Transcription of the mitochondrial genome is essential for the maintenance of oxidative phosphorylation (OXPHOS) and other functions directly related to this unique genome. Considerable evidence suggests that mitochondrial transcription is dysregulated in cancer and cancer metastasis and contributes significantly to cancer cell metabolism. RecThently, inhibitors of the mitochondrial DNA-dependent RNA polymerase (POLRMT) were identified as potentially attractive new anti-cancer compounds. These molecules (IMT1, IMT1B) inactivate cancer cell metabolism through reduced transcription of mitochondrially-encoded OXPHOS subunits such as ND1-5 (Complex I) and COI-IV (Complex IV).
- mitochondria
- mitochondrial transcription
- oxidative phosphorylation
- POLRMT
- ClpP
- mitochondrial DNA
1. Mitochondrial Transcription and Metabolism as Targets for New Anti-Cancer Approaches
2. Functional Roles of POLRMT in Mitochondria
POLRMT, the only RNA polymerase present in the mitochondria, transcribes and facilitates the replication of mtDNA [43]. POLRMT is therefore a pivotal enzyme for the expression of the genes encoded by mtDNA and for the maintenance of functional mitochondria and related metabolic processes [44]. During mitochondrial transcription, POLRMT works in concert with mitochondrial transcriptional factor A (TFAM) and mitochondrial dimethyladenosine transferase 2 (TFB2M) to initiate mitochondrial transcription. TFAM binds to mtDNA promoters to direct POLRMT to initiate transcription, while TFB2M aids POLRMT in melting and stabilizing the mtDNA strands to enable transcription (Figure 1) [9,43][9][43]. TFAM has an additional role in mtDNA maintenance. TFAM binds nonspecifically to mtDNA, causing the DNA to bend and allowing the packaging of genetic material into nucleoids [9,45][9][45]. TFAM typically binds to mtDNA every 15–20 kb, and the abundance of TFAM bound to mtDNA is critically important in mtDNA transcriptional regulation [46,47][46][47]. Overexpression of TFAM has been shown to reduce transcription and expression of mitochondrially-encoded genes, potentially due to TFAM blocking POLRMT access to mtDNA [47,48][47][48]. However, some tissues have been observed to upregulate expression of POLRMT, potentially as a mechanism to overcome high TFAM levels and promote increased transcription [47]. For example, transcription at LSP, HSP1, or HSP2 can be regulated by the TFAM protein level because each promotor is transcribed optimally at a different TFAM:mtDNA ratio [47,48][47][48]. These data suggest both TFAM and POLRMT play critical roles in regulating mitochondrial transcription and function.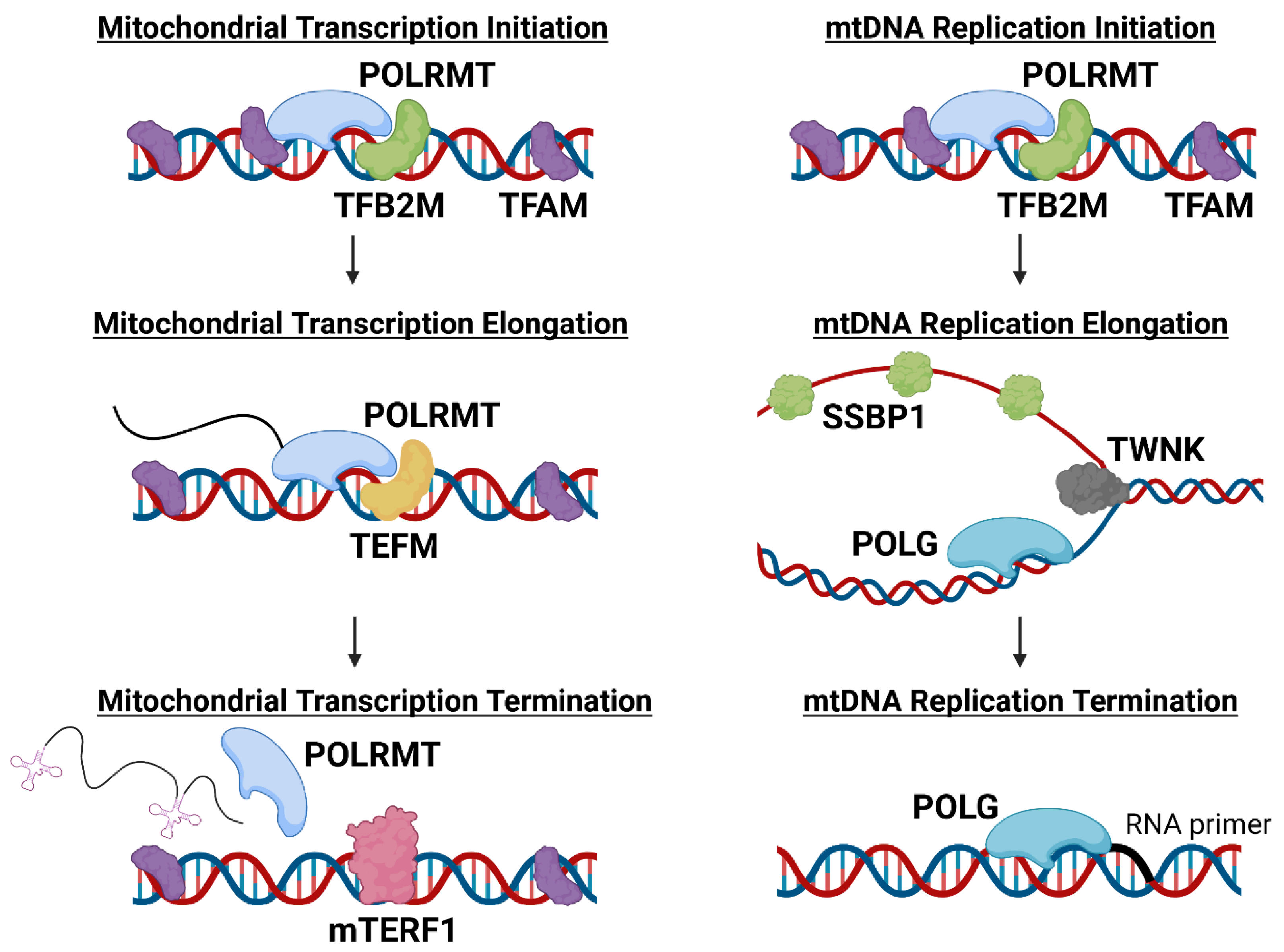
3. POLRMT Inhibition as an Anti-Cancer Strategy
4. Small Molecule ClpP Agonists as Anti-Cancer Compounds
