Your browser does not fully support modern features. Please upgrade for a smoother experience.
Please note this is a comparison between Version 1 by WenWei Li and Version 2 by Lindsay Dong.
L-Cysteine is a widely used unique sulfur-containing amino acid with wide application in the food, pharmaceutical, and agricultural industries.
- L-cysteine
- Escherichia coli
- metabolic engineering
- synthetic biology
1. Introduction
L-cysteine is a sulfur-containing amino acid that plays an important role in the folding of proteins, has a high redox activity in cellular metabolism, is a catalytic residue for a variety of enzymes, and is a sulfur donor compound that is required for the synthesis of Fe/S clusters, biotin, coenzyme A, and thiamine [1][2][3][1,2,3]. In addition to its roles in cellular metabolism, L-cysteine plays a variety of roles in metal binding, catalytic activity, and redox and has a vast array of industrial applications in the production of food, cosmetics, pharmaceuticals, and animal feed [4][5][6][4,5,6].
Chemical hydrolysis of proteins, which are typically extracted from the keratin of animal hair such as feathers, pig hair, etc., is the traditional method of industrial production of L-cysteine [7]. However, this method not only consumes a large amount of hydrochloric acid, but it also causes an unpleasant odor and wastewater treatment problems, which have a significant impact on the environment [7]. To avoid the environmental hazards of this method, scientists have explored biotechnological approaches to synthesize L-cysteine as an alternative to chemical hydrolysis. Fermentation and enzymatic biotransformation are the two most prominent biotechnological methods [8][9][8,9]. However, due to the presence of generated L-cysteine products that inhibit the activity of the enzymes, the enzyme bioconversion method presents difficulty in solving the problems of low yield and high cost.
Although fermentation offers a number of advantages, the design and construction of efficient microbial cell factories for fermentative production of L-cysteine remains challenging due to the high toxicity of L-cysteine and the complex regulation of its synthetic pathway [10]. The efficient production of L-cysteine on an industrial scale has not yet been achieved, which is a major challenge for the industrialization of L-cysteine [11][12][11,12]. Numerous microorganisms, including bacteria such as E. coli, C. glutamicum, and Pantoea ananatis, have been engineered to produce L-cysteine due to the rapid development of systems metabolic engineering and synthetic biology [13][14][15][16][17][13,14,15,16,17]. In comparison to other bacteria, E. coli has a rapid growth rate and more developed genetic engineering techniques, whereas C. glutamicum is a non-pathogenic, industrial microorganism with developed fermentation technology that is extensively employed in food processing and other industries [13][18][13,18]. Therefore, E. coli and C. glutamicum are the two most studied chassis cells that directly produce L-cysteine from glucose [19][20][21][19,20,21].
 Both L-ATC hydrolase (atcB) and S-carbamoyl-L-cysteine hydrolase (atcC) genes originated from Pseudomonas sp. strain BS. After sequencing by Japanese scientists, the amino acid sequence of the atcC gene product was found to be highly homologous to L-N-carbamoylase from other bacteria, but the amino acid sequence of the atcB gene was novel [8]. AtcB was initially identified as a gene encoding an enzyme that catalyzes the thiazoline ring-opening reaction and does not share a high degree of homology with previously described enzymes [26].
Enzymatic bioconversion is to some extent environmentally friendly and has lower energy consumption than hydrolysis of animal hair. However, the high toxicity of L-cysteine inhibits enzyme activity, leading to low efficiency and relatively high cost [8][9][8,9].
Both L-ATC hydrolase (atcB) and S-carbamoyl-L-cysteine hydrolase (atcC) genes originated from Pseudomonas sp. strain BS. After sequencing by Japanese scientists, the amino acid sequence of the atcC gene product was found to be highly homologous to L-N-carbamoylase from other bacteria, but the amino acid sequence of the atcB gene was novel [8]. AtcB was initially identified as a gene encoding an enzyme that catalyzes the thiazoline ring-opening reaction and does not share a high degree of homology with previously described enzymes [26].
Enzymatic bioconversion is to some extent environmentally friendly and has lower energy consumption than hydrolysis of animal hair. However, the high toxicity of L-cysteine inhibits enzyme activity, leading to low efficiency and relatively high cost [8][9][8,9].
