Your browser does not fully support modern features. Please upgrade for a smoother experience.
Please note this is a comparison between Version 1 by Diego Ruano and Version 2 by Jessie Wu.
SARS-CoV-2’s structure and mechanism of infection have been well characterized. The virus comprises a lipid envelope studded with spike (S) proteins. These spikes facilitate viral entry into host cells by binding to angiotensin-converting enzyme 2 (ACE2) receptors on the cell surface. Following attachment, the virus enters the cell by endocytosis. Its genetic material consists of a single-stranded RNA molecule, which encodes structural proteins, non-structural proteins (NSP), and accessory proteins. Once inside, the viral RNA is translated into proteins, including those for replication and the formation of new virus particles.
- COVID-19
- SARS-CoV-2
- protein quality control systems
- ER stress
- neurodegeneration
1. Endoplasmic Reticulum Stress and Unfolded Protein PResponse
Endoplasmic reticulum (ER) stress and unfolded protein response (UPR)UPR are common aspects that emerge during the appearance and progression of both neurodegenerative diseases [1][2][3][14,15,16] and viral infections [4][5][13,17]. Viral infections often trigger an upsurge in protein synthesis, potentially surpassing the folding capacity of the ER. Consequently, this imbalance leads to the accumulation of unfolded proteins, inducing ER stress [4][5][13,17]. In response to ER stress, cells activate a multifaceted signaling network known as UPR, an adaptive response aimed at mitigating the burden of unfolded proteins to sustain cellular viability and function [6][7][18,19].
The UPR is a sophisticated signaling pathway initiated by the activation of three primary UPR stress sensors (see Figure 1): inositol-requiring protein 1 (IRE1), protein kinase RNA-like ER kinase (PERK), and activating transcription factor 6 (ATF6) [4][6][8][9][10][13,18,20,21,22]. The glucose regulated protein-78 (GRP78, also known as BiP or HSPA5), is an ER chaperone that plays a pivotal role as the principal regulatory protein by binding to these sensors. In the absence of stress, GRP78 predominantly associates with these three proteins, effectively suppressing their activity and preventing the initiation of UPR signaling. However, under stress conditions characterized by the accumulation of misfolded proteins in the ER (as observed in neurodegenerative diseases and in viral infections), GRP78 interacts with these unfolded proteins, aiming to maintain their foldable state and resulting in the release of the three UPR mediators (Figure 1). The specific mechanisms of the UPR pathways include [8][9][20,21]:
- −
-
IRE1 activation and splicing of XBP1 (X-box binding protein 1) mRNA, resulting in the production of sXBP1, an active transcription factor. sXBP1 regulates the expression of chaperones and ERAD components, reinforcing ER’s protein-folding and -degradation capacity. IRE1 also catalyzes the degradation of a large number of mRNAs and some pre-microRNAs (pre-miRNAs). This process is called regulated IRE1-dependent decay (RIDD) [11][12][23,24]. On the other hand, it is known that IRE1 is capable of forming high-order complexes in the ER membrane and interacting with a large number of proteins, among which the tumor necrosis factor (TNF) receptor-associated factor 2 (TRAF2) stands out. This interaction activates a cascade of signaling that leads to the activation of c-Jun N-terminal kinase (JNK), which, in turn, can inhibit some anti-apoptotic members of the BCL-2 family while activating pro-apoptotic proteins. Together, these two events lead to the oligomerization of BCL-2-like protein 4 (BAX) and BCL-2 antagonist/killer (BAK), initiating the apoptosis process [11][12][13][23,24,25];
- −
-
PERK activation and the phosphorylation of the eukaryotic initiation factor 2α (eIF2α), a pivotal regulator of protein translation. Phosphorylated eIF2α reduces global protein synthesis, thereby alleviating the ER burden and allowing cells to cope with ER stress. However, this process can also induce apoptosis by upregulating the C/EBP homologous protein (CHOP). The prolonged upregulation of CHOP induces apoptosis through pathways involving the BCL2 binding component 3 (BBC3/PUMA) and the tribbles pseudokinase 3 (TRIB3).
- −
-
ATF6 activation and translocation to the Golgi apparatus, where it is cleaved by site-1 (S1P) and site-2 (S2P) proteases, releasing a 50 kDa N-terminal fragment that translocates to the nucleus. This fragment acts as a transcription factor that subsequently upregulates genes encoding ER chaperones and other proteins involved in ER quality control.