Recently, a novel class of mitochondria-targeting drugs were discovered based on their highly unusual property of activating the mitochondrial protease ClpP. ONC201 was first identified as a potential anti-cancer compound in a screen for small molecule inducers of tumor necrosis factor-related apoptosis-inducing ligand (TRAIL) [37]. ONC201 (Dordaviprone) and ONC206 have now progressed to phase I, II, and III clinical trials for gliomas, acute myeloid leukemia (AML), endometrial, breast, and other cancers [63,64,65][56][57][58]. More recently, Madera Therapeutics LLC developed scaffold analogs of ONC201 (TR compounds) which are highly potent and selective agonists of ClpP [33]. Studies confirmed that these compounds bind to ClpP, the proteolytic barrel of the ClpXP mitochondrial matrix protease. The ClpXP protease is comprised of two components, ClpP and the unfoldase ClpX, which plays an important role in protein quality control and mitochondrial proteostasis (Figure 2) [66,67][59][60].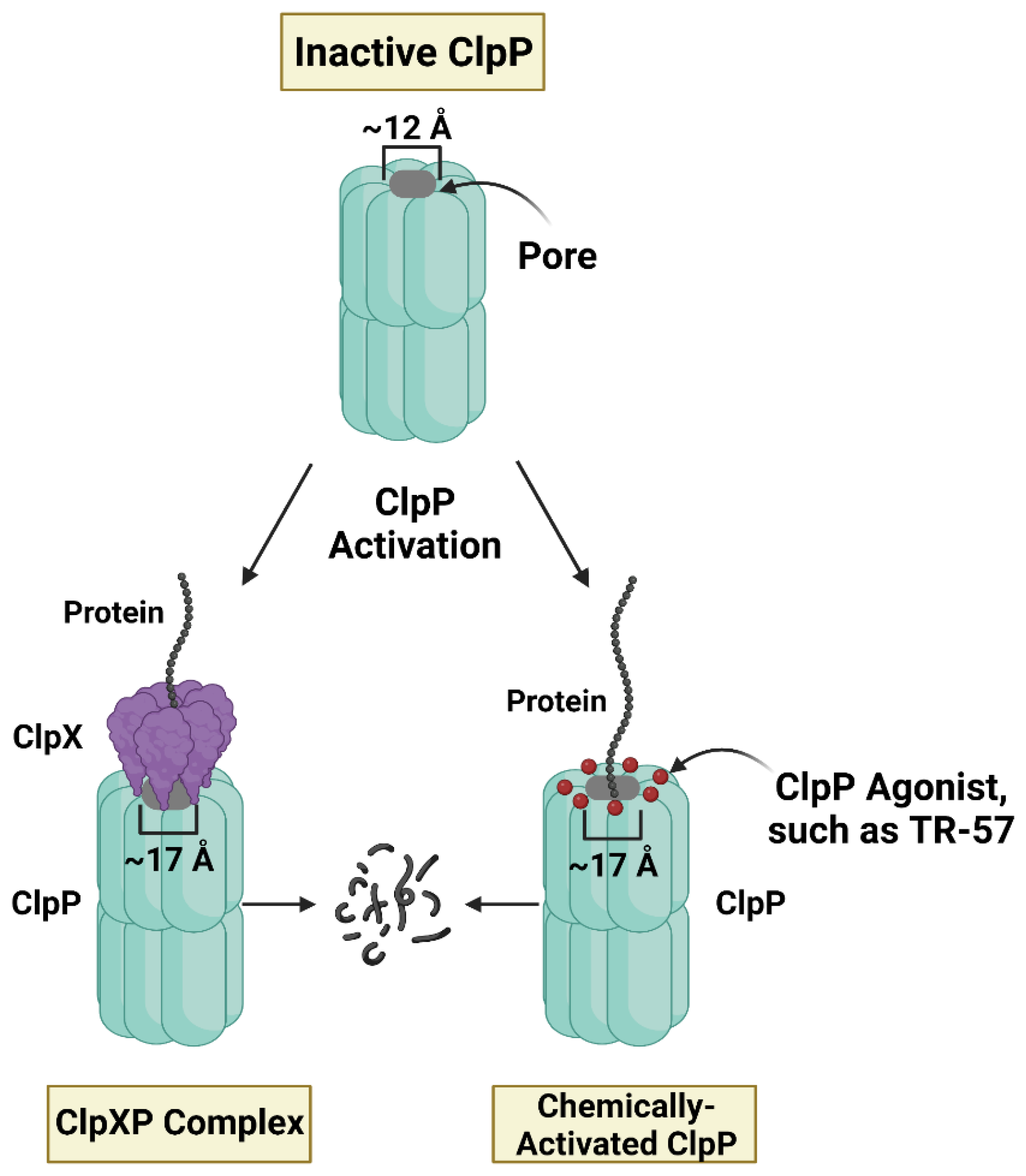
5. Mechanistic Similarities between ClpP Agonists and POLRMT Inhibitors
5.1. Inhibition of Cell Proliferation
5.2. Cytostatic to Cancer Cells, Harmless to Normal Cells
One of the unusual properties of ClpP agonists is that they appear to adversely affect cancer cells but have little or no effect on normal cells. Several groups have tested ClpP agonists on immortalized cell lines (i.e., HFF) or primary cells and observed no significant effects on cell proliferation, viability, or apoptosis as compared to the cancer cell models [42,68,75,84][42][61][63][73]. Additionally, ClpP agonists were reported to be well tolerated clinically, with few side effects observed in patients with refractory solid tumors and neuroendocrine tumors [64,65][57][58]. In contrast, other mitochondria-targeting drugs such as metformin and CPI-631 have shown negative side effects such as gastrointestinal tract disorders, nausea, and vomiting; however, these toxicities were not reported in ONC201 clinical trials [64,65,85,86][57][58][74][75].5.3. Dysregulation of Cancer Cell Metabolic Programs
Both ClpP agonists and POLRMT inhibitors induce major metabolic alterations in cancer cells, most prominently observed with the inhibition of OXPHOS [20,34,75,78,83][20][34][63][66][72]. Both classes of compounds reduce the mitochondrially-encoded protein components required for OXPHOS [20,34][20][34]. However, the mechanism by which OXPHOS is impaired in response to ClpP activation is less well understood than the proposed mechanism of POLRMT inhibitors. Loss of POLRMT activity results in the direct inhibition of mitochondrial transcription, which in turn prevents the expression of the 13 protein-coding genes essential for respiration [20]. Significant decreases in mRNA levels of several of these genes have been shown following IMT1 treatment [20[20][32],32], confirming inhibition of mitochondrial transcription following POLRMT inhibition. This