1. Introduction
Kidney cancer is the 14th most common cancer worldwide, and its incidence has continued to increase in recent years
[1]. To date, more than 0.4 million new cases of kidney cancer are diagnosed each year
[2][3][2,3]. Among them, more than 85% of patients present with renal cell carcinoma (RCC)
[4]. Based on the traditional histopathological classification, RCC can be divided into three main categories: clear cell carcinoma (ccRCC, 75%), papillary renal cell carcinoma (PRCC, 15–20%), and chromophobe cell renal carcinoma (chRCC, 5%)
[5]. Studies in recent years have found that RCC mostly occurs in older men
[6] and most cases are localized tumors, with only 17% of RCC patients having distant metastases at the time of diagnosis, which are mainly found in lung, bone, liver, lymph nodes, and adrenal gland
[1][4][1,4]. In 2020, a statistic by Padala SA showed that the 5-year survival rate for kidney cancer patients with metastatic disease was only 12%
[7]. Currently, there are more and more treatment modalities for patients with metastatic RCC (mRCC), with targeted therapies and immune checkpoint inhibitor-based immunotherapy gradually proving to be effective in the treatment of patients with mRCC and the survival rate of those patients greatly improving recently
[8][9][8,9].
Epigenetic alterations are considered to be a hallmark of cancer
[10]. However, recent studies found that RCC has multiple molecular alterations, such as DNA methylation and micro-RNA alterations in ccRCC, which could greatly affect the biological progression of these tumors
[11][12][11,12]. The 2022 World Health Organization (WHO) classification of pathological kidney tumors added new histopathological subtypes, including molecularly defined RCC
[5][8][5,8]. It includes transcription factor binding to
IGHM enhancer 3 (
TFE3)-rearranged renal cell carcinomas, transcription factor
EB (
TFEB)-altered renal cell carcinomas, elongin C (
ELOC)-mutated renal cell carcinoma, fumarate hydratase (
FH)-deficient renal cell carcinoma, succinate dehydrogenase (
SDH)-deficient renal cell carcinoma, anaplastic lymphoma kinase (
ALK)-rearranged renal cell carcinomas, and
SWI/SNF-related, matrix-associated, actin-dependent regulator of chromatin subfamily B member 1 (
SMARCB1)-deficient renal medullary carcinoma (see
Table 1)
[5]. These different molecularly defined histopathological subtypes of RCC are easily confused and may lead to suboptimal treatment outcomes as a result of misdiagnoses
[12].
Table 1. Genes of molecularly defined renal cell carcinoma and associated clinical syndromes.
2. TFE3-Rearranged Renal Cell Carcinomas
Transcription factor binding to
IGHM enhancer 3 (
TFE3) is an important regulator of the immune system and has now been shown to cooperate with transcription factor EB (
TFEB) to control and regulate carbohydrate and lipid metabolism and mitochondrial homeostasis
[13]. The
TFE3/TFEB rearrangement renal cell carcinoma is characterized by translocations involving the
TFE3 and
TFEB genes. They are both derived from the microphthalmia transcription (
MiT) family of heterotopic RCC according to the 2016 version of the WHO classification. The
MiT subfamily of transcription factors includes
TFE3,
TFEB,
TFEC, and
MITF [14].
TFE3- and
TFEB-rearranged RCC accounts for 1–4% of the newly diagnosed adult patients
[15]. Recent studies have shown that
TFE3/TFEB-rearranged RCC can be frequently detected in children
[16]. In adults RCC patients,
TFE3 ectopic fusions with chaperone genes are more commonly seen
[17], and there are no significant prognostic gender differences
[15] (
Figure 1). This ectopic fusion with a chaperone gene and the decreased immunity in adults
TFE3-rearranged RCC patients cause them to have a potentially more aggressive course compared to the pediatric patients
[16]. Current studies suggest that previous exposure to cytotoxic chemotherapy might be a predisposing factor
[18].
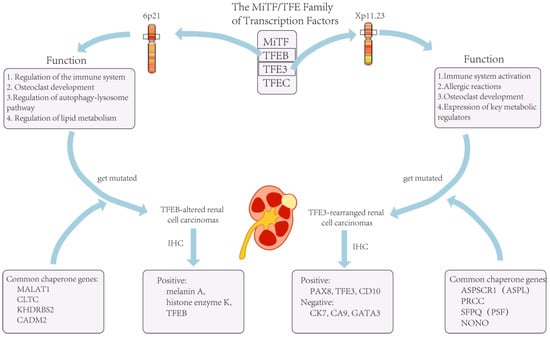
Figure 1. The role of TFE3 in the organism and tumors caused by its mutation.
