant-Associated Bacteria. In Plant Microbiome: Stress Response; Springer: Cham, Switzerland, 2018; pp. 45–80. ISBN 978-981-10-5513-3.
- Ahmad, M.; Nadeem, S.M.; Zahir, Z.A. Plant-Microbiome Interactions in Agroecosystem: An Application. In Microbiome in Plant Health and Disease; Springer: Singapore, 2019; pp. 251–291.
- Salazar-Cerezo, S.; Martínez-Montiel, N.; García-Sánchez, J.; Pérez-Y-Terrón, R.; Martínez-Contreras, R.D. Gibberellin biosynthesis and metabolism: A convergent route for plants, fungi and bacteria. Microbiol. Res. 2018, 208, 85–98.
- de Garcia Salamone, I.E.; Hynes, R.K.; Nelson, L.M. Role of Cytokinins in Plant Growth Promotion by Rhizosphere Bacteria. In PGPR: Biocontrol and Biofertilization; Siddiqui, Z.A., Ed.; Springer: Dordrecht, The Netherlands, 2006; pp. 173–195. ISBN 978-1-4020-4152-5.
- Ramadan, E.; Abdelhafez, A.; Enas, A.; Saber, F. Plant growth promoting rhizobacteria and their potential for biocontrol of phytopathogens. Afr. J. Microbiol. Res. 2016, 10, 486–504.
- Zhang, S.; Gan, Y.; Xu, B. Mechanisms of the IAA and ACC-deaminase producing strain of Trichoderma longibrachiatum T6 in enhancing wheat seedling tolerance to NaCl stress. BMC Plant Biol. 2019, 19, 22.
- Sakure, S.; Bhosale, S. Actinobacteria for Biotic Stress Management. In Plant Growth Promoting Rhizobacteria for Sustainable Stress Management; Springer: Cham, Switzerland, 2019; pp. 363–378. ISBN 9789811369858.
- Mathur, A.; Koul, A.; Hattewar, J. Plant Growth-Promoting Rhizobacteria (PGPRs): Significant Revolutionary Tools for Achieving Long-Term Sustainability and Combating the Biotic Stress Caused by the Attack of Pathogens Affecting Crops in Agriculture. In Plant Growth Promoting Rhizobacteria for Sustainable Stress Management; Springer: Cham, Switzerland, 2019; pp. 379–388. ISBN 9789811369858.
- Tandon, S.; Vats, S. Microbial Biosynthesis of Cadmium Sulfide (CDS) Nanoparticles and their Characterization. Eur. J. Pharm. Med. Res. 2016, 3, 545–550.
- Vats, S.; Bhargava, P. Alternate Energy: Fuel for “Modi’s India” and “Smart cities”. Int. J. Curr. Res. 2017, 9, 49090–49097.
- Saxena, P.; Srivastava, J.; Pandey, S.; Srivastava, S.; Maurya, N.; Chand, K.; Mishra, S.; Asthana, G.; Bhargava, P.; Kumar, R.; et al. Plants for Biocontrol and Biological Control of Plant Pathogens. In Plant Biotic Interactions; Springer: Cham, Switzerland, 2019; pp. 147–179. ISBN 978-3-030-26656-1.
- Arseneault, T.; Filion, M. Biocontrol through antibiosis: Exploring the role played by subinhibitory concentrations of antibiotics in soil and their impact on plant pathogens. Can. J. Plant Pathol. 2017, 39, 267–274.
- Ram, R.M.; Keswani, C.; Bisen, K.; Tripathi, R.; Singh, S.P.; Singh, H.B. Biocontrol Technology; Elsevier: Amsterdam, The Netherlands, 2018; pp. 177–190.
- Quiñones, B.; Dulla, G.; Lindow, S.E. Quorum sensing regulates exopolysaccharide production, motility, and virulence in Pseudomonas syringae. Mol. Plant Microbe Interact. 2005, 18, 682–693.
- Koutsoudis, M.D.; Tsaltas, D.; Minogue, T.D.; von Bodman, S.B. Quorum-sensing regulation governs bacterial adhesion, biofilm development, and host colonization in Pantoea stewartii subspecies stewartii. Proc. Natl. Acad. Sci. USA 2006, 103, 5983–5988.
