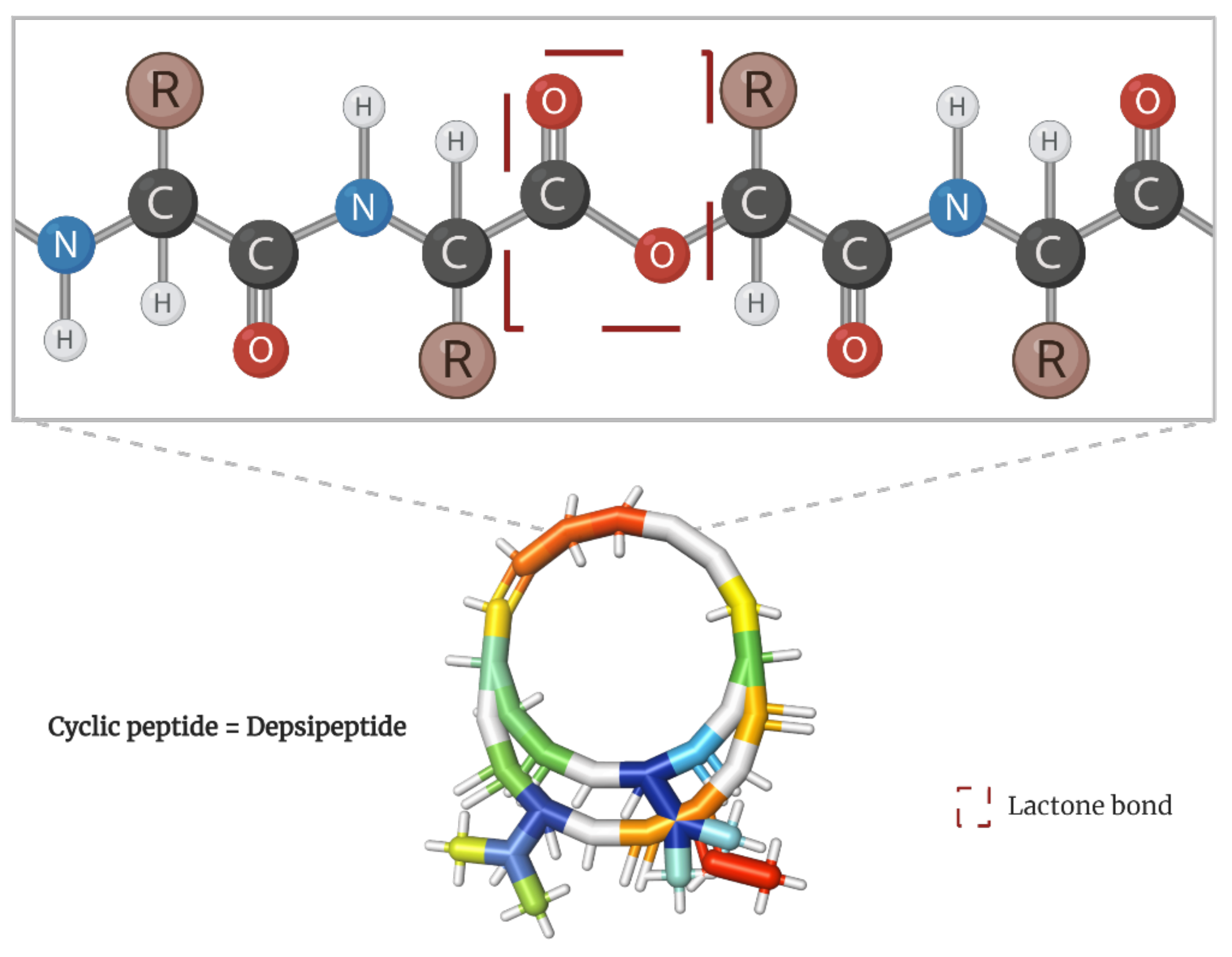
Cancer is currently considered one of the most threatening diseases worldwide. Diet could be one of the factors that can be enhanced to comprehensively address a cancer patient’s condition. Unfortunately, most molecules capable of targeting cancer cells are found in uncommon food sources. Among them, depsipeptides have emerged as one of the most reliable choices for cancer treatment. These cyclic amino acid oligomers, with one or more subunits replaced by a hydroxylated carboxylic acid resulting in one lactone bond in a core ring, have broadly proven their cancer-targeting efficacy, some even reaching clinical trials and being commercialized as “anticancer” drugs.
1. Introduction
Nowadays, cancer is considered the first or second major cause of death in multiple countries around the world
[1,2][1][2]. In 2020, more than 19 million new cases and nearly 10 million deaths associated with this disease were reported
[3,4,5][3][4][5]. Unsurprisingly, cancer cases are expected to reach the 24 million mark by 2035, and their incidence to triple by 2050
[6,7][6][7].
Although recognized cancer therapies are being further enhanced
[8], the consumption of enriched foods to boost the recovery of cancer patients is also gaining attention
[9]. To date, nearly 35% of chemotherapeutic drugs are nature-derived products, making these an important source of oncological pharmaceuticals
[10]. Nonetheless, only a limited number of recognized molecules found in foods, especially in animals, have been shown to possess promising cancer cell-targeting abilities in preclinical trials
[11].