combined with the loss of their cognate proteins (i.e., COI) could explain the observed inhibition of OXPHOS in these cells [20]. Another common metabolic alteration observed following ClpP activation or POLRMT inhibition is AMPK activation [20,75][20][63]. Greer et al. (2018) observed a decline in ATP levels and an increase in AMPK phosphorylation (Thr172) following ONC201 treatment [75][63]. Similar effects were observed with IMT1 treatment [20]. AMPK activation is mediated by a high cellular AMP/ATP ratio [87][76], further indicating that inhibition of OXPHOS and/or other mitochondrial processes required for ATP generation is occurring in response to these two different mitochondrial targeting approaches. As an expected consequence of OXPHOS inhibition, ClpP agonists have been shown to cause a compensatory increase in glycolysis [34,75,78][34][63][66]. A similar compensatory response has not been demonstrated following POLRMT inhibition, although it may be predicted.5.4. Loss of mtDNA Content
An important similarity between ClpP agonists and POLRMT inhibitors is the corresponding reduction in mtDNA content, despite observations that the mtDNA copy number appears to change at different rates. IMT1 treatment reportedly took ~96 h to reduce the mtDNA copy number to ~25% of its original number [20], whereas ONC201 treatment required less time (~48 h) to reduce the mtDNA copy number to the same level [75][63]. This may in part be determined by the rate of TFAM protein loss. TFAM expression is closely correlated to the mtDNA copy number, likely because it binds to and packages mtDNA [98][77]. ClpP activation results in a time- and dose-dependent decline in TFAM protein that correlates with observed decreases in mtDNA content [33[33][34][63][66],34,75,78], as do POLRMT inhibitors (Daglish, unpublished observations (IMT1)).5.5. Inhibition of Mitochondrial Transcription
It has been shown that POLRMT and TFAM protein expression decreases after treatment with ClpP agonists, but it has not been established that mitochondrial transcription is inhibited. The scholars predicted that mitochondrial transcription would be impaired by ClpP agonists due to their ability to deplete POLRMT, TFAM, and TEFM proteins which are all essential for mitochondrial transcription (Figure 1). Both TR-57 and IMT1 inhibited mitochondrial transcription in the tumorigenic immortalized HEK293T cells line (Figure 3) [101,102][78][79]. mRNA transcript levels of two mitochondrially-encoded protein genes required for OXPHOS (ND1 and ND6) were measured by quantitative PCR to monitor mitochondrial transcription from the heavy and light strands of mtDNA, respectively. Both treatments significantly decreased ND1 and ND6 transcript levels after 3 and 6 h, although there was a greater decrease observed following IMT1 treatment (Figure 3).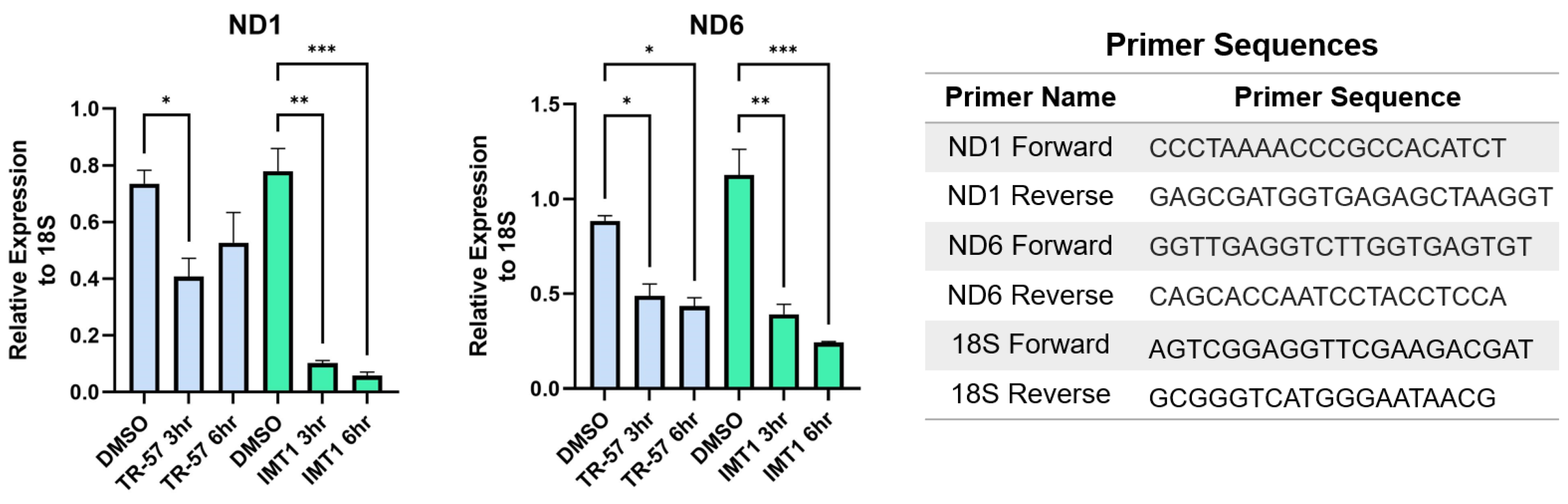