- Hussain, M.B.B.M.; Zhang, H.-B.; Xu, J.-L.; Liu, Q.; Jiang, Z.; Zhang, L.-H. The acyl-homoserine lactone-type quorum-sensing system modulates cell motility and virulence of Erwinia chrysanthemi pv. zeae. J. Bacteriol. 2008, 190, 1045–1053.
- Kang, B.R.; Anderson, A.J.; Kim, Y.C. Hydrogen Cyanide Produced by Pseudomonas chlororaphis O6 Exhibits Nematicidal Activity against Meloidogyne hapla. Plant Pathol. J. 2018, 34, 35–43.
- Yu, Y.; Gui, Y.; Li, Z.; Jiang, C.; Guo, J.; Niu, D. Induced Systemic Resistance for Improving Plant Immunity by Beneficial Microbes. Plants 2022, 11, 386.
- Lekired, A.; Cherif-Silini, H.; Silini, A.; Ben Yahia, H.; Ouzari, H.-I. Comparative genomics reveals the acquisition of mobile genetic elements by the plant growth-promoting Pantoea eucrina OB49 in polluted environments. Genomics 2023, 115, 110579.
- Maheshwari, M.; Abulreesh, H.H.; Khan, M.S.; Ahmad, I.; Pichtel, J. Horizontal Gene Transfer in Soil and the Rhizosphere: Impact on Ecological Fitness of Bacteria. In Agriculturally Important Microbes for Sustainable Agriculture: Volume I: Plant-Soil-Microbe Nexus; Meena, V.S., Mishra, P.K., Bisht, J.K., Pattanayak, A., Eds.; Springer: Singapore, 2017; pp. 111–130. ISBN 978-981-10-5589-8.
- Sánchez-Salazar, A.M.; Taparia, T.; Olesen, A.K.; Acuña, J.J.; Sørensen, S.J.; Jorquera, M.A. An overview of plasmid transfer in the plant microbiome. Plasmid 2023, 127, 102695.
- Ku, Y.-S.; Wang, Z.; Duan, S.; Lam, H.-M. Rhizospheric Communication through Mobile Genetic Element Transfers for the Regulation of Microbe–Plant Interactions. Biology 2021, 10, 477.
- Popowska, M.; Krawczyk-Balska, A. Broad-host-range IncP-1 plasmids and their resistance potential. Front. Microbiol. 2013, 4, 44.
- Ma, W.; Charles, T.C.; Glick, B.R. Expression of an exogenous 1-aminocyclopropane-1-carboxylate deaminase gene in Sinorhizobium meliloti increases its ability to nodulate alfalfa. Appl. Environ. Microbiol. 2004, 70, 5891–5897.
- Serova, T.A.; Tikhonovich, I.A.; Tsyganov, V.E. Analysis of nodule senescence in pea (Pisum sativum L.) using laser microdissection, real-time PCR, and ACC immunolocalization. J. Plant Physiol. 2017, 212, 29–44.
- Camerini, S.; Senatore, B.; Lonardo, E.; Imperlini, E.; Bianco, C.; Moschetti, G.; Rotino, G.L.; Campion, B.; Defez, R. Introduction of a novel pathway for IAA biosynthesis to rhizobia alters vetch root nodule development. Arch. Microbiol. 2008, 190, 67–77.
- Defez, R.; Esposito, R.; Angelini, C.; Bianco, C. Overproduction of Indole-3-Acetic Acid in Free-Living Rhizobia Induces Transcriptional Changes Resembling Those Occurring in Nodule Bacteroids. Mol. Plant Microbe Interact. 2016, 29, 484–495.
- Defez, R.; Andreozzi, A.; Romano, S.; Pocsfalvi, G.; Fiume, I.; Esposito, R.; Angelini, C.; Bianco, C. Bacterial IAA-Delivery into Medicago Root Nodules Triggers a Balanced Stimulation of C and N Metabolism Leading to a Biomass Increase. Microorganisms 2019, 7, 403.
- Nascimento, F.X.; Brígido, C.; Glick, B.R.; Oliveira, S.; Alho, L. Mesorhizobium ciceri LMS-1 expressing an exogenous 1-aminocyclopropane-1-carboxylate (ACC) deaminase increases its nodulation abilities and chickpea plant resistance to soil constraints. Lett. Appl. Microbiol. 2012, 55, 15–21.