Among the spectrum of potential therapeutic molecules, the tumor-targeting capability of natural cyclic peptides has been deeply explored
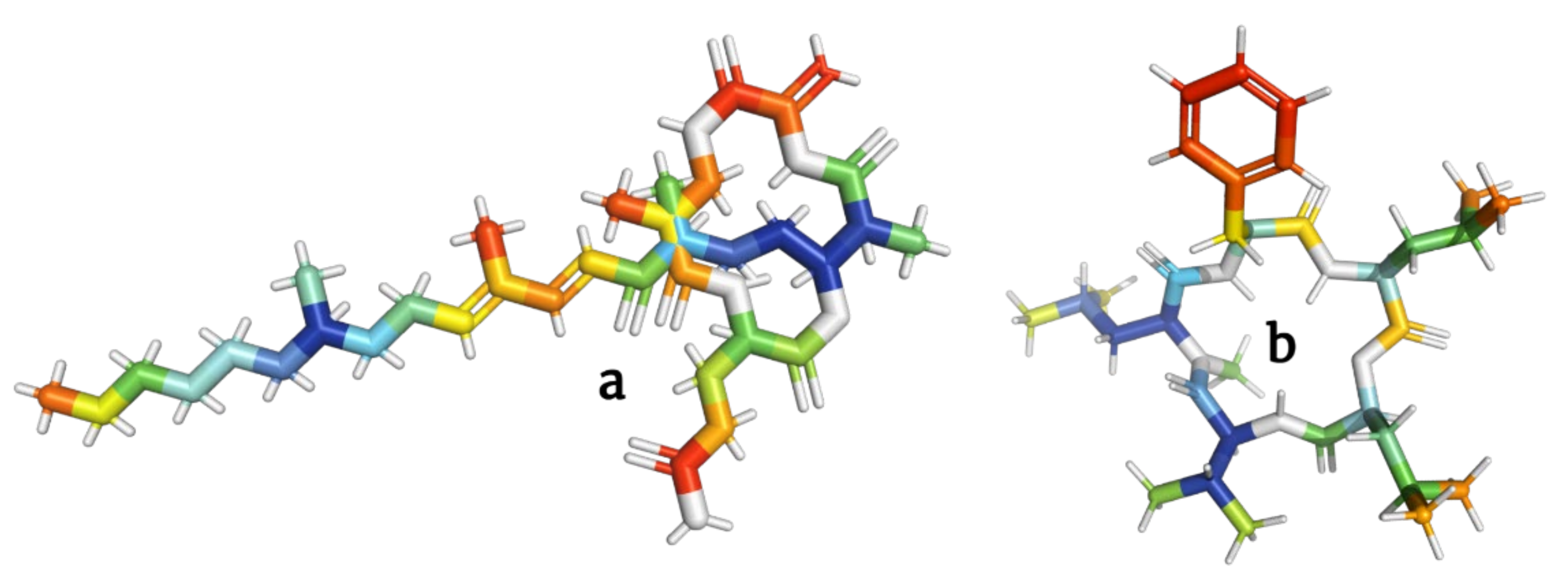
[12,13][12][13]. Scientifically known as depsipeptides, these peptide cyclo-oligomers are characterized by having one or more amino acids replaced by a hydroxylated carboxylic acid, resulting in one lactone bond in a core ring (
Figure 1)
[14]. Interestingly, they account for more than 1300 natural molecules reported to contain different acyl groups and other moieties
[15,16][15][16] as well as to perform varied bioactivities
[17,18][17][18].
Figure 1.
Peptide bond structure of depsipeptides. Lactone bond in the core ring marked in red.
Hitherto, few authors have recognized that the vast majority of depsipeptides are found in marine species, most of which are not present in easily accessible areas and are hard to reproduce in the laboratory environment without proper instruction
[10,19][10][19]. Some other authors have highlighted that the knowledge on “anticancer” mechanisms, known limitations, and faced challenges of using these molecules is minimal
[20].
2. Depsipeptides with a Recognized Mechanism of Targeting Cancer Cells
Depsipeptides induce cytotoxicity by various mechanisms, with their principal features being the capacity to perforate cell membranes and tubulin–microtubule imbalance, eventually inducing apoptosis
[21].
2.1. Depsipeptides Inducing Apoptosis in Tumor Cells
Apoptosis, as the most prevalent and well-defined form of programmed cell death
[22], plays an important role in regulating tumor cells
[23]. Unsurprisingly, resistance to apoptosis—due to the inactivation or loss of caspases
[24] and dysregulation of the mitochondrial pathway
[25]—is considered a major advantage for oncogenic cells
[26].
In general, these depsipeptides potently activate caspase-3, Fas/FasL, p21, and p27, and inhibit histone deacetylases, all of which result in pro-apoptotic signaling
[27,28,29,30][27][28][29][30] Here, the researchers present the state-of-the-art on depsipeptides that induce apoptosis in tumor cells (
Table 1 and
Figure 2).
Figure 2. Models of depsipeptides inducing apoptosis in tumor cells. (a) Apratoxin A; (b) Aurilides; (c) Beauvericins; (d) Dehydrodidemnin B; (e) Enniatins; (f) Lagunamides; (g) Coibamide A; (h) Grassypeptolides; (i) Hantupeptins; (j) Tiahuramides.
Table 1.
Formula and molecular weight of depsipeptides inducing apoptosis in tumor cells.
|
Key
|
Depsipeptide
|
IUPAC Condensed Formula
|
Molecular Weight
(g/mol)
|
Ref.
|
|
a
|
Apratoxin
|
Cyclo[N(Me)Ala-N(Me)Ile-Pro-Unk-Tyr(Me)]
|
840.1
|
[31]
|
|
b
|
Aurilide
|
Cyclo[N(Me)Ala-Unk-Val-D-N(Me)Leu-Sar-Val]
|
834.1
|
[21]
|
|
c
|
Beauvericin
|
Cyclo[D-Oval-N(Me)Phe-D-Oval-N(Me)Phe-D-Oval-N(Me)Phe]
|
783.9 |
[95][47].
