
Video Upload Options
Receptor-like kinases (RLKs) constitute a large group of cell surface receptors that play crucial roles in multiple biological processes. However, the function of most RLKs in plants has not been extensively explored, and much less for the class of cell wall associated kinases (WAKs) and WAK-like kinases (WAKLs). In this study, analyses of developmental expression patterns uncovered a putative role of AtWAKL10 in modulating leaf senescence, which was further investigated at physiological and molecular levels. The expression level of AtWAKL10 increased with the developmental progression and was rapidly upregulated in senescing leaf tissues. The promoter of AtWAKL10 contains various defense and hormone responsive elements, and its expression could be significantly induced by exogenous ABA, JA and SA. Moreover, the loss-of-function atwakl10 mutant showed earlier senescence along the course of natural development and accelerated leaf senescence under darkness and hormonal stresses, while plants overexpressing AtWAKL10 showed an opposite trend.
1. Introduction
Leaf development is a strictly programmed process that includes the early stages of leaf development such as initiation, growth, and maturation, and the later leaf senescence. Due to the sessile lifestyle of plants, leaf senescence is not only regulated by intrinsic and genetic factors, but also affected by various environmental stimuli, such as drought, shade, extreme temperature, and pathogen infections [1][2]. Leaf senescence is also a complicated multi-step process. In contrast to nutrient assimilation at early development stages, leaf senescence is mainly accompanied by highly ordered degradation of organelles and cellular compositions, subsequent remobilization of valuable resources from senescing leaves to vigorous tissues and organs, and eventually, the suspension of photosynthesis and programmed cell death [3][4][5]. Thus, following the initiation of leaf senescence, leaves will experience constant chlorophyll loss, reduced efficiency of photosynthesis, and increased accumulation of reactive oxygen species (ROS) [5][6][7]. To cope with the substantial metabolic loss during leaf senescence, plants have established sophisticated transcriptional regulatory networks and signaling mechanisms to support plant growth and ensure reproduction by stimulating the expression of senescence associated genes (SAGs) [2][8][9].
The initiation of natural leaf senescence is mainly dependent on the age [10], but can also be triggered by various exogenous stimuli. As crucial tissues for photosynthesis, leaves subjected to prolonged light deprivation such as complete darkness show accelerated yellowing, which also exhibit a considerable overlap in differentially expressed genes and signaling pathways with natural senescence [11]. However, it is worth noting that such senescence phenotypes are more suitable for individual and detached leaves but not the whole plant [12][13]. In addition to dark treatment, plant hormones are also commonly used as effective inducers of senescence. Hormone signaling pathways, as a comprehensive mechanism, are involved in regulating both developmental and environmental responses. Many studies have found that the levels of hormones such as abscisic acid (ABA), jasmonic acid (JA) and salicylic acid (SA) are increased in senescing leaves [14][15][16], which may promote senescence by inducing SAGs [17][18]. Correspondingly, the transcription levels of genes encoding key components for hormonal biosynthesis and signal transduction are also altered during leaf senescence [16][19][20].
During leaf senescence, genes involved in catabolism and defense responses undergo large-scale transcriptional reprogramming, which requires hierarchical but also coordinated regulation by multiple transcription factors (TFs) [11][16][21]. It is well exemplified by the regulation of chlorophyll degradation. Specifically, chlorophyll catabolic genes such as NON-YELLOWING1 (NYE1)/STAY-GREEN1 (SGR1) and PHEOPHORBIDE A OXYGENASE (PAO) can be induced by increased ABA accumulation [22]. The ABA biosynthesis gene ALDEHYDE OXIDASE3 (AAO3) is a transcription target of NAC-LIKE, ACTIVATED BY AP3/PI (NAP/ANAC029) in Arabidopsis, while the promoter of AtNAP can be bound by ANAC016, which is also a transcription regulator of NYE1 [23][24]. Moreover, NYE1 and PAO are positively regulated by MYC2, MYC3, and MYC4, three positive TFs of the JA signaling pathway [25], while they are also integrated into the regulatory loop of ethylene-induced chlorophyll degradation with ETHYLENE-INSENSITIVE3 (EIN3) and ORESARA 1 (ORE1)/ANAC092 [26]. In addition to NACs, WRKY TFs also constitute a large set of SAGs, some of which also play a role in other biological processes, typically, hormone mediated defense responses [11]. For example, WRKY6 is upregulated upon pathogen infection and leaf senescence [27][28], and overexpression or lack of WRKY6 facilitates and retards dark-induced senescence [29]. It is also required for SA-mediated leaf senescence, and transcriptionally activated by WRKY46 upon SA treatment [30]. Both WRKY6 and WRKY46 are downstream of NONEXPRESSER OF PR GENES 1 (NRP1), and act synergistically with NRP1 to positively regulate leaf senescence.
Plant receptor-like kinases (RLKs) are the largest serine/threonine protein kinase subfamily, which can mediate extracellular signals and then activate intracellular signaling pathways, thus playing a crucial role in cell-cell communications and plant-pathogen interactions [31][32][33]. Wall associated kinases (WAKs) and WAK-like kinases (WAKLs) are a small subset of RLKs with a total of 26 members in Arabidopsis, which possess a unique extracellular region that harbors several epidermal growth factor (EGF) repeats and is closely associated with the cell wall [34][35]. In addition, the composition of the extracellular domain of each member is also different, which may be due to their different ligand types and regulatory functions [35][36]. WAKs are essential for cell expansion, and some members, such as WAK1, are primarily involved in defense responses and can be up-regulated by fungal pathogens, exogenous SA, MeJA, and systemically acquired resistance (SAR) [37][38][39][40]. AtWAKL10 has similar molecular weight and structural characteristics to WAK1 and has been identified as a twin-domain signaling molecule [41][42]. Its intracellular domain has both kinase activity and functional Guanylyl cyclase (GC) activity, which can catalyze the formation of guanine nucleoside 3’,5’-cyclomonophosphate (cGMP), an important signaling molecule involved in a variety of physiological and biological processes in higher plants [41]. A more recent study also showed that AtWAKL10 could be induced by nitric oxide (NO) and played a positive role in basal defense, effector-triggered immunity and salt stress but negatively regulated drought stress [43].