- Nascimento, F.X.; Tavares, M.J.; Franck, J.; Ali, S.; Glick, B.R.; Rossi, M.J. ACC deaminase plays a major role in Pseudomonas fluorescens YsS6 ability to promote the nodulation of Alpha- and Betaproteobacteria rhizobial strains. Arch. Microbiol. 2019, 201, 817–822.
- Alemneh, A.A.; Zhou, Y.; Ryder, M.H.; Denton, M.D. Mechanisms in plant growth-promoting rhizobacteria that enhance legume-rhizobial symbioses. J. Appl. Microbiol. 2020, 129, 1133–1156.
- Orozco-Mosqueda, M.d.C.; Glick, B.R.; Santoyo, G. ACC deaminase in plant growth-promoting bacteria (PGPB): An efficient mechanism to counter salt stress in crops. Microbiol. Res. 2020, 235, 126439.
- Alemneh, A.A.; Zhou, Y.; Ryder, M.H.; Denton, M.D. Is phosphate solubilizing ability in plant growth-promoting rhizobacteria isolated from chickpea linked to their ability to produce ACC deaminase? J. Appl. Microbiol. 2021, 131, 2416–2432.
- Nukui, N.; Minamisawa, K.; Ayabe, S.-I.; Aoki, T. Expression of the 1-aminocyclopropane-1-carboxylic acid deaminase gene requires symbiotic nitrogen-fixing regulator gene nifA2 in Mesorhizobium loti MAFF303099. Appl. Environ. Microbiol. 2006, 72, 4964–4969.
- Balla, A.; Silini, A.; Cherif-Silini, H.; Chenari Bouket, A.; Alenezi, F.N.; Belbahri, L. Recent Advances in Encapsulation Techniques of Plant Growth-Promoting Microorganisms and Their Prospects in the Sustainable Agriculture. Appl. Sci. 2022, 12, 9020.
- Pereira, J.F.; Oliveira, A.L.M.; Sartori, D.; Yamashita, F.; Mali, S. Perspectives on the Use of Biopolymeric Matrices as Carriers for Plant-Growth Promoting Bacteria in Agricultural Systems. Microorganisms 2023, 11, 467.
- Lobo, C.B.; Juárez Tomás, M.S.; Viruel, E.; Ferrero, M.A.; Lucca, M.E. Development of low-cost formulations of plant growth-promoting bacteria to be used as inoculants in beneficial agricultural technologies. Microbiol. Res. 2019, 219, 12–25.
- Marcelino, P.R.F.; Milani, K.M.L.; Mali, S.; Santos, O.J.A.P.D.; de Oliveira, A.L.M. Formulations of polymeric biodegradable low-cost foam by melt extrusion to deliver plant growth-promoting bacteria in agricultural systems. Appl. Microbiol. Biotechnol. 2016, 100, 7323–7338.
- Panichikkal, J.; Prathap, G.; Nair, R.A.; Krishnankutty, R.E. Evaluation of plant probiotic performance of Pseudomonas sp. encapsulated in alginate supplemented with salicylic acid and zinc oxide nanoparticles. Int. J. Biol. Macromol. 2021, 166, 138–143.
- Zago, S.; Fonseca dos Santos, M.; Konrad, D.; Fiorini, A.; Rosado, F.; Missio, R.; Vendruscolo, E. Shelf Life of Azospirillum brasilense in Alginate Beads Enriched with Trehalose and Humic Acid. J. Agric. Sci. 2019, 11, 269.
- Jurić, S.; Đermić, E.; Topolovec-Pintarić, S.; Bedek, M.; Vinceković, M. Physicochemical properties and release characteristics of calcium alginate microspheres loaded with Trichoderma viride spores. J. Integr. Agric. 2019, 18, 2534–2548.
- Kumar, A.; Dubey, A. Rhizosphere microbiome: Engineering bacterial competitiveness for enhancing crop production. J. Adv. Res. 2020, 24, 337–352.
- Bano, S.; Wu, X.; Zhang, X. Towards sustainable agriculture: Rhizosphere microbiome engineering. Appl. Microbiol. Biotechnol. 2021, 105, 7141–7160.
- Haskett, T.L.; Tkacz, A.; Poole, P.S. Engineering rhizobacteria for sustainable agriculture. ISME J. 2021, 15, 949–964.