2.3. Depsipeptides Inhibiting Elastases
Elastases are a group of proteases that degrade and lyse elastin, one of the main components of the extracellular matrix
[96][48]. These enzymes are also involved in tumor cell migration
[97][49]. Consequently, any dysregulation of elastase may result in the genesis of various diseases such as cancer
[98][50] as well as metastasis
[97][49].
Here, the researchers elaborate on the depsipeptides with elastase-inhibiting potential (
Table 3 and
Figure 4).
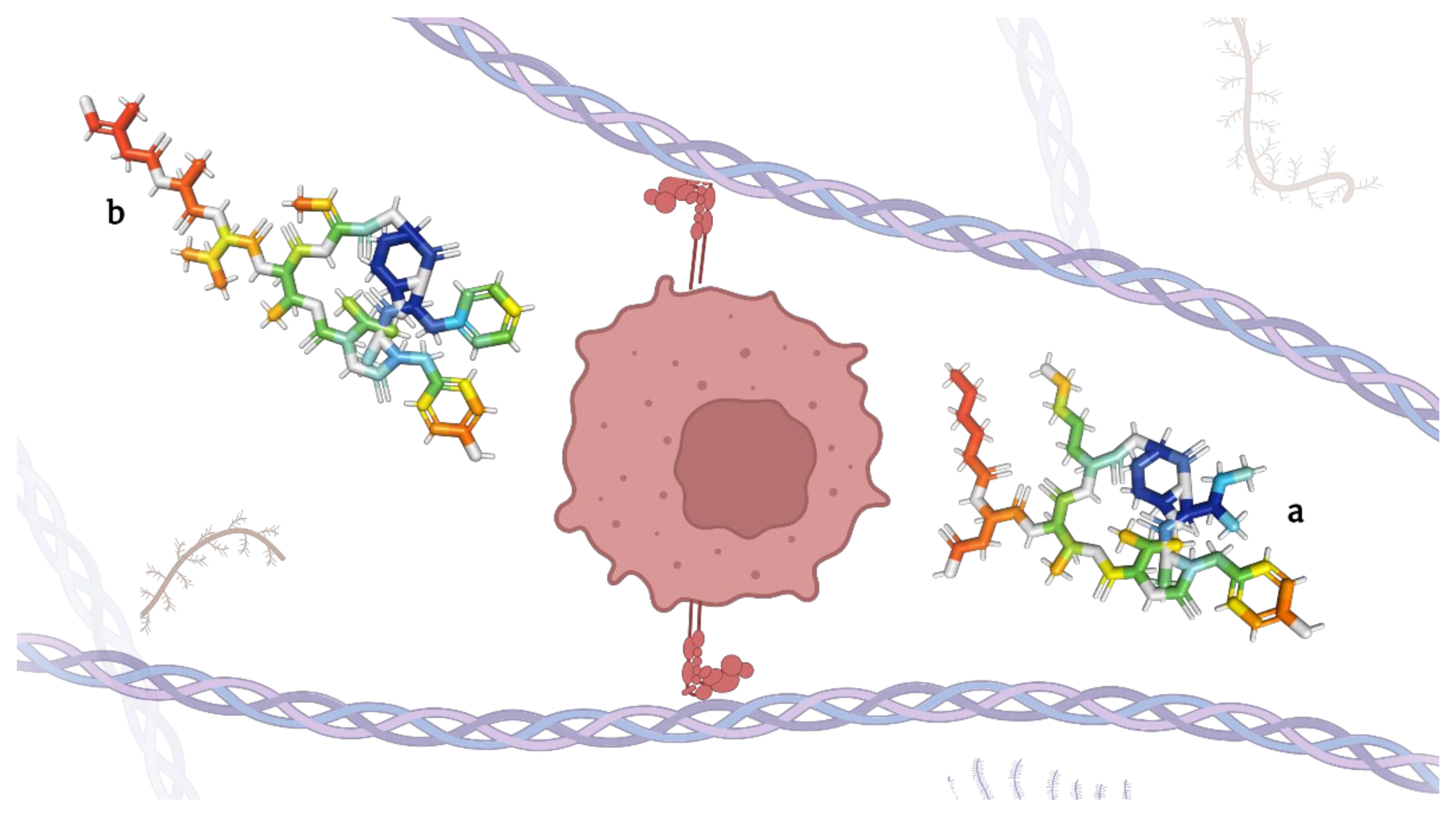
Figure 4.
Micropeptins (
a
) and tutuilamides (
b
), depsipeptides capable of inhibiting elastases.