Leaf senescence is a natural and orderly physiological process, which is of great significance to agricultural production. Under unfavorable conditions, the timely initiation of senescence is beneficial to plant viability and successful regeneration. Here, based on tissue expression pattern analyses on a variety of RLKs, AtWAKL10 was found to be upregulated in senescing leaves. This was further confirmed by quantification of its transcript abundance in leaf tissues of different degrees of senescence. Characterization of the loss-of-function atwakl10 mutant and plants overexpressing AtWAKL10 found that they were sensitive and resistant to natural, dark-, and hormone-induced leaf senescence, respectively. Moreover, several defense and senescence associated WRKYs could bind to the promoter of AtWAKL10, while expression of some WRKYs could also be affected by the levels of AtWAKL10, further indicating that AtWAKL10 was involved in regulating leaf senescence. In addition, deletion and overexpression of AtWAKL10 caused significant changes in expression of cell expansins and defense response genes, respectively, which only occurred during leaf senescence. These results indicated that AtWAKL10 played a negative role in leaf senescence.
2. Functional Domain Analysis and Subcellular Localization of AtWAKL10
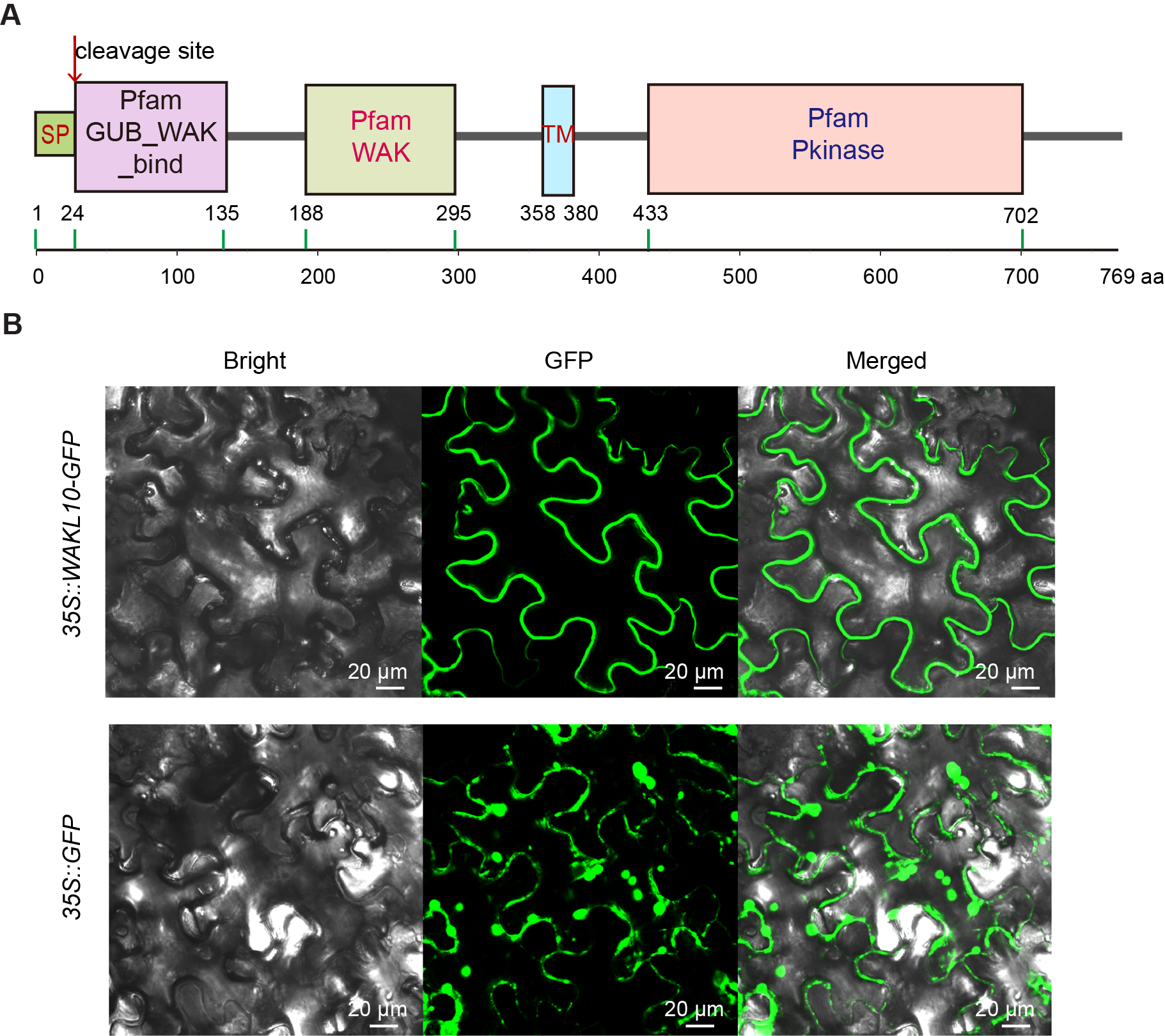

3. AtWAKL10 Was Highly Induced during Leaf Senescence
To identify putative roles of AtWAKL10 in plant development, tissue expression pattern of AtWAKL10 was analyzed through Arabidposis eFB Brower (http://bar.utoronto.ca, accessed on 8 November 2020) and GENEVESTIGATOR v3 (https://genevestigator.com, accessed on 20 December 2020). AtWAKL10 showed relatively low expression in most plant tissues and developmental stages but was clearly upregulated in senescing leaves (Supplementary Figure S1, supplementary could be found in https://www.mdpi.com/1422-0067/22/9/4885#supplementary).
To confirm the expression pattern, rosette leaves 5 and 6 from A. thaliana wild type (Col-0) plants at four different developmental stages were collected, including young leaf stage (S1), non-senescence mature leaf stage (S2), early senescence stage (S3), and late senescence stage (S4) (Figure 2A). A constant decrease in chlorophyll concentrations in leaves from S1 to S4 indicated the progression of leaf senescence (Figure 2B). This was further supported by the expression pattern of typical senescence marker genes (Figure 2C). The expression levels of senescence marker genes PAO and NYE1/SGR1 increased slightly during the transition from young leaves to mature leaves, and then increased continuously and rapidly with the developmental stages (Figure 2C). Correspondingly, the senescence-opposite maker gene, the small subunit of 1,5-ribulose bisphosphate carboxylase oxygenase (RBCS3B) consistently declined (Figure 2C). A sharp decrease in its expression from S2 to S3 stage also indicated the early onset of leaf senescence. It was not surprising that the age-related senescence marker gene SAG12 was specifically and substantially upregulated in leaves at both early and late senescence stages (Figure 2C). For the AtWAKL10 transcript, it showed highly consistent expression patterns with SAG12, which was rapidly induced from the initial leaf senescence (Figure 2C), suggesting that AtWAKL10 may be involved in the regulation of leaf senescence.