6. Mechanistic Differences between ClpP Agonists and POLRMT Inhibitors
6.1. Differences in Treatment Response Times
Despite the many mechanistic similarities between ClpP agonists and POLRMT inhibitors, there are several notable differences. Primarily, ClpP agonists inhibit cell proliferation much more robustly than POLRMT inhibitors. TR-57 strongly inhibits HEK293T cell proliferation following 72 h treatment, whereas IMT1 treatment requires a minimum of 120 h to show a similar inhibition of cell proliferation (Figure 4). In terms of potency, the TR compounds are more potent inhibitors of HEK293T growth; the IC50 of TR-57 in HEK293T cells is ~15 nM after 72 h, whereas the POLRMT inhibitor IMT1 has an IC50 of ~190 nM after 120 h of treatment (Figure 4).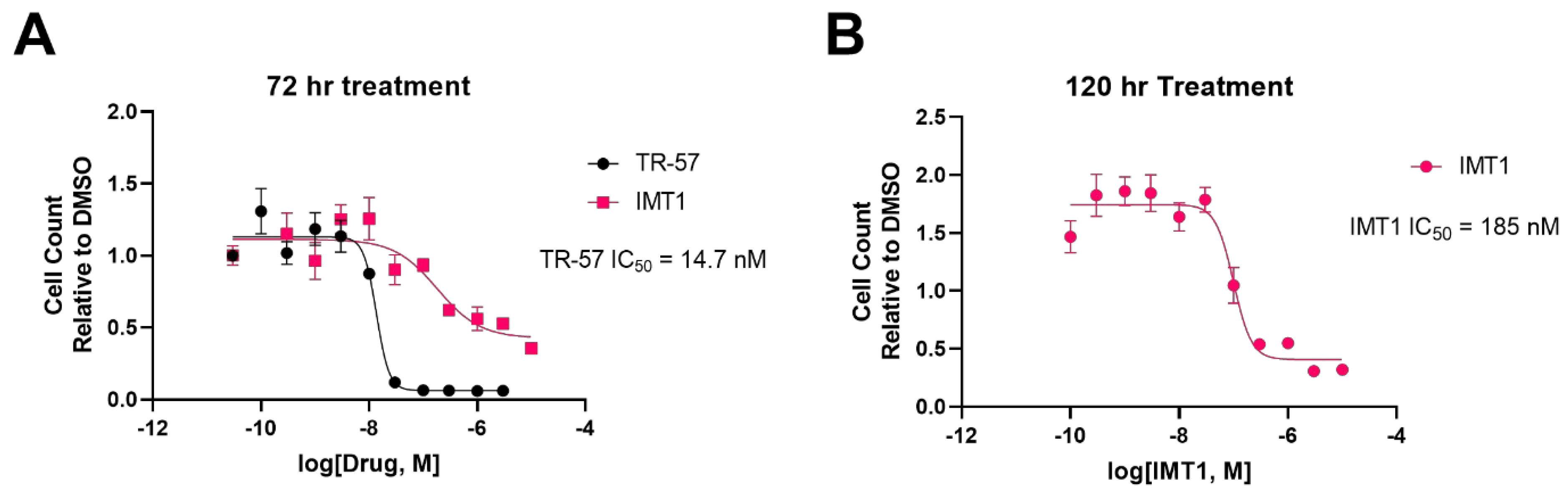
6.2. Known and Unknown Mechanisms of Resistance
There are a few known cancer cell mechanisms of resistance to ClpP agonists, including p0 cells, fumarate hydratase knockout cell lines, and ClpP knockout or ClpP mutant cell lines [61][80]. The specific ClpP mutation, D190A, is known to confer resistance to ClpP agonists [68][61]. Comparatively, there are specific POLRMT mutations that confer resistance to IMT1. Bonekamp et al. (2020) reported six mutations in POLRMT that lead to IMT1-resistance [20]. A CRISPR screen aimed at identifying mechanisms of resistance against IMT1 found that loss of von Hippel-Landau protein (VHL) and mammalian target of rapamycin complex 1 (mTORC1) expression produced IMT1-resistance in RKO cells [104][81].
6.3. Inhibiting One Protein versus Degrading Many Proteins
References
- Chakrabarty, R.P.; Chandel, N.S. Beyond ATP, New Roles of Mitochondria. Biochemist 2022, 44, 2–8.
- Haynes, C.M.; Petrova, K.; Benedetti, C.; Yang, Y.; Ron, D. ClpP Mediates Activation of a Mitochondrial Unfolded Protein Response in C. Elegans. Dev. Cell 2007, 13, 467–480.
- Giampazolias, E.; Zunino, B.; Dhayade, S.; Bock, F.; Cloix, C.; Cao, K.; Roca, A.; Lopez, J.; Ichim, G.; Proïcs, E.; et al. Mitochondrial Permeabilization Engages NF-ΚB-Dependent Anti-Tumour Activity under Caspase Deficiency. Nat. Cell Biol. 2017, 19, 1116–1129.
- Collins, Y.; Chouchani, E.T.; James, A.M.; Menger, K.E.; Cochemé, H.M.; Murphy, M.P. Mitochondrial Redox Signalling at a Glance. J. Cell Sci. 2012, 125, 801–806.
- Riley, J.S.; Tait, S.W. Mitochondrial DNA in Inflammation and Immunity. EMBO Rep. 2020, 21, e49799.
- Qureshi, M.A.; Haynes, C.M.; Pellegrino, M.W. The Mitochondrial Unfolded Protein Response: Signaling from the Powerhouse. J. Biol. Chem. 2017, 292, 13500–13506.
- Bueno, M.J.; Ruiz-Sepulveda, J.L.; Quintela-Fandino, M. Mitochondrial Inhibition: A Treatment Strategy in Cancer? Curr. Oncol. Rep. 2021, 23, 49.
- Rackham, O.; Filipovska, A. Organization and Expression of the Mammalian Mitochondrial Genome. Nat. Rev. Genet. 2022, 23, 606–623.