Table 3. Formula and molecular weight of depsipeptides inhibiting elastases in tumor cells.
|
Key
|
Depsipeptide
|
IUPAC Formula
|
Molecular Weight
(g/mol)
|
Ref.
|
|
a
|
Micropeptins
|
(3S)-4-[[(2S,5S,8S,11R,12S,15S,18S,21R)-15-(4-aminobutyl)-2-[(2S)-butan-2-yl]-21-hydroxy-5-[(4-hydroxyphenyl)methyl]-4,11-dimethyl-3,6,9,13,16,22-hexaoxo-8-propan-2-yl-10-oxa-1,4,7,14,17-pentazabiCyclo [16.3.1]docosan-12-yl]amino]-3-(hexanoylamino)-4-oxobutanoic acid
|
945.1
|
[99][51]
|
|
b
|
Tutuilamides
|
(2S)-N-[(2S,5S,8S,11R,12S,15Z,18S,21R)-2-benzyl-15-ethylidene-21-hydroxy-5-[(4-hydroxyphenyl)methyl]-4,11-dimethyl-3,6,9,13,16,22-hexaoxo-8-propan-2-yl-10-oxa-1,4,7,14,17-pentazabiCyclo [16.3.1]docosan-12-yl]-2-[[(2S)-2-[[I-4-chloro-3-methylbut-3-enoyl]amino]propanoyl]amino]-3-methylbutanamide
|
1007.6
|
[100][52]
|
2.4. Depsipeptides Inhibiting Histone Deacetylases
Histone deacetylases constitute a group of enzymes catalyzing the deacetylation of lysine residues found in histones and other proteins
[106][53]. The acetylation of these residues results in a less condensed chromatin structure due to increased space between the nucleosome and DNA, thus modulating cell cycle progression
[107][54]. Interestingly, histone deacetylase inhibitors can induce apoptosis, delay cell cycle progression, and inhibit cancer cell differentiation
[30].
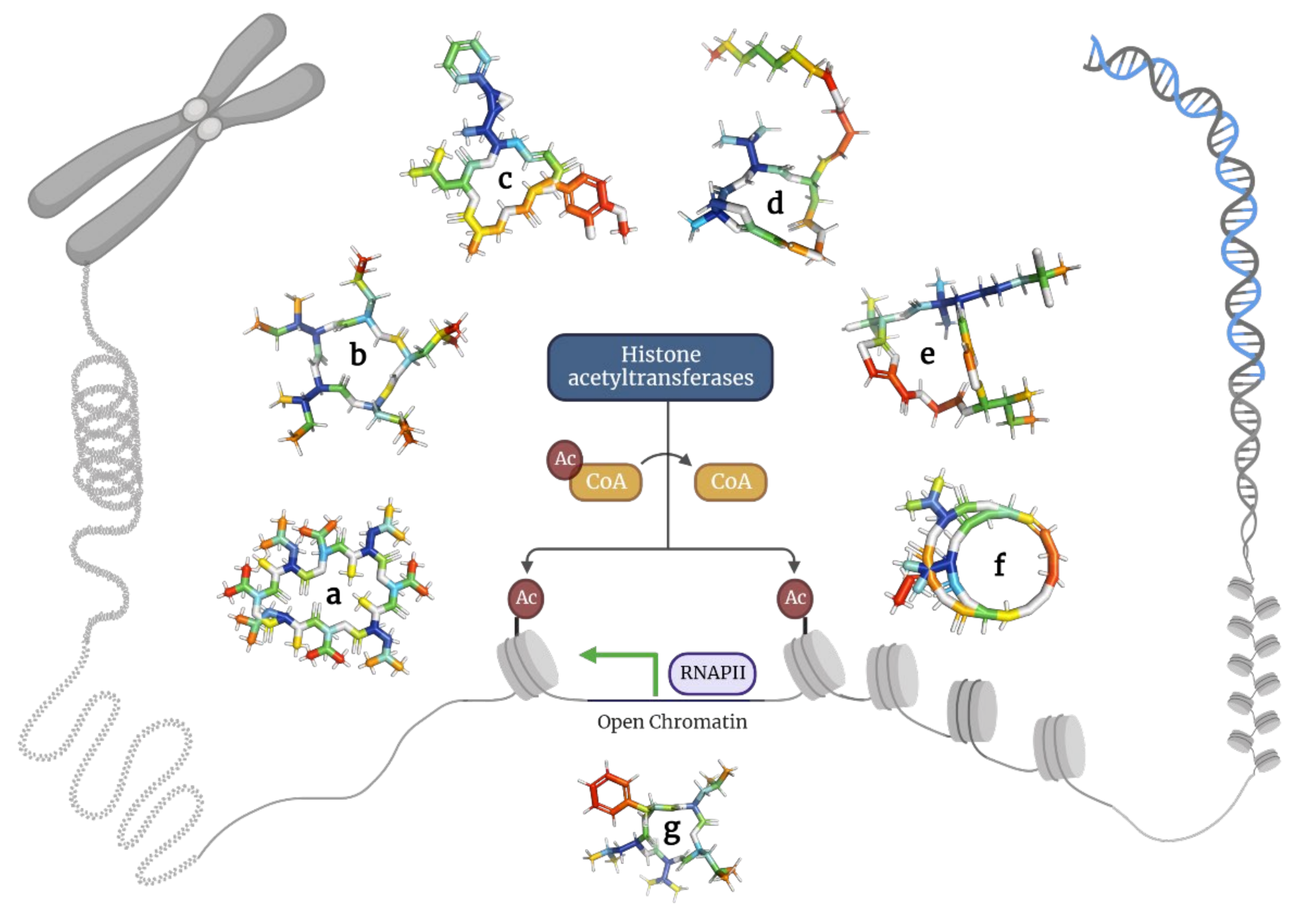
Here, the researchers describe depsipeptides that inhibit histone deacetylases in cancer cells (
Figure 5 and
Table 4).