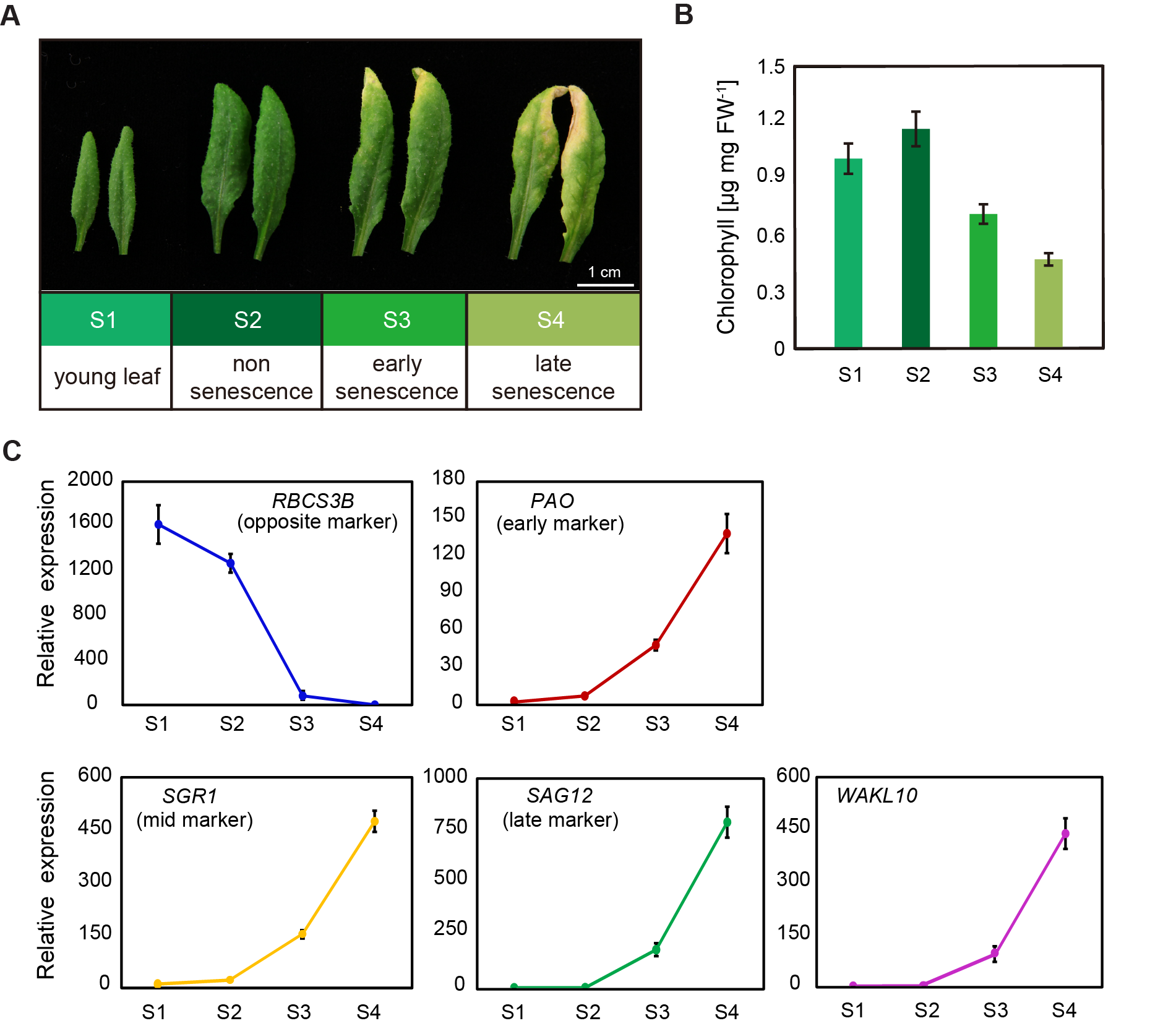
Figure 2. The specifically upregulated expression of AtWAKL10 in leaves of A. thaliana wild type (Col-0) plants during natural senescence. (A) Representative images of fifth and sixth rosette leaves from A. thaliana wild type (Col-0) plants at different developmental stages. S1, young leaf; S2, non-senescence leaves with true leaves fully expanded; S3, early senescent leaf with yellowing leaf area less than 25% from the leaf tip; S4, more than 40% of leaf area yellowing. (B) The chlorophyll concentrations in leaves 5 and 6 at S1, S2, S3, and S4 stages. (C) The transcript abundance of AtWAKL10 and leaf senescence marker genes. Data were given as averages ± SE of three biological replicates. For RBCS3B, gene expression was calculated relative to S4 stage, or otherwise, was relative to S1 stage.
4. AtWAKL10 Negatively Regulates Natural Leaf Senescence
To investigate how AtWAKL10 functions in regulating natural leaf senescence, a homozygous mutant line designated as atwakl10 (SALK_132887) was screened by PCR (Supplementary Figure S2). Sanger sequencing revealed that the mutant contained a T-DNA insertion in the promoter region, specifically, 205 bp upstream of the start codon (Figure 4A, Supplementary Figure S2). Meanwhile, two overexpression lines for AtWAKL10 were generated in the A. thaliana wild type (Col-0) background, designated as AtWAKL10 OE1 and AtWAKL10 OE2 (Figure 4A). Both the knock-out mutant and overexpression lines were confirmed by quantitative Real-time PCR (Figure 4B).
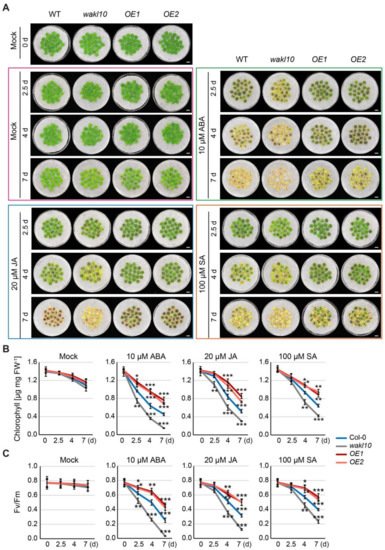
Soil-based phenotyping was performed on the wild type, atwakl10 mutant, and AtWAKL10 OE lines, and no obvious differences in leaf size, leaf number, and flowering time were observed. Then, a closer analysis of developmental progression revealed that natural leaf senescence seemed to occur earlier for the loss-of-function atwakl10 mutant, while overexpression of AtWAKL10 greatly inhibited leaf senescence (Figure 4C). Specifically, compared with the wild type, the 7-week-old atwakl10 mutant had about two more yellowing leaves and a greater degree of chlorophyll loss in the same leaf position, while more than 90% of the true leaves from the AtWAKL10 OE plants remained green (Figure 4D). Again, the rosette leaves from a single plant were divided into four groups based on the leaf age. In agreement with the severe leaf senescence phenotype, leaves of each group from atwakl10 mutant had the lowest chlorophyll concentrations and quantum efficiency of photosystem II (Fv/Fm) (Figure 4E,F). Meanwhile, it was also supported by the higher expression of SAG12 and lower expression of RBCS3B in almost all groups. One exception for the lower expression of SAG12 and SAG113 in the oldest leaves (group1) was due to the loss of leaf vitality [46]. On the other hand, not surprisingly, the values for various physiological parameters and the transcription abundance of senescence marker genes in all groups of AtWAKL10 OE plants were significantly greater than that in wild type (Figure 4E,F,H). In addition, with the course of leaf senescence, ion leakage increases due to disruption of cell membranes and increased membrane permeability [47]. As was shown, the accelerated senescence symptoms of the atwakl10 mutant were accompanied by the higher membrane ion leakage rates (Figure 4G), whereas two overexpression lines always had lower ion leakage rates compared with the wild type. These results further suggested that AtWAKL10 exerted negative control on natural leaf senescence.