2. Advances in the Biosynthesis of L-Cysteine
2.1. Enzyme Biotransformation—Asymmetrical Hydrolysis of DL-2-amino-Δ
2
-thiazoline-4- Carboxylic Acid
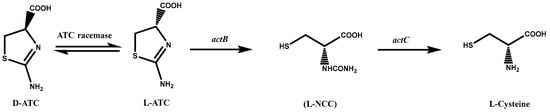
Since L-cysteine is traditionally obtained by hydrolyzing animal hair, the extraction of 1 kg of L-cysteine requires approximately 10 kg of animal hair and 2.7 kg of hydrochloric acid, a process that not only has a low yield but also produces foul odors and wastewater, causing severe environmental damage [22]. The transformation method uses Pseudomonas to enzymatically convert DL-2-amino-Δ2-thiazoline-4-carboxylic acid (DL-ATC) to L-cysteine. This method of converting DL-ATC to cysteine involves three enzymes: ATC racemase, L-ATC hydrolase, and S-carbamoyl-L-cysteine hydrolase [23][24][23,24]. The complete procedure consists of three stages (Figure 1): (i) conversion of D-ATC to L-ATC by ATC racemase, (ii) ring-opening of L-ATC by L-ATC hydrolase to produce N-carbamoyl-L-cysteine (L-NCC), and (iii) final hydrolysis of L-NCC to L-cysteine by S-carbamoyl-L-cysteine hydrolase [25].
Figure 1. A metabolic pathway of DL-2-amino-Δ2-thiazoline-4-carboxylic acid (DL-ATC) to L-cysteine via N-carbamyl-L-cysteine (L-NCC) in Pseudomonas species. atcB gene encoding L-ATC acid hydrolase; atcC gene encoding L-NCC amidohydrolase.
2.2. Biological Fermentation Methods
2.2.1. L-Cysteine Biosynthesis in E. coli
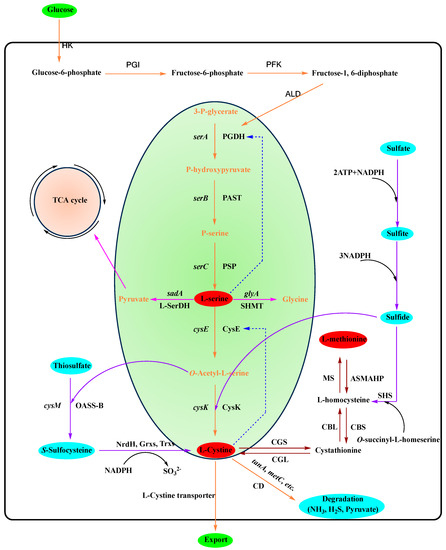
It is well known that in most microorganisms and plants, L-serine is the precursor substance for the synthesis of L-cysteine. The biosynthetic pathway of L-cysteine has been widely reported after many years of research [4][27][28][4,27,28]. In gut bacteria, L-serine is synthesized via a three-step pathway from the glycolytic intermediate 3-phosphoglycerate, and L-cysteine is synthesized via a two-step pathway from L-serine [28]. Firstly, the glycolytic intermediate 3-phosphoglycerate is converted to L-serine by a three-step reaction catalyzed by 3-phosphoglycerate dehydrogenase (PGDH), phosphoserine aminotransferase (PAST), and phosphoserine phosphatase (PSP) (Figure 2) [27].
Figure 2. The metabolic pathway of L-cysteine in E. coli and C. glutamicum. Brown arrows refer to metabolic pathways in P. aeruginosa. The solid purple line indicates the conversion of L-serine to other productsDashed lines represent feedback inhibition. The solid orange line indicates the major metabolic pathway of L-cysteine. The solid purple line indicates the source of the sulfide. Italicized fonts on the same arrow line indicate genes encoding corresponding enzymes. HK, hexokinase; PGI, phosphoglucose isomerase; PFK, phosphofructokinase; ALD, aldolase; PGDH, 3-phosphoglycerate dehydrogenase; PAST, phosphoserine aminotransferase; PSP, phosphoserine phosphatase; CysE, serine O-acetyltransferase; CysK, cysteine synthase; L-SerDH, L-serine dehydratase; SHMT, serine hydroxymethyl transferase; CD, L-cysteine desulfhydrases; OASS-B, O-acetyl-L-serine sulfhydrylase-A; NrdH and Grxs, glutaredoxins; Trxs, thioredoxins; CGS, cystathionine γ-synthase; CBL, cystathionine β-lyase; MS, methionine synthase; SHS, O-succinyl-L-homoserine sulfhydrylase; ASMAHP, S-adenosylmethionine synthase-methyltransferases-S-adenosylhomocysteine hydrolase pathway; CBS, cystathionine β-synthase; CGL, cystathionine γ-lyase.