Figure 5. Depsipeptides inhibiting histone acetyltransferases in tumor cells. (a) Bassianolide; (b) Clavatustides; (c) Cryptophycins; (d) Largazole; (e) Lyngbyabellins; (f) Romidepsin; and (g) Sansalvamide.
Table 4. Formula and molecular weight of depsipeptides inhibiting histone deacetylases in tumor cells.
|
Key
|
Depsipeptide
|
IUPAC Formula
|
Molecular Weight
(g/mol)
|
Ref.
|
|
|
d
|
Nobilamide
|
Propionyl-D-Phe-D-Leu-Phe-D-aThr-Val-Ala-Abu(2,3-dehydro)-OH
|
836.0
|
[145][67]
|
2.6. Depsipeptides Inhibiting Cell Growth
The loss of normal cell cycle regulation is one of the cornerstones of human cancer emergence
[156][68]. Thus, compounds that inhibit or terminate mitosis have been the emphasis of clinical trials for many years, with some achieving significant success
[157][69]. Certain depsipeptides are of particular interest in this field
[120[70][71],
122], mainly skyllamycins and stereocalpin compounds that inhibit transcription factors and survival signaling
[158,159][72][73].
Thus, the researchers delved into the depsipeptides that inhibit cell growth (
Table 6 and
Figure 7).
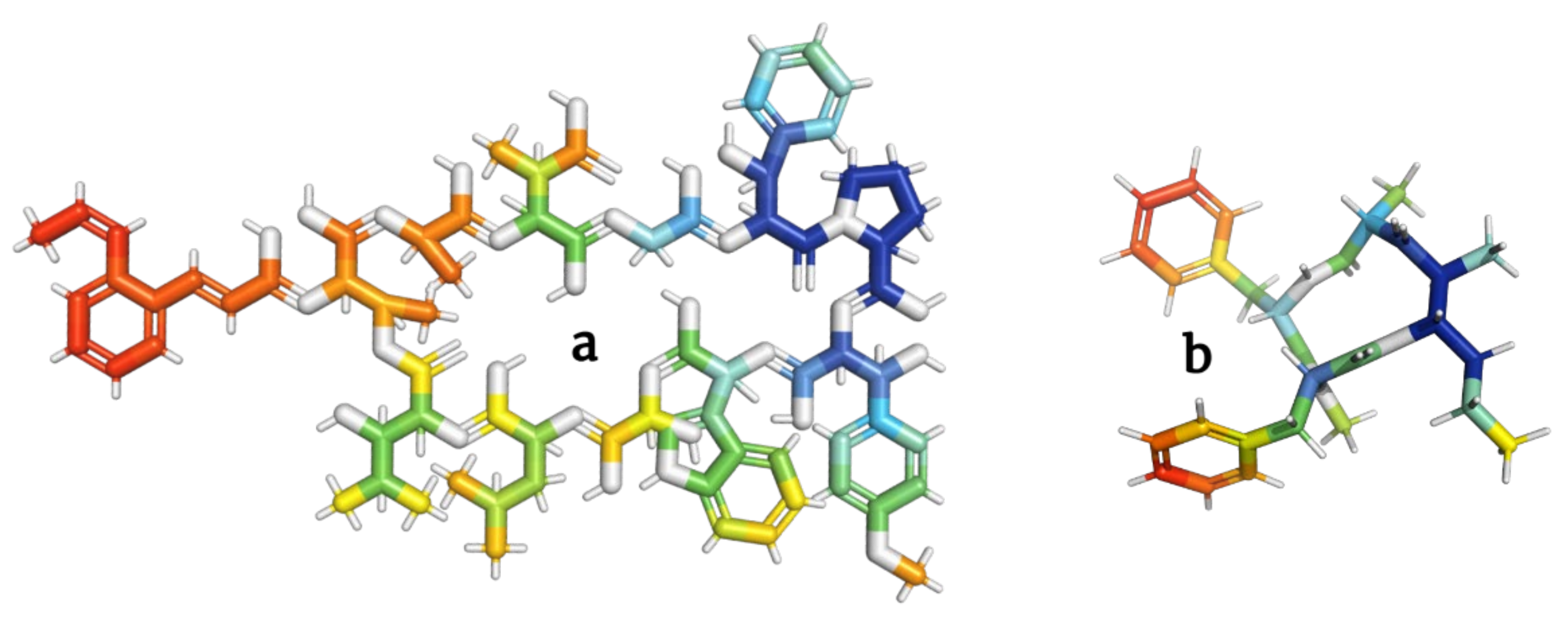
Figure 7.
Models of skyllamycins (
a
) and stereocalpin (
b
) depsipeptides inhibiting the growth of tumor cells.
Table 6. Formula and molecular weight of depsipeptides inhibiting the growth of tumor cells.
|
Key
|
Depsipeptide
|
IUPAC Formula
|
Molecular Weight
(g/mol)
|
Ref.
|
|
a
|
Bassianolide
|
Cyclo[N(Me)Leu-D-Oval-N(Me)Leu-D-Oval-N(Me)Leu-D-Oval-N(Me)Leu-D-Oval]
|
909.2
|
[108][55]
|
|
b
|
Clavatustide
|
Cyclo [2Abz-2Abz-D-Ophe-N(Et)Gly]
|
471.5
|
[ |
139][61]. Many depsipeptides derived from bacteria exert activities against microfilaments, causing complete depolymerization of microtubules and hyper-polymerization of actin
[140][62]. Moreover, they have shown the ability to destabilize microtubules, inducing caspase and Bcl-2 activation, eventually resulting in cell cycle arrest and apoptosis
[141][63].