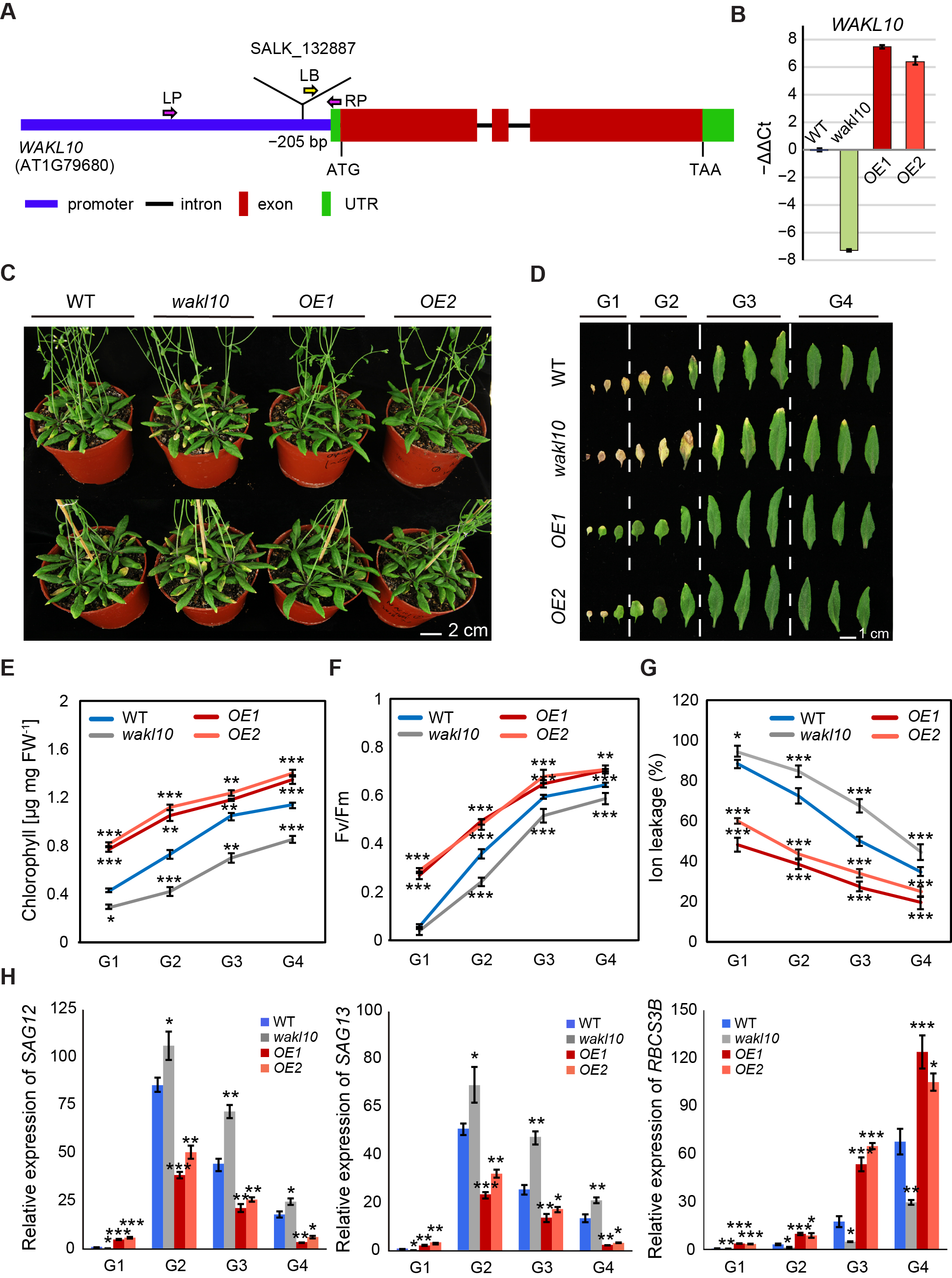
Figure 4. AtWAKL10 negatively regulated natural leaf senescence. (A) Schematic gene model of AtWAKL10. The T-DNA insertion site was shown, and homozygous mutants were screened using specific primers (Supplementary Table S1). LP (left primer), RP (right primer), left border primer (LB), ATG (start codon), TAA (stop codon) and UTR (untranslated region). (B) Quantification of AtWAKL10 in atwakl10 mutant and AtWAKL10 overexpression lines. Expression levels of AtWAKL10 were determined by qRT-PCR and presented as values of −ΔΔCt. Data shown were means ± SE of three independents plants. (C) Phenotypes of 7-week-old wild type (Col-0), atwakl10 mutant and AtWAKL10 overexpression lines (OE1 and OE2), and (D) a series of developmental stages of all leaves as indicated. Rosette leaves were divided into four groups according to the degree of senescence. Quantification of chlorophyll concentrations (E), PSII maximum efficiency (Fv/Fm) (F), ion leakage (G) and relative expression of SAG12, SAG13 and RBCS3B (H) in leaves from each group. Mean values ± SE were shown for three biological replicates. Significant differences (* p < 0.05, ** p < 0.01, and *** p < 0.001) compared with the wild type in each group were determined by Student’s t-test.
5. AtWAKL10 Had a Negative Effect on Dark-Induced Senescence
To further elucidate the roles of AtWAKL10 in leaf senescence, darkness stress was applied to induce leaf senescence. To start, dark-induced senescence was performed on attached leaves 4, 5, and 6 from 25-day-old wild type, atwakl10 mutant, and AtWAKL10 OE plants by covering leaves with aluminum foil paper for 7 d (Figure 5A). Although no differences were observed in chlorophyll concentrations and photosynthetic efficiency (Fv/Fm) among these genotypes under control conditions, the values of these parameters were significantly lower in atwakl10 mutant but higher in both AtWAKL10 OE lines compared with wild type after 7 d of dark treatment (Figure 5B,C). In addition, three senescence associated marker genes, including WRKY22, CYSTEINE PROTEASE 1 (CP1), and SAG12, were upregulated in all genotypes after 7 d of darkness stress, and the corresponding expression levels were always the highest in the atwakl10 mutant but the lowest in both AtWAKL10 OE lines.