- Hillen, H.S.; Temiakov, D.; Cramer, P. Structural Basis of Mitochondrial Transcription. Nat. Struct. Mol. Biol. 2018, 25, 754–765.
- Hashimoto, Y.; Kurita, M.; Aiso, S.; Nishimoto, I.; Matsuoka, M. Humanin Inhibits Neuronal Cell Death by Interacting with a Cytokine Receptor Complex or Complexes Involving CNTF Receptor Alpha/WSX-1/Gp130. Mol. Biol. Cell 2009, 20, 2864–2873.
- Ro, S.; Ma, H.-Y.; Park, C.; Ortogero, N.; Song, R.; Hennig, G.W.; Zheng, H.; Lin, Y.-M.; Moro, L.; Hsieh, J.-T.; et al. The Mitochondrial Genome Encodes Abundant Small Noncoding RNAs. Cell Res. 2013, 23, 759–774.
- Kummer, E.; Ban, N. Mechanisms and Regulation of Protein Synthesis in Mitochondria. Nat. Rev. Mol. Cell Biol. 2021, 22, 307–325.
- Sainero-Alcolado, L.; Liaño-Pons, J.; Ruiz-Pérez, M.V.; Arsenian-Henriksson, M. Targeting Mitochondrial Metabolism for Precision Medicine in Cancer. Cell Death Differ. 2022, 29, 1304–1317.
- Wheaton, W.W.; Weinberg, S.E.; Hamanaka, R.B.; Soberanes, S.; Sullivan, L.B.; Anso, E.; Glasauer, A.; Dufour, E.; Mutlu, G.M.; Budigner, G.S.; et al. Metformin Inhibits Mitochondrial Complex I of Cancer Cells to Reduce Tumorigenesis. eLife 2014, 3, e02242.
- Moullan, N.; Mouchiroud, L.; Wang, X.; Ryu, D.; Williams, E.G.; Mottis, A.; Jovaisaite, V.; Frochaux, M.V.; Quiros, P.M.; Deplancke, B.; et al. Tetracyclines Disturb Mitochondrial Function across Eukaryotic Models: A Call for Caution in Biomedical Research. Cell Rep. 2015, 10, 1681–1691.
- Dijk, S.N.; Protasoni, M.; Elpidorou, M.; Kroon, A.M.; Taanman, J.-W. Mitochondria as Target to Inhibit Proliferation and Induce Apoptosis of Cancer Cells: The Effects of Doxycycline and Gemcitabine. Sci. Rep. 2020, 10, 4363.
- Rivas, M.O.G.; Stuart, S.D.; Thach, D.; Dahan, M.; Shorr, R.; Zachar, Z.; Bingham, P.M. Evidence for a Novel, Effective Approach to Targeting Carcinoma Catabolism Exploiting the First-in-Class, Anti-Cancer Mitochondrial Drug, CPI-613. PLoS ONE 2022, 17, e0269620.
- Delaunay, S.; Pascual, G.; Feng, B.; Klann, K.; Behm, M.; Hotz-Wagenblatt, A.; Richter, K.; Zaoui, K.; Herpel, E.; Münch, C.; et al. Mitochondrial RNA Modifications Shape Metabolic Plasticity in Metastasis. Nature 2022, 607, 593–603.
- Bergbrede, T.; Hoberg, E.; Larsson, N.-G.; Falkenberg, M.; Gustafsson, C.M. An Adaptable High-Throughput Technology Enabling the Identification of Specific Transcription Modulators. SLAS Discov. 2017, 22, 378–386.
- Bonekamp, N.A.; Peter, B.; Hillen, H.S.; Felser, A.; Bergbrede, T.; Choidas, A.; Horn, M.; Unger, A.; Di Lucrezia, R.; Atanassov, I.; et al. Small-Molecule Inhibitors of Human Mitochondrial DNA Transcription. Nature 2020, 588, 712–716.
- Reznik, E.; Miller, M.L.; Şenbabaoğlu, Y.; Riaz, N.; Sarungbam, J.; Tickoo, S.K.; Al-Ahmadie, H.A.; Lee, W.; Seshan, V.E.; Hakimi, A.A.; et al. Mitochondrial DNA Copy Number Variation across Human Cancers. eLife 2016, 5, e10769.
- van Osch, F.H.M.; Voets, A.M.; Schouten, L.J.; Gottschalk, R.W.H.; Simons, C.C.J.M.; van Engeland, M.; Lentjes, M.H.F.M.; van den Brandt, P.A.; Smeets, H.J.M.; Weijenberg, M.P. Mitochondrial DNA Copy Number in Colorectal Cancer: Between Tissue Comparisons, Clinicopathological Characteristics and Survival. Carcinogenesis 2015, 36, 1502–1510.
- Yuan, Y.; Ju, Y.S.; Kim, Y.; Li, J.; Wang, Y.; Yoon, C.J.; Yang, Y.; Martincorena, I.; Creighton, C.J.; Weinstein, J.N.; et al. Comprehensive Molecular Characterization of Mitochondrial Genomes in Human Cancers. Nat. Genet. 2020, 52, 342–352.
- Tseng, L.-M.; Yin, P.-H.; Chi, C.-W.; Hsu, C.-Y.; Wu, C.-W.; Lee, L.-M.; Wei, Y.-H.; Lee, H.-C. Mitochondrial DNA Mutations and Mitochondrial DNA Depletion in Breast Cancer. Genes Chromosom. Cancer 2006, 45, 629–638.
- Wallace, D.C. Mitochondria and Cancer. Nat. Rev. Cancer 2012, 12, 685–698.
- Vyas, S.; Zaganjor, E.; Haigis, M.C. Mitochondria and Cancer. Cell 2016, 166, 555–566.
- Zong, W.-X.; Rabinowitz, J.D.; White, E. Mitochondria and Cancer. Mol. Cell 2016, 61, 667–676.
- Filograna, R.; Mennuni, M.; Alsina, D.; Larsson, N.-G. Mitochondrial DNA Copy Number in Human Disease: The More the Better? FEBS Lett. 2021, 595, 976–1002.