Here, the researchers explore the depsipeptides that disrupt microfilaments in tumor cells (
Figure 6 and
Table 5).
Figure 6.
Models of depsipeptides inducing microfilament disruption in tumor cells. (
a
) Desmethoxymajusculamide C; (
b
) Dolastatin 10; (
c
) Miuraenamide; (
d
) Nobilamide I.
Table 5. Formula and molecular weight of depsipeptides inducing microfilament disruption in tumor cells.
|
Key
|
Depsipeptide
|
IUPAC Formula
|
Molecular Weight
(g/mol)
|
Ref.
|
|
a
|
Skyllamycins
|
N-[I-3-[2-[(Z)-1-Propenyl]phenyl]propenoyl]-Cyclo[L-Thr*-L-Ala-[(3S)-3-Me-L-Asp-]-Gly-[(βS)-β-hydroxy-L-Phe-]-L-Pro-[(βS)-β-hydroxy-O-Me-L-Tyr-]-D-Trp-[(2S)-2-hydroxy-Gly-]-D-Leu-[(3S)-3-hydroxy-D-Leu-]-]
|
1483.6
|
[160][74]
|
|
b
|
Stereocalpin
|
Cyclo[Phe-N(Me)Phe-Unk]
|
492.6
|
[161][75]
|
2.7. Depsipeptides Inhibiting Topoisomerases
Topoisomerases are enzymes that relieve torsional stress within DNA during replication and transcription by cleaving one strand to unwind the supercoiled DNA
[167,168][76][77]. Inhibiting these enzymes leads to DNA damage, nicotinamide adenine dinucleotide phosphate oxidase-dependent generation of reactive oxygen species, p21 activation, and eventual cell cycle arrest and senescence
[169][78].
Here, the researchers describe the relevant depsipeptides targeting topoisomerases in cancer cells (
Table 7 and
Figure 8).
Figure 8.
Models of fusaristatin A (
a
) and neo-N-methylsansalvamide (
b
) depsipeptides targeting topoisomerases in tumor cells.
Table 7. Formula and molecular weight of depsipeptides targeting topoisomerases in tumor cells.
|
Key
|
|
a
|
Desmethoxymajusculamide C
|
Cyclo[Ala-Unk-Ala-Unk-Gly-N(Me)Ile-Gly-N(Me)Val-N(Me)Phe]
|
955.2
|
[142][64]
|
|
b
|
Dolastatin
|
N(Me2)Val-Val-Unk
|
785.1
| 109][56]
|
|
| [ | 143 | ] | [65]
|
|
c
|
Cryptophycin |
|
c
|
|
Miuraenamide
10-[(3-chloro-4-methoxyphenyl)methyl]-6-methyl-3-(2-methylpropyl)-16-[1-(3-phenyloxiran-2-yl)ethyl]-1,4-dioxa-8,11-diazacyclohexadec-13-ene-2,5,9,12-tetrone
|
(3E,15E)-6-[(3-bromo-4-hydroxyphenyl)methyl]-3-[methoxy(phenyl)methylidene]-7,9,16,19-tetramethyl-1-oxa-4,7,10-triazacyclononadec-15-ene-2,5,8,11-tetrone
655.2
|
[ |
684.6
| 32]
|
|
| [ | 110 | ] | [57]
|
|
| [ | 144 | ] | [66]
|
d
|
Coibamide
|
d |
N(Me2)Val-Oval-N(Me)Ser(Me)-N(Me)Leu-N(Me)Thr(1)-N(Me)Ser(Me)-N(Me)Ile-Ala-N(Me)Leu-Tyr(Me)-N(Me)Ala-(1)
|
|
Largazole
|
1287.6
|
|
S-[I-4-[(5R[33] |
|