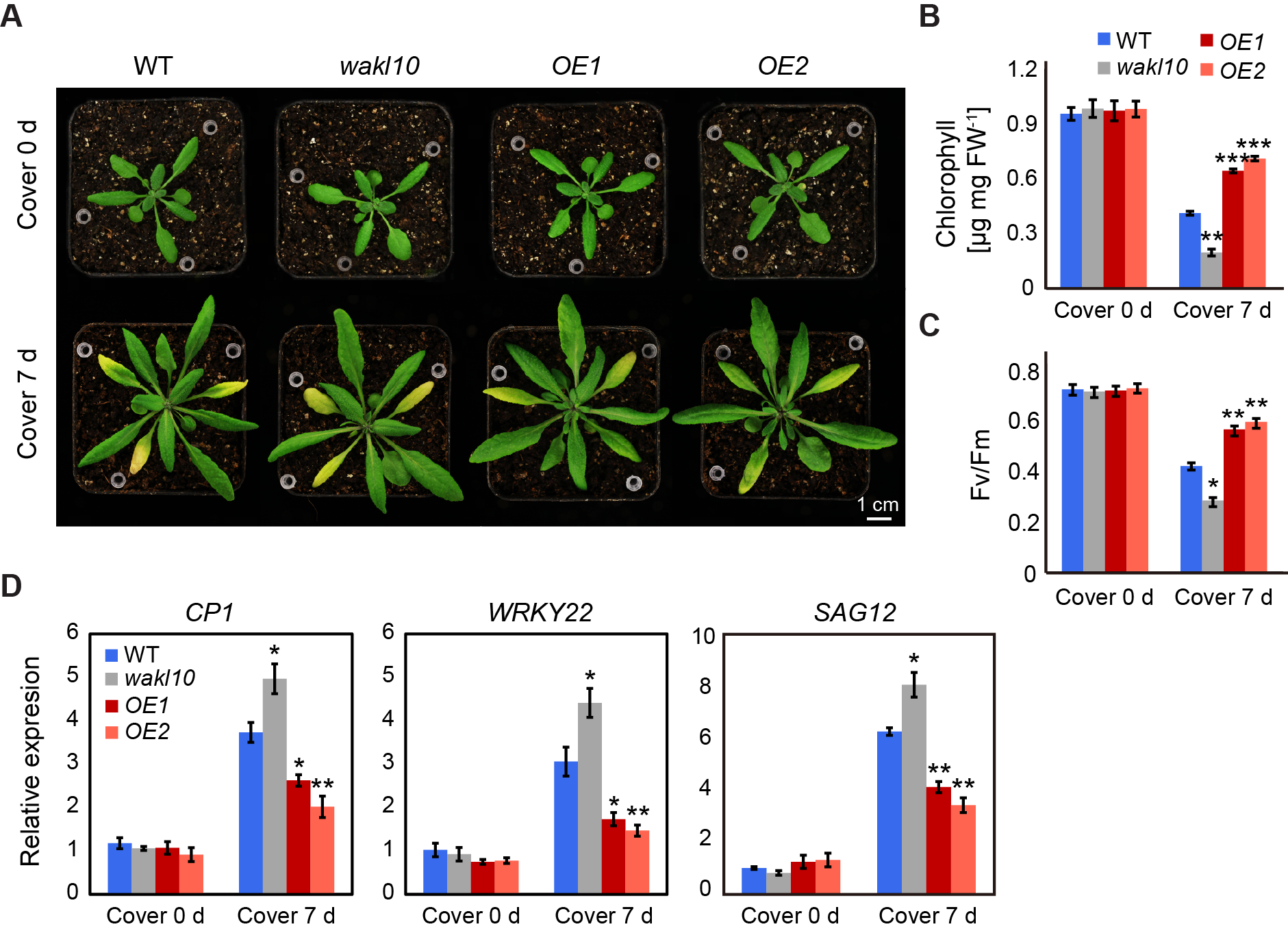
Figure 5. AtWAKL10 effectively delayed dark-induced senescence in attached leaves. (A) Phenotypes of rosette leaves from A. thaliana wild type (Col-0), atwakl10 mutant, and AtWAKL10 overexpression lines under control growth conditions and after 7 d of darkness treatment. The 25-day-old uniformly grown plants were designated as cover 0 d, and those subjected to a follow-up extra 7 d of dark acclimation were indicated as cover 7 d. Attached leaves 4, 5, and 6 were exposed to darkness stress by being covered with aluminum foil paper. The tested leaves were indicated by white pipette tips. Quantification of chlorophyll concentrations (B), chlorophyll fluorescence efficiency (Fv/Fm) (C), and expression levels of dark-induced or natural senescence related genes (D) in tested leaves before and after 7 d of darkness stress. Data were given as averages ± SE of five biological replicates. Significant differences compared with wild type under each condition were indicated (* p < 0.05, ** p < 0.01, and *** p < 0.001, Student’s t-test).
6. Discussion
For the response to unexpected stresses and changing environmental challenges, plants have evolved special mechanisms, such as cell wall sensing systems, to relay external signals to intracellular effectors to trigger defense responses and minimize potential damage. Receptor-like kinases are indispensable sensors that contribute to intercellular communication, cellular signaling and plant-pathogen interactions [31][32][33]. Most of the well characterized RLKs belong to the leucine-rich repeat RLKs (LRR-RLKs) and are mainly involved in tissue and organ development, hormone perception, and stress response [33][46][47]; only a few RLKs are identified to participate in the regulation of leaf senescence [48][49][50][51][52]. Since an appropriate delay of senescence can improve crop yield potential and agronomic traits due to prolonged photosynthesis, exploring more senescence related RLKs and underlying mechanisms can provide practical theoretical guidance for agricultural production. In this study, we found that AtWAKL10, encoding a member of the cell wall-associated class of RLKs, was highly up-regulated in senescing leaves (Figure 1 and Figure 2). Deletion of AtWAKL10 exacerbated age-dependent and dark-induced leaf senescence, with the loss-of-function mutant having higher expression of senescence marker genes and lower chlorophyll content compared with wild type during natural leaf senescence (Figure 3 and Figure 4). In parallel, plants overexpressing AtWAKL10 exhibited the oppositely delayed senescence phenotype, which further confirmed that AtWAKL10 exerted negative control over leaf senescence.