- Grandhi, S.; Bosworth, C.; Maddox, W.; Sensiba, C.; Akhavanfard, S.; Ni, Y.; LaFramboise, T. Heteroplasmic Shifts in Tumor Mitochondrial Genomes Reveal Tissue-Specific Signals of Relaxed and Positive Selection. Hum. Mol. Genet. 2017, 26, 2912–2922.
- Lax, N.Z.; Turnbull, D.M.; Reeve, A.K. Mitochondrial Mutations. Neuroscientist 2011, 17, 645–658.
- Van Gisbergen, M.W.; Voets, A.M.; Starmans, M.H.W.; De Coo, I.F.M.; Yadak, R.; Hoffmann, R.F.; Boutros, P.C.; Smeets, H.J.M.; Dubois, L.; Lambin, P. How Do Changes in the MtDNA and Mitochondrial Dysfunction Influence Cancer and Cancer Therapy? Challenges, Opportunities and Models. Mutat. Res. Rev. Mutat. Res. 2015, 764, 16–30.
- Li, S.; Ou, L.; Zhang, Y.; Shen, F.; Chen, Y. A First-in-Class POLRMT Specific Inhibitor IMT1 Suppresses Endometrial Carcinoma Cell Growth. Cell Death Dis. 2023, 14, 152.
- Graves, P.R.; Aponte-Collazo, L.J.; Fennell, E.M.J.; Graves, A.C.; Hale, A.E.; Dicheva, N.; Herring, L.E.; Gilbert, T.S.K.; East, M.P.; McDonald, I.M.; et al. Mitochondrial Protease ClpP Is a Target for the Anticancer Compounds ONC201 and Related Analogues. ACS Chem. Biol. 2019, 14, 1020–1029.
- Fennell, E.M.J.; Aponte-Collazo, L.J.; Wynn, J.D.; Drizyte-Miller, K.; Leung, E.; Greer, Y.E.; Graves, P.R.; Iwanowicz, A.A.; Ashamalla, H.; Holmuhamedov, E.; et al. Characterization of TR-107, a Novel Chemical Activator of the Human Mitochondrial Protease ClpP. Pharmacol. Res. Perspect. 2022, 10, e00993.
- Mabanglo, M.F.; Wong, K.S.; Barghash, M.M.; Leung, E.; Chuang, S.H.W.; Ardalan, A.; Majaesic, E.M.; Wong, C.J.; Zhang, S.; Lang, H.; et al. Potent ClpP Agonists with Anticancer Properties Bind with Improved Structural Complementarity and Alter the Mitochondrial N-Terminome. Structure 2022, 31, 185–200.
- Wang, P.; Zhang, T.; Wang, X.; Xiao, H.; Li, H.; Zhou, L.-L.; Yang, T.; Wei, B.; Zhu, Z.; Zhou, L.; et al. Aberrant Human ClpP Activation Disturbs Mitochondrial Proteome Homeostasis to Suppress Pancreatic Ductal Adenocarcinoma. Cell Chem. Biol. 2022, 29, 1396–1408.e8.
- Allen, J.E.; Krigsfeld, G.; Patel, L.; Mayes, P.A.; Dicker, D.T.; Wu, G.S.; El-Deiry, W.S. Identification of TRAIL-Inducing Compounds Highlights Small Molecule ONC201/TIC10 as a Unique Anti-Cancer Agent That Activates the TRAIL Pathway. Mol. Cancer 2015, 14, 99.
- Fennell, E.M.J.; Aponte-Collazo, L.J.; Pathmasiri, W.; Rushing, B.R.; Barker, N.K.; Partridge, M.C.; Li, Y.-Y.; White, C.A.; Greer, Y.E.; Herring, L.E.; et al. Multi-Omics Analyses Reveal ClpP Activators Disrupt Essential Mitochondrial Pathways in Triple-Negative Breast Cancer. Front. Pharmacol. 2023, 14, 1136317.
- Kline, C.L.B.; Van den Heuvel, A.P.J.; Allen, J.E.; Prabhu, V.V.; Dicker, D.T.; El-Deiry, W.S. ONC201 Kills Solid Tumor Cells by Triggering an Integrated Stress Response Dependent on ATF4 Activation by Specific EIF2α Kinases. Sci. Signal. 2016, 9, ra18.
- Ray, J.E.; Ralff, M.D.; Jhaveri, A.; Zhou, L.; Dicker, D.T.; Ross, E.A.; El-Deiry, W.S. Antitumorigenic Effect of Combination Treatment with ONC201 and TRAIL in Endometrial Cancer in Vitro and in Vivo. Cancer Biol. Ther. 2021, 22, 554–563.
- Yuan, X.; Kho, D.; Xu, J.; Gajan, A.; Wu, K.; Wu, G.S. ONC201 Activates ER Stress to Inhibit the Growth of Triple-Negative Breast Cancer Cells. Oncotarget 2017, 8, 21626–21638.
- Allen, J.E.; Crowder, R.; El-Deiry, W.S. First-in-Class Small Molecule ONC201 Induces DR5 and Cell Death in Tumor but Not Normal Cells to Provide a Wide Therapeutic Index as an Anti-Cancer Agent. PLoS ONE 2015, 10, e0143082.
- Hillen, H.S.; Morozov, Y.I.; Sarfallah, A.; Temiakov, D.; Cramer, P. Structural Basis of Mitochondrial Transcription Initiation. Cell 2017, 171, 1072–1081.e10.
- Shutt, T.E.; Bestwick, M.; Shadel, G.S. The Core Human Mitochondrial Transcription Initiation Complex. Transcription 2011, 2, 55–59.
- Kaufman, B.A.; Durisic, N.; Mativetsky, J.M.; Costantino, S.; Hancock, M.A.; Grutter, P.; Shoubridge, E.A. The Mitochondrial Transcription Factor TFAM Coordinates the Assembly of Multiple DNA Molecules into Nucleoid-like Structures. Mol. Biol. Cell 2007, 18, 3225–3236.