,8 | S | ,11S)-5-Me-6,9,13-trioxo-8-propan-2-yl-10-oxa-3,17-dithia-7,14,19,20-tetrazatriCyclo [14.2.1.12,5]icosa-1(18),2(20),16(19)-trien-11-yl]but-3-enyl] octanethioate
|
622.9
|
[111][58]
|
e
|
Dehydrodidemnin
|
Pyruvoyl-Pro-D-N(Me)Leu-D-Thr(1)-Unk-Leu-Pro-DL-N(Me)Tyr(Me)-(1)
|
1110.3
|
[34]
|
|
e
|
Lyngbyabellin
|
(7 |
f
|
Enniatin
|
Cyclo[DL-Oval-DL-N(Me)xiIle-DL-Oval-DL-N(Me)xiIle-DL-Oval-DL-N(Me)xiIle]
|
681.9
|
[35]
|
|
g
|
Grassypeptolide
|
Cyclo[D-N(Me)Leu-D-aThr-Unk-N(Me)Val-Pro-Unk]
|
1102.4
|
[36]
|
|
h
|
Hantupeptin
|
Cyclo[N(Me)Ile-Ophe-Pro-N(Me)Val-Unk-Val]
|
740.9
|
[37]
|
|
i
|
Lagunamide
|
Cyclo[Ala-D-N(Me)Phe-Sar-aIle-N(Me)Ala-Unk]
|
842.1
|
[38]
|
|
j
|
Tiahuramide
|
Cyclo[N(Me)Ile-Unk-Val-N(Me)Val-Ophe-Pro]
|
736.9
|
[39]
|
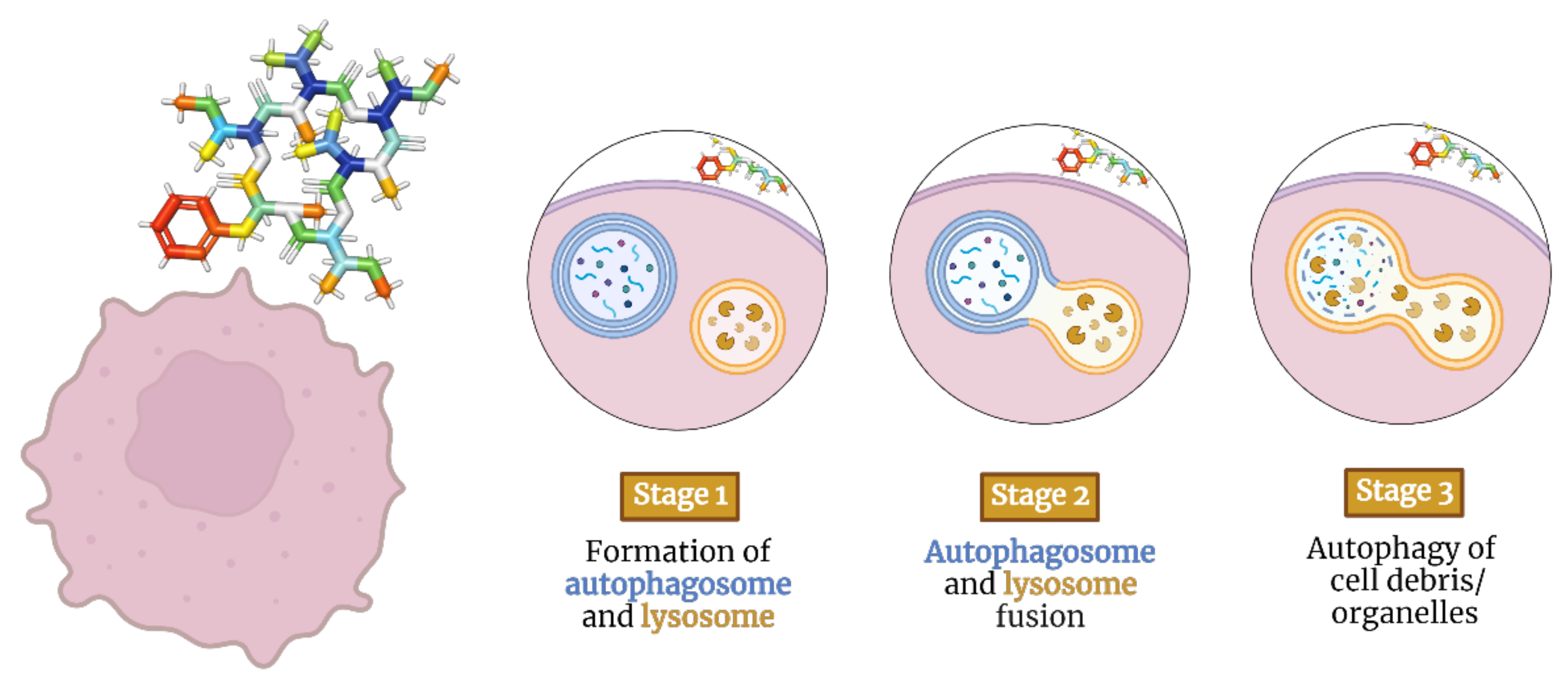
2.2. Depsipeptides Inducing Autophagy in Tumor Cells
Autophagy is the process that uses autophagosomes to deliver cytoplasmic material to the lysosome for degradation
[89][40]. Autophagy-inducing depsipeptides can activate intracellular messengers such as adenosine monophosphate-activated protein kinase and transcription factor EB
[90][41] (
Figure 3). They can also inhibit survival-inducing molecules such as hypoxia-inducible factor 1-alpha and mammalian target of rapamycin
[29], thereby inducing autophagy through various mechanisms
[91,92][42][43].
Figure 3.
Staging model of depsipeptides inducing autophagy in tumor cells.
Beauvenniatins
Initially isolated from a mycelial extract of
Acremonium spp., with additional analogs also produced by
Fusarium and
Isaria spp.
[93][44], the structure of these peptides is a hybrid between the aromatic and aliphatic cyclodepsipeptides, with moieties of N-Me-valine, N-Me-leucine, and N-Me-phenylalanine (
Table 2)
[54][45].
Table 2. Formula and molecular weight of beauvenniatin, a depsipeptide triggering autophagy in tumor cells.
Beauvenniatin F has exhibited strong cytotoxicity against adriamycin-resistant myelogenous leukemia cells (K-562), with an IC
50 of 2.78 μM, and autophagy-inducing activity at a concentration of 20 μM in green fluorescent protein-light chain 3-stable HeLa cells
[94][46]. These depsipeptides also displayed cytotoxicity against epithelial carcinoma (KB), lymphoma (BC), and lung carcinoma cells (NCI-H187), with IC
50 values ranging from 1.00 to 2.29 μM as well as against kidney epithelial cells (Vero) at 1.9–5.5 μM
|
| Depsipeptide
|
IUPAC Formula
|
Molecular Weight
(g/mol)
|
Ref.