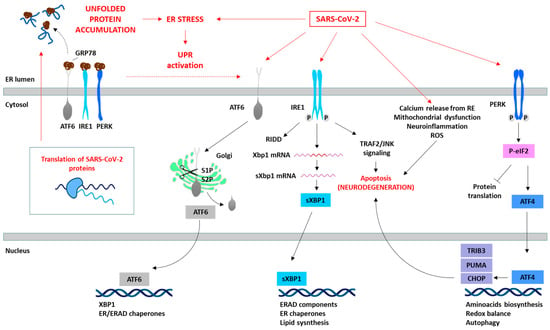
Figure 1. SARS-CoV-2 induces ER stress and activates the UPR. The translation of SARS-CoV-2 proteins generates ER stress, which activates the UPR as an antiviral host cell response. However, SARS-CoV-2 is able to manipulate the different pathways of the UPR (red arrows) depending on the viral requirements for its replication. The specific proteins involved in the SARS-CoV-2 and UPR interaction are summarized in Table 1. The prolonged activation of the UPR leads to the initiation of pro-apoptotic signaling pathways (e.g., IRE1/TRAF2/JNK or PERK/ATF4/CHOP), disturbances in cellular calcium levels due to ER release, mitochondrial dysfunction, and consequently ROS generation, and the induction of oxidative stress, culminating in cell death.
These sensors orchestrate adaptive processes through both transcriptional and non-transcriptional responses to restore ER homeostasis (Figure 1). This involves adapting protein synthesis, enhancing protein folding capacity, and increasing the efficiency of ERAD [8][9][10][20,21,22]. The activation of these three UPR pathways leads to diverse downstream consequences, depending on the nature and intensity of the stimuli and the specific cell type involved. Although the primary objective of the UPR is to reinstate ER homeostasis, prolonged or severe ER stress can activate apoptotic pathways through factors such as activating transcription factor 4 (ATF4) and CHOP, ultimately culminating in cell death, as observed in neurodegenerative processes [4][8][9][10][13,20,21,22].
2. Endoplasmic Reticulum-Associated DegradationRAD and Ubiquitin-Proteasome PSystem
ERAD is a crucial process in the quality control of ER proteins, facilitating the elimination of aberrant proteins by the UPS [14][49]. This process is essential for maintaining cellular proteostasis, as misfolded or unassembled proteins can compromise the cell function. This process consists of three phases: recognition of misfolded or damaged proteins in the ER, retrotranslocation into the cytosol, and ubiquitin-dependent degradation by the proteasome (Figure 2). ER luminal chaperones both prevent improper folding and identify terminally misfolded proteins, directing them for ERAD. Some ERAD components include the EDEM family members, such as EDEM1, EDEM2, and EDEM3, as well as the Osteosarcoma 9 (OS9) protein, the chaperone GRP78, and protein disulfide-isomerases (PDIs), such as the ER DnaJ domain-containing protein 5 (ERdj5). These components facilitate the recognition, recruitment and disposal of the misfolded proteins [15][16][50,51] for retrotranslocation via the ERAD complex. This complex involves the Sec61 translocon, DERL1-3 and valosin-containing protein (p97), together with the suppressor enhancer Lin12 1-like (SEL1L) and the ERAD-associated E3 ubiquitin–protein ligase (HRD1) that contribute to the ubiquitination process [16][17][51,52]. After (poly)ubiquitination, the proteins are recognized and degraded by the proteasome (Figure 1). The ubiquitination process (which can take place in the ER or in the cytosol) involves the covalent attachment of ubiquitin molecules to lysine (K) residues on the substrate protein, through an enzymatic cascade involving E1 (ubiquitin-activating), E2 (ubiquitin-conjugating), and E3 (ubiquitin ligase) enzymes. The specificity of ubiquitination is determined by the type of linkage formed between ubiquitin molecules [18][53]. Hence, the E3 ubiquitin ligases play a pivotal role in specifying the target proteins for degradation. Their activity, expression, and turnover are rigorously regulated to prevent inappropriate ubiquitination and maintain cellular function [14][49]. The type of ubiquitination will direct the protein to a specific destination. For example, K48 linkages are tagged to UPS degradation, while K63 is tagged to autophagy and non-proteolytic signaling pathways [18][19][53,54]. The proteasome, a large multi-catalytic protease, is responsible for the degradation of ERAD