- Kukat, C.; Wurm, C.A.; Spåhr, H.; Falkenberg, M.; Larsson, N.-G.; Jakobs, S. Super-Resolution Microscopy Reveals That Mammalian Mitochondrial Nucleoids Have a Uniform Size and Frequently Contain a Single Copy of MtDNA. Proc. Natl. Acad. Sci. USA 2011, 108, 13534–13539.
- Bonekamp, N.A.; Jiang, M.; Motori, E.; Villegas, R.G.; Koolmeister, C.; Atanassov, I.; Mesaros, A.; Park, C.B.; Larsson, N.-G. High Levels of TFAM Repress Mammalian Mitochondrial DNA Transcription in Vivo. Life Sci. Alliance 2021, 4, 11.
- Lodeiro, M.F.; Uchida, A.; Bestwick, M.; Moustafa, I.M.; Arnold, J.J.; Shadel, G.S.; Cameron, C.E. Transcription from the Second Heavy-Strand Promoter of Human MtDNA Is Repressed by Transcription Factor A in Vitro. Proc. Natl. Acad. Sci. USA 2012, 109, 6513–6518.
- Salem, A.F.; Whitaker-Menezes, D.; Howell, A.; Sotgia, F.; Lisanti, M.P. Mitochondrial Biogenesis in Epithelial Cancer Cells Promotes Breast Cancer Tumor Growth and Confers Autophagy Resistance. Cell Cycle 2012, 11, 4174–4180.
- Sotgia, F.; Whitaker-Menezes, D.; Martinez-Outschoorn, U.E.; Salem, A.F.; Tsirigos, A.; Lamb, R.; Sneddon, S.; Hulit, J.; Howell, A.; Lisanti, M.P. Mitochondria “Fuel” Breast Cancer Metabolism: Fifteen Markers of Mitochondrial Biogenesis Label Epithelial Cancer Cells, but Are Excluded from Adjacent Stromal Cells. Cell Cycle 2012, 11, 4390–4401.
- Han, Q.-C.; Zhang, X.-Y.; Yan, P.-H.; Chen, S.-F.; Liu, F.-F.; Zhu, Y.-R.; Tian, Q. Identification of Mitochondrial RNA Polymerase as a Potential Therapeutic Target of Osteosarcoma. Cell Death Discov. 2021, 7, 393.
- Zhou, T.; Sang, Y.-H.; Cai, S.; Xu, C.; Shi, M. The Requirement of Mitochondrial RNA Polymerase for Non-Small Cell Lung Cancer Cell Growth. Cell Death Dis. 2021, 12, 751.
- Wang, Y.; Ou, L.; Li, X.; Zheng, T.; Zhu, W.; Li, P.; Wu, L.; Zhao, T. The Mitochondrial RNA Polymerase POLRMT Promotes Skin Squamous Cell Carcinoma Cell Growth. Cell Death Discov. 2022, 8, 347.
- Sarrazin, C.; Hézode, C.; Zeuzem, S.; Pawlotsky, J.-M. Antiviral Strategies in Hepatitis C Virus Infection. J. Hepatol. 2012, 56, S88–S100.
- Arnold, J.J.; Smidansky, E.D.; Moustafa, I.M.; Cameron, C.E. Human Mitochondrial RNA Polymerase: Structure–Function, Mechanism and Inhibition. Biochim. Et Biophys. Acta (BBA)—Gene Regul. Mech. 2012, 1819, 948–960.
- Home—ClinicalTrials.Gov. Available online: https://clinicaltrials.gov/ (accessed on 27 January 2023).
- Arrillaga-Romany, I.; Odia, Y.; Prabhu, V.V.; Tarapore, R.S.; Merdinger, K.; Stogniew, M.; Oster, W.; Allen, J.E.; Mehta, M.; Batchelor, T.T.; et al. Biological Activity of Weekly ONC201 in Adult Recurrent Glioblastoma Patients. Neuro Oncol. 2020, 22, 94–102.
- Stein, M.N.; Bertino, J.R.; Kaufman, H.L.; Mayer, T.; Moss, R.; Silk, A.; Chan, N.; Malhotra, J.; Rodriguez, L.; Aisner, J.; et al. First-in-Human Clinical Trial of Oral ONC201 in Patients with Refractory Solid Tumors. Clin. Cancer Res. 2017, 23, 4163–4169.
- Fei, X.; Bell, T.A.; Jenni, S.; Stinson, B.M.; Baker, T.A.; Harrison, S.C.; Sauer, R.T. Structures of the ATP-Fueled ClpXP Proteolytic Machine Bound to Protein Substrate. eLife 2020, 9, e52774.
- Szczepanowska, K.; Trifunovic, A. Mitochondrial Matrix Proteases: Quality Control and Beyond. FEBS J. 2022, 289, 7128–7146.
- Ishizawa, J.; Zarabi, S.F.; Davis, R.E.; Halgas, O.; Nii, T.; Jitkova, Y.; Zhao, R.; St-Germain, J.; Heese, L.E.; Egan, G.; et al. Mitochondrial ClpP-Mediated Proteolysis Induces Selective Cancer Cell Lethality. Cancer Cell 2019, 35, 721–737.e9.
- Mabanglo, M.F.; Bhandari, V.; Houry, W.A. Substrates and Interactors of the ClpP Protease in the Mitochondria. Curr. Opin. Chem. Biol. 2022, 66, 102078.
- Greer, Y.E.; Porat-Shliom, N.; Nagashima, K.; Stuelten, C.; Crooks, D.; Koparde, V.N.; Gilbert, S.F.; Islam, C.; Ubaldini, A.; Ji, Y.; et al. ONC201 Kills Breast Cancer Cells in Vitro by Targeting Mitochondria. Oncotarget 2018, 9, 18454–18479.
- Zhang, J.; Luo, B.; Sui, J.; Qiu, Z.; Huang, J.; Yang, T.; Luo, Y. IMP075 Targeting ClpP for Colon Cancer Therapy in Vivo and in Vitro. Biochem. Pharmacol. 2022, 204, 115232.
- Fan, Y.; Wang, J.; Fang, Z.; Pierce, S.R.; West, L.; Staley, A.; Tucker, K.; Yin, Y.; Sun, W.; Kong, W.; et al. Anti-Tumor and Anti-Invasive Effects of ONC201 on Ovarian Cancer Cells and a Transgenic Mouse Model of Serous Ovarian Cancer. Front. Oncol. 2022, 12, 789450.