|
|---|
|
a
|
Fusaristatin A
|
3-[6,13-dimethyl-10-methylidene-2,5,9,12-tetraoxo-14-[(5E,7E)-3,7,11-trimethyl-4-oxoheptadeca-5,7-dienyl]-1-oxa-4,8,11-triazacyclotetradec-3-yl]propanamide
|
658.9
|
[170][79]
|
|
b
|
N-methylsansalvamide
|
Cyclo[Leu-Oleu-Val-N(Me)Leu-Phe]
|
S |
,14 |
S | ,18 | S)-7-[(2S)-butan-2-yl]-14-(4,4-dichloropentyl)-18-(2-hydroxypropan-2-yl)-15,15-dimethyl-13,17-dioxa-9,20-dithia-3,6,22,23-tetrazatriCyclo [17.2.1.18,11]tricosa-1(21),8(23),10,19(22)-tetraene-2,5,12,16-tetrone
|
691.7
|
[18]
|
|
| 600.8 |
|
| [28]
|
f
|
Romidepsin
|
7-ethylidene-4,21-di(propan-2-yl)-2-oxa-12,13-dithia-5,8,20,23-tetrazabiCyclo [8.7.6]tricos-16-ene-3,6,9,19,22-pentone
|
540.7
|
[112][59]
|
|
g
|
Sansalvamide
|
Cyclo[L-leucyl-N-oxa-L-leucyl-L-valyl-L-leucyl-L-phenylalanyl] |
2.8. Oncosis
Oncosis is a form of programmed cell death characterized by organelle swelling and membrane disruption
[177][80]. It is a type of cell injury that involves energy depletion, impairment of ionic pumps, swelling, dilation of the Golgi apparatus and endoplasmic reticulum, chromatin clumping, and the formation of cytoplasmic blebs
[178][81].
Compared with apoptosis, oncosis eventually results in necrosis with karyolysis and cell swelling, instead of karyorrhexis and cell shrinkage
[179][82]. One of the most studied oncosis-inducing depsipeptides in cancer cells is kahalalide F (
Figure 9 and
Table 8), a molecule that causes disruption of the mitochondrial membrane potential and alters the permeability of lysosomal membranes
[91][42].
Figure 9.
Model of kahalalide F: the only depsipeptide inducing oncosis in tumor cells.
Table 8. Formula and molecular weight of kahalalide F, the only depsipeptide inducing oncosis in tumor cells.
|
Depsipeptide
|
IUPAC Formula
|
Molecular Weight
(g/mol)
|
Ref.
|
|
Kahalalide F
|
L-Val, N-(5-Me-1-oxohexyl)-D-valyl-l-threonyl-l-valyl-D-valyl-D-prolyl-l-ornithyl-D-alloisoleucyl-D-allothreonyl-D-alloisoleucyl-D-valyl-l-phenylalanyl-(2z)-2-amino-2-butenoyl-, (13->8)-lactone
|
1477.9
|
[180][83]
|
|
|
|
|
586.8 |
|
|
|
| [ | 113 | ] | [60]
|
2.5. Depsipeptides Disrupting Microfilaments
Microtubule–actin interactions are fundamental for many cellular processes such as cell motility, cell division, and cytoskeleton dynamics
[
3. Current Methods for Purification and Synthesis of Depsipeptides
3.1. Proved Methods for Purifying Depsipeptides
Nowadays, depsipeptides can be purified using different methods
[218,219][84][85]. When derived from natural sources, the organism-produced depsipeptides can be analyzed using high-performance liquid chromatography (HPLC)
[220,221][86][87]. Some studies have coupled a photo-diode array and used ultra-performance liquid chromatography in tandem with high-resolution mass spectrometry
[222][88].
Frequently, in vitro and even in vivo assays are performed simultaneously during purification to analyze the biological properties of interest
[223][89]. Yu et al. used reversed-phase HPLC (RP-HPLC) to purify and quantify depsipeptides and short oligomers of glycine
[220][86]. In 2006, Matsuo et al. also used RP-HPLC to purify urukthapelstatin
[224][90], a depsipeptide with inhibitory effects on human lung cancer (A549) cells
[221][87].
RP-HPLC was also used to purify trikoveramides A–C, peptides that belong to the kulolide superfamily and inhibited the growth of human leukemia cells (MOLT-4)
[225][91]. The well-known anticancer peptide, coibamide A, was also purified by RP-HPLC
[226][92]. In 2015, Kaur et al. achieved the purification of the novel cyclic depsipeptide, YM-280193, with the ability to inhibit platelet aggregation
[227][93].
3.2. Synthesis of Depsipeptides
Depsipeptides were originally synthesized to improve the solubility of ultra-long peptides by adding an ester-linked residue that would alter the linear structure
[228][94]. In addition to the presence of at least one ester and one amide bond in their structures, most depsipeptides contain a wide variety of acyl groups and moieties added through the assembly line
[16].
Certain authors have proposed the synthesis of these peptides using solid-phase peptide synthesis with a fluorenylmethoxycarbonyl (Fmoc)-protected N-terminus, lysine, or aspartic acid as the charged entities, and lactic acid as the ester moiety to maintain hydrophobicity
[219][85]. A stepwise Fmoc-based solid-phase methodology to create a highly complex depsipeptide has been reported, with the addition of small amounts of organic acids to improve this stepwise approach and minimize secondary reactions
[229][95].
For instance, coibamide A has been synthesized using Fmoc-based solid-phase synthesis, but with the initial step being the attachment of the Fmoc-Tyr-allyl ester to the free hydroxyl group on a 2-chlorotrityl chloride resin
[226][92]. Another depsipeptide, teixobactin, and its analogs have also been synthesized using Fmoc-based solid-phase synthesis
[230][96].