substrates, as well as other UPS-tagged proteins [20][55]. The proteasome consists of a catalytic core complex (20S proteasome) and regulatory subunits such as 19S or 11S particles. The catalytic core is the 20S proteasome, a hollow barrel-shaped structure comprising four rings with α and β subunits. The α-rings control substrate access, and the β-rings house catalytic subunits (β1, β2, and β5). This 20S proteasome degrades non-ubiquitinated misfolded or damaged proteins [4][13]. Additionally, the proteasome can associate with regulatory subunits, forming different types of proteasomes (mainly 26S or 30S, immunoproteasome, hybrid proteasomes) with distinct functions. The regulatory particle 19S forms the 26S or 30S proteasome when associated with the 20S proteasome. The 19S particle facilitates binding, deubiquitination, unfolding, and the channeling of target proteins for degradation [4][20][13,55]. Additionally, deubiquitinating enzymes (DUBs) can modulate ubiquitination and the ERAD process by removing ubiquitin chains from substrate proteins [21][56]. Besides this, cytokines such as interferon α (IFNα), IFNγ and TNFα induce the replacement of constitutive catalytic subunits (β1, β2, and β5) in the 20S proteasome with inducible subunits β1i, β2i, and β5i, and this results in the formation of the immunoproteasome (Figure 2). The immunoproteasome, expressed in immune cells, exhibits distinct proteolytic activities compared to the standard 20S proteasome. It plays essential roles in antigen presentation, γ-interferon-mediated microglial activation, cytokine production by microglial cells, and the regulation of T-cell populations, highlighting its significance in immune responses and cellular regulation [22][57].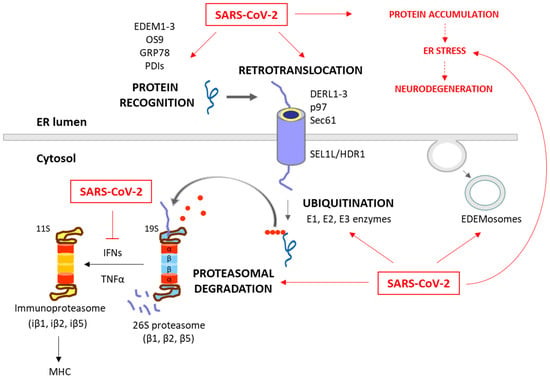
Figure 2. Degradation of unfolded protein through the ERAD/UPS pathways and SARS-CoV-2 interplay. When chaperones are not able to achieve the proper folding of proteins within the ER, these misfolded proteins are recognized and isolated by lectins and ERAD components, including EDEM1-3, OS9, PDIs and GRP78. The targeted proteins are then recruited to the ERAD complex and retrotranslocated to the cytosol. This complex involves Sec61, DERL1-3 and p97, together with SEL1L and HRD1, which assist in the ubiquitination process. After (poly)ubiquitination, the proteins are recognized by the 26S proteasome and deubiquitinated for degradation. While these processes are common antiviral responses of host cells, SARS-CoV-2 is able to positively or negatively manipulate this machinery to ensure its replication, escape and survival (red arrows). Consequently, the ERAD and UPS systems are not efficient in solving protein accumulation, leading to neurodegeneration. Moreover, SARS-CoV-2 can inhibit IFN production, and therefore the formation of immunoproteasome and antigen presentation.
The ERAD is also modulated by the ERAD tuning process, the regulation of ERAD activity by segregating ERAD components (like EDEM proteins) into specific ER-derived vesicles, named EDEMosomes (Figure 2). This segregation prevents the premature degradation of certain proteins, allowing the cell to regulate ERAD more precisely [23][24][58,59]. A huge number of associations between the ERAD pathway and human diseases have been established, highlighting the former’s significance in health and disease. Neurodegenerative diseases are characterized by disruptions in both ERAD and the UPS. These disruptions contribute to the accumulation of abnormal proteins and the formation of toxic protein aggregates, which are commonly associated with the pathogenesis of these conditions [25][26][27][60,61,62]. Also, beyond their fundamental roles in cell proteostasis, the ERAD and UPS have emerged as critical players in the intricate interaction between viruses and host cells during the process of viral infection [28][29][30][31][32][33][34][35][36][37][38][39][40][41][42][43][44][45][46][63,64,65,66,67,68,69,70,71,72,73,74,75,76,77,78,79,80,81].