- Greer, Y.E.; Hernandez, L.; Fennell, E.M.J.; Kundu, M.; Voeller, D.; Chari, R.; Gilbert, S.F.; Gilbert, T.S.K.; Ratnayake, S.; Tang, B.; et al. Mitochondrial Matrix Protease ClpP Agonists Inhibit Cancer Stem Cell Function in Breast Cancer Cells by Disrupting Mitochondrial Homeostasis. Cancer Res. Commun. 2022, 2, 1144–1161.
- Mishukov, A.; Odinokova, I.; Mndlyan, E.; Kobyakova, M.; Abdullaev, S.; Zhalimov, V.; Glukhova, X.; Galat, V.; Galat, Y.; Senotov, A.; et al. ONC201-Induced Mitochondrial Dysfunction, Senescence-like Phenotype, and Sensitization of Cultured BT474 Human Breast Cancer Cells to TRAIL. Int. J. Mol. Sci. 2022, 23, 15551.
- Ishizawa, J.; Kojima, K.; Chachad, D.; Ruvolo, P.; Ruvolo, V.; Jacamo, R.O.; Borthakur, G.; Mu, H.; Zeng, Z.; Tabe, Y.; et al. ATF4 Induction through an Atypical Integrated Stress Response to ONC201 Triggers P53-Independent Apoptosis in Hematological Malignancies. Sci. Signal. 2016, 9, ra17.
- Jhaveri, A.V.; Zhou, L.; Ralff, M.D.; Lee, Y.S.; Navaraj, A.; Carneiro, B.A.; Safran, H.; Prabhu, V.V.; Ross, E.A.; Lee, S.; et al. Combination of ONC201 and TLY012 Induces Selective, Synergistic Apoptosis in Vitro and Significantly Delays PDAC Xenograft Growth in Vivo. Cancer Biol. 2021, 22, 607–618.
- Rumman, M.; Buck, S.; Polin, L.; Dzinic, S.; Boerner, J.; Winer, I.S. ONC201 Induces the Unfolded Protein Response (UPR) in High- and Low-Grade Ovarian Carcinoma Cell Lines and Leads to Cell Death Regardless of Platinum Sensitivity. Cancer Med. 2021, 10, 3373–3387.
- Tu, Y.; He, J.; Liu, H.; Lee, H.C.; Wang, H.; Ishizawa, J.; Allen, J.E.; Andreeff, M.; Orlowski, R.Z.; Davis, R.E.; et al. The Imipridone ONC201 Induces Apoptosis and Overcomes Chemotherapy Resistance by Up-Regulation of Bim in Multiple Myeloma. Neoplasia 2017, 19, 772–780.
- Prabhu, V.V.; Morrow, S.; Rahman Kawakibi, A.; Zhou, L.; Ralff, M.; Ray, J.; Jhaveri, A.; Ferrarini, I.; Lee, Y.; Parker, C.; et al. ONC201 and Imipridones: Anti-Cancer Compounds with Clinical Efficacy. Neoplasia 2020, 22, 725–744.
- Cole, A.; Wang, Z.; Coyaud, E.; Voisin, V.; Gronda, M.; Jitkova, Y.; Mattson, R.; Hurren, R.; Babovic, S.; Maclean, N.; et al. Inhibition of the Mitochondrial Protease ClpP as a Therapeutic Strategy for Human Acute Myeloid Leukemia. Cancer Cell 2015, 27, 864–876.
- Lycan, T.; Pardee, T.; Petty, W.; Bonomi, M.; Alistar, A.; Lamar, Z.; Isom, S.; Chan, M.; Miller, A.; Ruiz, J. A Phase II Clinical Trial of CPI-613 in Patients with Relapsed or Refractory Small Cell Lung Carcinoma. PLoS ONE 2016, 11, e0164244.
- Brown, J.R.; Chan, D.K.; Shank, J.J.; Griffith, K.A.; Fan, H.; Szulawski, R.; Yang, K.; Reynolds, R.K.; Johnston, C.; McLean, K.; et al. Phase II Clinical Trial of Metformin as a Cancer Stem Cell–Targeting Agent in Ovarian Cancer. JCI Insight 2020, 5, e133247.
- Mihaylova, M.M.; Shaw, R.J. The AMP-Activated Protein Kinase (AMPK) Signaling Pathway Coordinates Cell Growth, Autophagy, & Metabolism. Nat. Cell Biol. 2011, 13, 1016–1023.
- Ekstrand, M.I.; Falkenberg, M.; Rantanen, A.; Park, C.B.; Gaspari, M.; Hultenby, K.; Rustin, P.; Gustafsson, C.M.; Larsson, N.-G. Mitochondrial Transcription Factor A Regulates MtDNA Copy Number in Mammals. Hum. Mol. Genet. 2004, 13, 935–944.
- Stepanenko, A.A.; Dmitrenko, V.V. HEK293 in Cell Biology and Cancer Research: Phenotype, Karyotype, Tumorigenicity, and Stress-Induced Genome-Phenotype Evolution. Gene 2015, 569, 182–190.
- Shen, C.; Gu, M.; Song, C.; Miao, L.; Hu, L.; Liang, D.; Zheng, C. The Tumorigenicity Diversification in Human Embryonic Kidney 293 Cell Line Cultured in Vitro. Biologicals 2008, 36, 263–268.
- Wedam, R.; Greer, Y.E.; Wisniewski, D.J.; Weltz, S.; Kundu, M.; Voeller, D.; Lipkowitz, S. Targeting Mitochondria with ClpP Agonists as a Novel Therapeutic Opportunity in Breast Cancer. Cancers 2023, 15, 1936.
- Mennuni, M.; Filograna, R.; Felser, A.; Bonekamp, N.A.; Giavalisco, P.; Lytovchenko, O.; Larsson, N.-G. Metabolic Resistance to the Inhibition of Mitochondrial Transcription Revealed by CRISPR-Cas9 Screen. EMBO Rep. 2022, 23, e53054.