Our results showed that GFP signals from 35S::GFP were cytosolic while signals from 35S::AtWAKL10-GFP were mostly in membrane systems. A prediction through the UniProt program (https://www.uniprot.org/, accessed on 19 April 2021) also shows that AtWAKL10 is localized in the plasma membrane. Moreover, other members of its family such as WAKL4 [53] and WAKL6 [54] in Arabidopsis as well as its potential homologs in Brassica napus [55] were reported to be present on cell membranes. Further analyses are needed to confirm the subcellular localization of AtWAKL10.
WAKs and WAKLs are a special class of RLKs that are divided into four groups based on the sequence and structure similarities in Arabidopsis [35]. AtWAKL10 is one of the six members of Group III with WAKL9, WAKL11, WAKL13, WAKL17, and WAKL18, which have similar structure features to the Group II members (WAKL1-6 and WAKL22). In addition to the protein sequence, some stress- and growth-associated motifs identified in the AtWAKL10 promoter were also present in other WAK members [36][54], suggesting that these members may be involved in similar biological processes and probably have similar transcriptional regulatory control. Typically, a positive role of AtWAKL10 in regulating early defense responses had also been confirmed for WAK group members, with some being greatly up-regulated by fungal and bacterial pathogens, SAR inducing conditions, and defense signaling molecules SA and MeJA [38][39][40][41]. In fact, the case seems to be more complicated for AtWAKL10, which is required to respond to intricate conditions such as convergence of different pathogens and microbes [41], and here, multiple hormones involved in leaf senescence (Figure 8 and Figure 9). Since the interrelated transcription of AtWAKL10 and genes encoding other PM localized receptor kinases was likely to promote the recognition and transduction of external adverse signals and activation of cellular signaling [41][56][57], identifying AtWAKL10-associated proteins and the key transcription factors or cascades for coordinately regulating hormone signals is necessary.
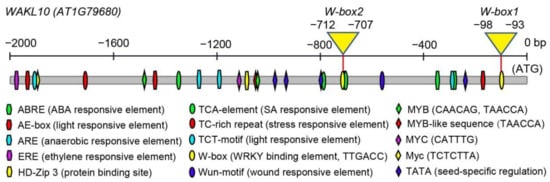
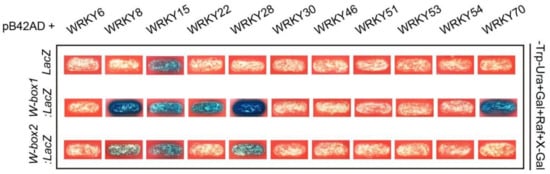
Transcriptomic analyses of leaves of different developmental stages and those under darkness stress have identified numerous WRKY TFs differentially expressed during leaf senescence [16][58][59]. These WRKYs play distinct regulatory roles in leaf senescence. For example, WRKY54 and WRKY70 act as negative regulators, whereas WRKY53 is identified as an accelerator of the onset of senescence [60][61][62]. WRKY22, a target gene of WRKY53, is induced by darkness stress, while WRKY22, similar to WRKY6, exerts positive control over dark-induced senescence [63][64]. WRKY28 contributes to SA biosynthesis and positively regulates age- and light-mediated leaf senescence [65][66][67]. In fact, most senescence-regulating WRKYs are involved in SA-related biological processes, which further support the large overlapping of transcriptomic changes and signaling pathways between SA-mediated stresses and age-dependent leaf senescence [11][68]. Similarly, AtWAKL10 can be induced by pathogen infection and its promoter is rich in WRKY TFBS and SA responsive elements (Figure 7), strengthening the possibility that AtWAKL10 is involved in the regulation of leaf senescence. Previously, several WRKY TFs have been predicted as transcription regulators of AtWAKL10, including WRKY2/3/4/6/10/14/18/26/33/40/56/61/65/71 [36][37], and WRKY28 and WRKY15 were highly co-expressed with AtWAKL10 [35]. These findings are partially in accordance with our results, especially for the WRKY28, exhibiting a strong binding to the W-box1 nearby the start codon (Figure 10). WRKY15 may be a candidate but it showed a weak binding and can be self-activated. WRKY70, a well-known TF involved in defense and stress response [69][70][71][72], is bound to the W-box1 on the AtWAKL10 promoter. Since these multifunctional WRKYs are recruited to assist a variety of biological processes, their coordinated regulation of AtWAKL10 places it as a key node that integrates and responds to signals of different levels [64][66][73][74][75][76]. It could not be ignored that these WRKYs play different regulatory roles in leaf senescence, and some factors such as WRKY54 did not affect its transcriptional expression, but may exert controls by physical interactions, which makes the control of leaf senescence by AtWAKL10 more complicated. In addition, given the limited study scope, more senescence-related WRKYs such as WRKY33 [77], WRKY45 [78], WRKY42 [79], and WRKY75 [80] should be validated.
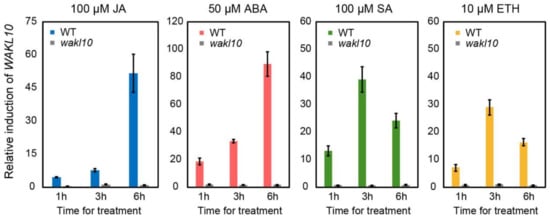
As previously reported, AtWAKL10 was mainly induced by SA-related stimuli [41]. Here, we also showed that it could be significantly activated by multiple hormones, though to varying degrees (Figure 8). Surprisingly, although there was no JA responsive element but four SA responsive elements in the promoter of AtWAKL10, it was maximally induced by ABA and JA but not SA. This was probably attributed to the concentration of the inducer. For example, SA functions in a concentration-dependent manner in which it promotes growth and development at low concentrations but acts as a stress inducer at high concentrations [81][82]. In addition, unlike the continuous up-regulation of AtWAKL10 under ABA and JA treatments, SA was inclined to trigger short-term and periodic response, which was consistent with previous studies [43].