The list of chaperone genes has been growing and evolving, with more than a dozen having been reported
[18]. The three most common translocations currently include a fusion of the
PRCC and
TFE3 genes, a fusion of the
ASPL (
ASPSCR1) and
TFE3 genes, and a fusion of the
SFPQ and
TFE3 genes
[14]. In addition to this, there are also genes such as
CLTC,
PARP14,
RBM10,
NONO, and
MED15 that can be fused with ectopic
TFE3 [19]. However, current studies suggest that different chaperone genes may exhibit different oncological behaviors and tumor morphologies, and these features vary depending on the type of the involved chaperone genes
[18]. For example,
TFE3 is more likely to exhibit lymph node metastasis when fused with
PRCC than when fused with
ASPSCR3 [20].
In terms of histopathological morphology, the characteristics of
TFE3 fusion usually presents with transparent eosinophils, a papillary architecture, and psammoma bodies under the microscope
[17][21][17,21]. However, due to chaperone genes, RCC with
TFE3 rearrangement may also resemble other types of RCC, including ccRCC, PRCC, and epithelioid vascular smooth muscle lipoma
[14]. Therefore, attention should be paid and the impact of the genes that are fused with should be determined as much as possible both in the diagnosis and in the treatment of
TFE3-rearranged renal cell carcinoma.
When facing
TFE3-rearranged renal cell carcinoma, immunohistochemistry (IHC) is the most commonly used examination for diagnosis
[13]. If IHC is not used at the time of diagnosis, a large proportion of TFE3-rearranged renal cell carcinomas are likely to be misdiagnosed as ccRCC
[19]. For most other types of RCC, the positive IHC markers are
cytokeratin 7 (
CK7),
carbonic anhydrase 9 (
CA9), and
GATA3. However, these are not expressed in
TFE3-rearranged RCC and are usually positive for
histone K [19]. In a recent review article, IHC data from nearly 400 cases of
TFE3-rearranged RCC patients were analyzed, and the biomarkers with the highest probability of positivity were found to be
PAX8 (100%),
TFE3 (95%),
CD10 (89%), and
achromatase (82%)
[22]. However,
TFE3-rearranged RCC did not always exhibit
TFE3 overexpression, and lower
TFE3 expression at the time of detection often resulted in false-positive or false-negative results; thus, this could limit the sensitivity and specificity of IHC for detecting
TFE3-rearranged RCC
[23]. In addition to this, the accuracy of IHC might be affected by the technique and be influenced by the formalin fixation time
[18]. On the other hand, there has been no consensus or standardized guidelines regarding the judgment of
TFE3 staining results
[18], and different pathologists might give completely opposite judgments if specimens show heterogeneous or focal staining. Lee HJ et al. used tissue specimens from 303 RCC patients for IHC testing and found that 23.2% of IHC-negative
TFE3 tumors were eventually diagnosed as
TFE3-rearranged RCC
[24]. Therefore, in clinical practice, a negative
TFE3 IHC result alone did not exclude the possibility of a
TFE3-rearranged RCC case. Thus, in some cases, a combination of clinical presentation and other examination results might be needed.
The current literature suggests that the detection of
TFE3 gene rearrangements by fluorescence in situ hybridization (FISH) is more sensitive and advantageous in experimental manipulation than traditional IHC
[23], and its results are more stable in formalin-fixed tissues
[14]. Therefore, FISH is currently considered as the gold standard for the diagnosis of
TFE3-rearranged RCC
[23]. However, some chaperone genes, such as
NONO,
RBM10, and
GRIPAP1, after fusing with
TFE3, may not be detected by traditional FISH assays for significant
TFE3-positive results.
[18]. In addition, similar to IHC testing, the current standard definition of a positive FISH result varies widely among laboratories, from as low as 10% up to 30%
[17]. These results suggest that, although the FISH test is currently the gold standard for the diagnosis of
TFE3-rearranged RCC, in clinical practice, it should be carefully used together with other test results. For example, the previously mentioned IHC and FISH tests should be considered along with the option of gene probes or alternative molecular techniques
[18]. In fact, FISH cannot provide information about fused genes, so in order to further confirm the diagnosis in clinical practice, RNA sequencing is often used to identify the gene involved in the translocation
[17]. Recently,
TRIM63 determination by RNA in situ hybridization (RNA-ISH) was proposed as an alternative diagnostic tool for
TFE3- and
TFEB-rearranged RCC
[25], but no strong evidence is available from in vitro studies.