Scientists have conducted studies to address the aforementioned causes of L-cysteine biosynthesis blockage. Table 12 summarizes the research progress on the fermentation synthesis of L-cysteine by different engineering strains.
Table 12.
Progress of fermentation synthesis of L-cysteine by different engineered strains.
| Bacterial Strain | Metabolic Strategy | L-Cysteine Production (g/L) | Productivity (g/(L·h)) | References |
|---|---|---|---|---|
| E. coli JM240 | Enhancing biosynthesis | 0.03 | / | [29][31] |
| E. coli JM39 | Enhancing biosynthesis | 0.20 | 0.003 | [30][32] |
| E. coli W3110 | Enhancing excretion | 0.07 | 0.003 | [31][33] |
| E. coli W3110 | Enhancing excretion | 0.15 | 0.007 | [32][34] |
| E. coli JM39 | Enhancing biosynthesis and weakening degradation | 0.60 | 0.013 | [33][35] |
| E. coli MG1655 | Enhancing biosynthesis and excretion and weakening degradation | 1.20 | 0.025 | [34][30] |
| E. coli BW25113 | Enhancing biosynthesis and excretion | 1.23 | 0.026 | [35][36] |
| E. coli BW25113 | Enhancing biosynthesis and excretion/weakening degradation | 1.72 | 0.024 | [20] |
| E. coli JM109 | Enhancing the sulfur conversion rate | 7.50 | 0.341 | [14] |
| E. coli BW25113 | Enhancing biosynthesis and thiosulfate assimilation and weakening degradation | 8.34 | 0.321 | [10] |
| E. coli W3110 | Balancing carbon and sulfur module conversion rate | 11.94 | 0.254 | [36][37] |
| C. glutamicum IR33 | Enhancing biosynthesis | 0.29 | 0.004 | [37][38] |
| C. glutamicum ATCC13032; C. glutamicum ATCC21586 |
Enhanced sulfur metabolism in biosynthesis | 0.06 | 0.004 | [27] |
| C. glutamicum NBRC12168 | Enhancing biosynthesis and weakening degradation | 0.20 | 0.017 | [16] |
| C. glutamicum CYS | Enhancing biosynthesis and excretion | 0.28 | 0.014 | [19] |
| C. glutamicum ATCC13032 | Enhancing precursor accumulation and weakening degradation | 0.95 | 0.026 | [15] |
| C. glutamicum Cys -10 |
Enhancing biosynthesis, excretion, and sulfur metabolism and weakening degradation | 5.92 | 0.082 | [38][39] |
| Pantoea ananatis | Weakening degradation and educing efflux | 2.20 | 0.079 | [17] |
2.2.2. L-Cysteine Biosynthesis in C. glutamicum
The metabolic pathway of L-cysteine in C. glutamicum is generally the same as that of E. coli, and L-serine is also synthesized from the glycolytic intermediate 3-phosphoglyceric acid via a three-step pathway, followed by further conversion to L-cysteine via CysE and CysK [39][43]. However, wild-type C. glutamicum is subjected to a feedback mechanism that produces almost no L-cysteine. In order to allow C. glutamicum to produce L-cysteine, the cysE gene (the gene is insensitive to feedback inhibition by L-cysteine) encoding an alteration of CysE in the E. coli Met256Ile mutant was introduced into C. glutamicum, which produces approximately 0.29 g/L of L-cysteine [37][38]. CysR is a transcriptional regulator that regulates sulfur metabolism in L-cysteine biosynthesis, and overexpression of the cysR gene in C. glutamicum resulted in a 2.7-fold higher intracellular sulfide concentration than that of the control strain (empty pMT-tac vector), and overexpression of the cysE, cysK, and cysR genes in C. glutamicum resulted in a 3-fold higher L-cysteine production than that of the control [27].