Leaf senescence is a degenerative process with large-scale degradation of organelles and structural components, leading to the destruction of the cell wall. Therefore, maintaining leaf cell elongation and cell wall structure is an effective means to prevent senescence. It was commonly accepted that WAKs were crucial for cell elongation, while decreased levels of WAKs caused dose-dependent inhibition of cell elongation and the resulting smaller rosette leaves [36][37]. However, there is so far no experimental evidence that AtWAKL10 is involved in regulating these processes. Cell elongation is controlled by different classes of wall proteins, among which expansins are mostly characterized factors that play crucial roles in organ growth, cell elongation, cell wall structure, and extensibility [83][84][85]. Here, deletion and overexpression of AtWAKL10 did not cause obvious alterations in the size and number of rosette leaves, organ morphology, bolting time, and expression of expansins compared with the wild type during the non-senescence developmental stages [43] (Supplementary Table S3). However, a significant down-regulation of expansins was specifically observed in the atwakl10 mutant upon leaf senescence, indicating that AtWAKL10 may be required for the expression of expansins, and such interconnection seemed to be age-dependent. Since the expression of expansins was not changed because of the upregulation of AtWAKL10, they could not be co-expressed with AtWAKL10. Unlike the WAKs and group II WAKL members, which could mediate common cell growth and organ development, the regulation of cell elongation and extension by AtWAKL10 was more like a stress responsive mechanism. Specifically, AtWAKL10 has been reported to mediate responses to pathogen infection and drought stress [43]. Both processes involve disruption of cell walls that was caused by invasion of external pathogens and altered cell tension, respectively. Expansins, as core regulators of cell wall extension, play crucial roles in resisting these stress conditions [86][87][88]. Still notably, there was a significant increase in expression of ABI5 in atwakl10 mutant during leaf senescence. As a core ABA signaling component, ABI5 is always upregulated to cope with drought stress [89][90][91][92]. Thereby, our results may provide a potential mechanism for the elevated resistance of the atwakl10 mutant to drought stress [43].
Another significant senescence-specific change was the consistent up-regulation of defense response genes in AtWAKL10 OE lines. Most of these genes such as PR1, PR5, CHI, and WRKY30/38/50/51 positively respond to both exogenous SA treatment and pathogen-induced endogenous SA biosynthesis, indicating that SA signaling is involved in AtWAKL10 mediated senescence regulatory mechanisms. However, it was interesting that genes encoding key enzymes for SA biosynthesis, e.g., Isochorismate Synthase 1 (ICS1) [93] and PHE AMMONIA LYASE 1 (PAL1), and core components for SA reception and signal transduction such as NONEXPRESSER OF PR GENES 1 (NPR1) [94], TGACG SEQUENCE-SPECIFIC BINDING PROTEIN 1 (TGA1), and TGA6 [95], were not differentially expressed compared with wild type. Since accumulation of SA can promote the transition from growth to senescence, it was possible that overexpression of AtWAKL10 triggered the downstream defense response, which acted as a feedback signal to inhibit excessive accumulation and signal transduction of SA. Notably, NAC6/ORE1/ANAC092, as the signal transduction junction of SA and ethylene to promote leaf senescence [26][96], was significantly down-regulated by elevated accumulation of AtWAKL10. Similarly, WRKY70 could bind to the promoter of AtWAKL10 (Figure 7), but the lack of AtWAKL10 during leaf senescence led to obvious down-regulation of WRKY70, which can generally be activated by SA [71]. The specific regulatory mechanism for these changes remains to be further studied.
References
- Gan, S.; Amasino, R.M. Making Sense of Senescence (Molecular Genetic Regulation and Manipulation of Leaf Senescence). Plant Physiol. 1997, 113, 313–319.
- Lim, P.O.; Kim, H.J.; Gil Nam, H. Leaf Senescence. Annu. Rev. Plant Biol. 2007, 58, 115–136.
- Himelblau, E.; Amasino, R.M. Nutrients mobilized from leaves of Arabidopsis thaliana during leaf senescence. J. Plant Physiol. 2001, 158, 1317–1323.
- Humbeck, K.; Quast, S.; Krupinska, K. Functional and molecular changes in the photosynthetic apparatus during senescence of flag leaves from field-grown barley plants. Plant Cell Environ. 1996, 19, 337–344.
- Buchanan-Wollaston, V.; Earl, S.; Harrison, E.; Mathas, E.; Navabpour, S.; Page, T.; Pink, D. The molecular analysis of leaf senescence—A genomics approach. Plant Biotechnol. J. 2003, 1, 3–22.
- Woo, H.R.; Kim, J.H.; Gil Nam, H.; Lim, P.O. The Delayed Leaf Senescence Mutants of Arabidopsis, ore1, ore3, and ore9 are Tolerant to Oxidative Stress. Plant Cell Physiol. 2004, 45, 923–932.
- Ougham, H.J.; Hörtensteiner, S.; Armstead, I.P.; Donnison, I.S.; King, I.P.; Thomas, H.; Mur, L.A.J. The control of chlorophyll catabolism and the status of yellowing as a biomarker of leaf senescence. Plant Biol. 2008, 10, 4–14.
- Guo, Y.; Gan, S. Leaf Senescence: Signals, Execution, and Regulation. Curr. Top. Dev. Biol. 2005, 71, 83–112.
- Hopkins, M.; Taylor, C.; Liu, Z.; Ma, F.; McNamara, L.; Wang, T.-W.; Thompson, J.E. Regulation and execution of molecular disassembly and catabolism during senescence. New Phytol. 2007, 175, 201–214.
- Buchanan-Wollaston, V. The molecular biology of leaf senescence. J. Exp. Bot. 1997, 48, 181–199.
- Buchanan-Wollaston, V.; Page, T.; Harrison, E.; Breeze, E.; Lim, P.O.; Gil Nam, H.; Lin, J.-F.; Wu, S.-H.; Swidzinski, J.; Ishizaki, K.; et al. Comparative transcriptome analysis reveals significant differences in gene expression and signalling pathways between developmental and dark/starvation-induced senescence in Arabidopsis. Plant J. 2005, 42, 567–585.
- Liebsch, D.; Keech, O. Dark-induced leaf senescence: New insights into a complex light-dependent regulatory pathway. New Phytol. 2016, 212, 563–570.
- Weaver, L.M.; Amasino, R.M. Senescence is induced in individually darkened Arabidopsis leaves but inhibited in whole darkened plants. Plant Physiol. 2001, 127, 876–886.
- Zhang, K.; Gan, S.-S. An Abscisic Acid-AtNAP Transcription Factor-SAG113 Protein Phosphatase 2C Regulatory Chain for Controlling Dehydration in Senescing Arabidopsis Leaves. Plant Physiol. 2012, 158, 961–969.
- Sarwat, M.; Naqvi, A.R.; Ahmad, P.; Ashraf, M.; Akram, N.A. Phytohormones and microRNAs as sensors and regulators of leaf senescence: Assigning macro roles to small molecules. Biotechnol. Adv. 2013, 31, 1153–1171.
- Breeze, E.; Harrison, E.; McHattie, S.; Hughes, L.; Hickman, R.; Hill, C.; Kiddle, S.; Kim, Y.-S.; Penfold, C.A.; Jenkins, D.; et al. High-Resolution Temporal Profiling of Transcripts during Arabidopsis Leaf Senescence Reveals a Distinct Chronology of Processes and Regulation. Plant Cell 2011, 23, 873–894.