Due to the rarity of
TFE3-rearranged RCC and the fact that it has not been previously considered as a specific tumor subtype, there are no treatment recommendations for it to date
[18]. Most previous treatment regimens are consistent with those for patients with ccRCC; however, due to recent developments in detection technology, its diagnosis has become more accurate, similarly to the detection of ccRCC. More importantly, drugs that normally treat ccRCC may not be effective against
TFE3-rearranged RCC
[16]. Additionally, Aldera AP et al. found that patients with
TFE3-rearranged RCC may develop metastases within 20–30 years after diagnosis, so such patients may also need long-term clinical follow-up
[26].
3. TFEB-Altered Renal Cell Carcinomas
As previously stated,
TFEB-altered RCC has been included in the
MiTF-translocated carcinoma family and
TFEB-overexpressing renal tumors were initially identified in pediatric patients. Nowadays, with the availability of accurate examinations, more and more adult RCC patients are diagnosed with
TFEB-altered RCC
[27]. Nevertheless, the number of
TFEB-altered RCC cases is still much lower than for
TFE3-rearranged RCC
[5]. There are two types of
TFEB-altered RCC, including
TFEB-rearranged RCC and
TFEB-amplified RCC. The
TFEB gene in
TFEB-rearranged RCC is located on chromosome 6 and is most often translocated into chromosome 11, fusing with the
MALAT1 gene. Therefore, it was previously called t(6;11) RCC
[19]. In the last few years, researchers have identified cases of RCC related to
TFEB amplification, and after further testing and analysis, it was found that both genetic alteration patterns could co-exist in one case
[5]. Due to the rarity of the disease, there are few studies on the distinction between different subtypes of
TFEB-altered RCC, and current case studies show that the mean age of diagnosis for
TFEB-amplified RCC is 62.5–64 years, while the mean age of diagnosis for
TFEB translocated RCC is 32.8–34 years
[28][29][28,29].
Similar to
TFE3-rearranged RCC,
TFEB can also be ectopically fused to chaperone genes
[28] (
Figure 1). Furthermore, for
TFEB-amplified RCC, in addition to the possible elevated expression of
TFEB, they are often accompanied by the amplification of other oncogenes, such as vascular endothelial growth factor A (
VEGFA) and G1 S specific cyclin D3 (
CCND3)
[30]. It has been shown that these two genes are associated with aggressive oncological behavior
[27], which would precisely explain the severe clinical symptoms and poor prognosis of patients with
TFEB-amplified RCC.
Although
TFEB genes are altered in
TFEB-altered RCC, the characteristics of tumor growth vary considerably between different patterns of alteration. It has been widely reported that in
TFEB-translocated RCC, the most commonly found morphology is a biphasic growth pattern consisting of large and small tumor cells
[27], with smaller cells around the basement membrane-like structures. In addition to this, extensive hyalinization, a papillary architecture, and a clear cell morphology can be seen
[31]. However, in
TFEB-amplified RCC, this pattern is less common. Gupta S et al. investigated 37 patients with
TFEB-altered RCC and found that nearly half of the patients had renal tubular structures and prominent cytoplasmic eosinophilia of tumor cells in their tumor specimens
[27].
IHC and FISH are commonly used tests to detect
TFEB-altered RCC; however, when assessing whether the
TFEB gene is amplified or translocated, the markers used in the detection are quite similar. For
TFEB-altered RCC, it has been shown that the staining results for both histone K and Melan-A are positive
[31]. Similarly, Gupta S et al. and Wyvekens N et al. studied
TFEB-amplified RCC and
TFEB-translocated RCC, respectively, and they found that both types of tumors typically express melanin A and histone enzyme K. The difference was that tumor cells in
TFEB-amplified RCC were usually diffusely or patchily positive when tested for
TFEB levels
[27]. However, there was also a subset of
TFEB-amplified RCC that had lower
TFEB expression levels than
TFEB-translocated RCC
[29]. Therefore, the type of
TFEB gene alteration cannot be distinguished by a
TFEB-specific assay alone. If a type of
TFEB gene alteration is suspected, it should also be demonstrated using a FISH breakdown test or identified by RNA sequencing with a gene fusion examination
[31]. In clinical practice, such detailed testing and diagnosis is not always necessary for all patients because of the very low incidence of the disease, the high cost of FISH, and the use of sequencing tests.
In addition to this, it has also been found that
TFEB-amplified RCC exhibits a higher tumor aggressiveness than
TFEB-rearranged tumors, and the 5-year survival rate for
TFEB-amplified RCC is only 48%
[32], while
TFEB-translocated RCC progresses more slowly than
TFE3-rearranged RCC. Therefore, in clinical practice, physicians should distinguish
TFEB-amplified RCC from
TFEB-translocated RCC. Since
TFEB-altered RCC has often been previously diagnosed as ccRCC, its current treatment modality still differs little from the standard treatment for patients with ccRCC, which may also contribute to the poor prognostic outcome for patients with
TFEB-altered RCC.