- Hu, Y.; Jiang, Y.; Han, X.; Wang, H.; Pan, J.; Yu, D. Jasmonate regulates leaf senescence and tolerance to cold stress: Crosstalk with other phytohormones. J. Exp. Bot. 2017, 68, 1361–1369.
- He, Y.; Fukushige, H.; Hildebrand, D.F.; Gan, S. Evidence Supporting a Role of Jasmonic Acid in Arabidopsis Leaf Senescence. Plant Physiol. 2002, 128, 876–884.
- Van Der Graaff, E.; Schwacke, R.; Schneider, A.; DeSimone, M.; Flügge, U.-I.; Kunze, R. Transcription Analysis of Arabidopsis Membrane Transporters and Hormone Pathways during Developmental and Induced Leaf Senescence. Plant Physiol. 2006, 141, 776–792.
- Kong, Z.; Li, M.; Yang, W.; Xu, W.; Xue, Y. A Novel Nuclear-Localized CCCH-Type Zinc Finger Protein, OsDOS, Is Involved in Delaying Leaf Senescence in Rice. Plant Physiol. 2006, 141, 1376–1388.
- Pruneda-Paz, J.L.; Breton, G.; Nagel, D.H.; Kang, S.E.; Bonaldi, K.; Doherty, C.J.; Ravelo, S.; Galli, M.; Ecker, J.R.; Kay, S.A. A Genome-Scale Resource for the Functional Characterization of Arabidopsis Transcription Factors. Cell Rep. 2014, 8, 622–632.
- Jiang, H.; Li, M.; Liang, N.; Yan, H.; Wei, Y.; Xu, X.; Liu, J.; Xu, Z.; Chen, F.; Wu, G. Molecular cloning and function analysis of the stay green gene in rice. Plant J. Cell Mol. Biol. 2007, 52, 197–209.
- Yang, J.; Worley, E.; Udvardi, M. A NAP-AAO3 Regulatory Module Promotes Chlorophyll Degradation via ABA Biosynthesis in Arabidopsis Leaves. Plant Cell 2014, 26, 4862–4874.
- Sakuraba, Y.; Han, S.-H.; Lee, S.-H.; Hörtensteiner, S.; Paek, N.-C. Arabidopsis NAC016 promotes chlorophyll breakdown by directly upregulating STAYGREEN1 transcription. Plant Cell Rep. 2015, 35, 155–166.
- Zhu, X.; Chen, J.; Xie, Z.; Gao, J.; Ren, G.; Gao, S.; Zhou, X.; Kuai, B. Jasmonic acid promotes degreening via MYC2/3/4-and ANAC019/055/072-mediated regulation of major chlorophyll catabolic genes. Plant J. 2015, 84, 597–610.
- Qiu, K.; Li, Z.; Yang, Z.; Chen, J.; Wu, S.; Zhu, X.; Gao, S.; Gao, J.; Ren, G.; Kuai, B.; et al. EIN3 and ORE1 Accelerate Degreening during Ethylene-Mediated Leaf Senescence by Directly Activating Chlorophyll Catabolic Genes in Arabidopsis. PLoS Genet. 2015, 11, e1005399.
- Robatzek, S.; Somssich, I.E. Targets of AtWRKY6 regulation during plant senescence and pathogen defense. Genes Dev. 2002, 16, 1139–1149.
- Robatzek, S.; Somssich, I.E. A new member of the Arabidopsis WRKY transcription factor family, AtWRKY6, is associated with both senescence- and defence-related processes. Plant J. 2001, 28, 123–133.
- Zhang, Y.; Liu, Z.; Wang, X.; Wang, J.; Fan, K.; Li, Z.; Lin, W. DELLA proteins negatively regulate dark-induced senescence and chlorophyll degradation in Arabidopsis through interaction with the transcription factor WRKY6. Plant Cell Rep. 2018, 37, 981–992.
- Zhang, D.; Zhu, Z.; Gao, J.; Zhou, X.; Zhu, S.; Wang, X.; Wang, X.; Ren, G.; Kuai, B. The NPR1-WRKY46-WRKY6 signaling cascade mediates probenazole/salicylic acid-elicited leaf senescence in Arabidopsis thaliana. J. Integr. Plant Biol. 2020.
- Tang, D.; Wang, G.; Zhou, J.-M. Receptor Kinases in Plant-Pathogen Interactions: More Than Pattern Recognition. Plant Cell 2017, 29, 618–637.
- Cano-Delgado, A.; Penfield, S.; Smith, C.; Catley, M.; Bevan, M. Reduced cellulose synthesis invokes lignification and defense responses in Arabidopsis thaliana. Plant J. 2003, 34, 351–362.
- Gish, L.A.; Clark, S.E. The RLK/Pelle family of kinases. Plant J. 2011, 66, 117–127.
- He, Z.H.; Cheeseman, I.; He, D.Z.; Kohorn, B.D. A cluster of five cell wall-associated receptor kinase genes, Wak1-5, are expressed in specific organs of Arabidopsis. Plant Mol. Biol. 1999, 39, 1189–1196.
- Verica, J.A.; He, Z.-H. The Cell Wall-Associated Kinase (WAK) and WAK-Like Kinase Gene Family. Plant Physiol. 2002, 129, 455–459.
- Wagner, T.A.; Kohorn, B.D. Wall-associated kinases are expressed throughout plant development and are required for cell expansion. Plant Cell 2001, 13, 303–318.
- Lally, D.; Ingmire, P.; Tong, H.Y.; He, Z.H. Antisense expression of a cell wall-associated protein kinase, WAK4, inhibits cell elongation and alters morphology. Plant Cell 2001, 13, 1317–1331.
- He, Z.; He, D.; Kohorn, B.D. Requirement for the induced expression of a cell wall associated receptor kinase for survival during the pathogen response. Plant J. 1998, 14, 55–63.
- Maleck, K.; Levine, A.; Eulgem, T.; Morgan, A.; Schmid, J.; Lawton, K.A.; Dangl, J.L.; Dietrich, R.A. The transcriptome of Arabidopsis thaliana during systemic acquired resistance. Nat. Genet. 2000, 26, 403–410.
- Schenk, P.M.; Kazan, K.; Wilson, I.; Anderson, J.P.; Richmond, T.; Somerville, S.C.; Manners, J.M. Coordinated plant defense responses in Arabidopsis revealed by microarray analysis. Proc. Natl. Acad. Sci. USA 2000, 97, 11655–11660.
- Meier, S.; Ruzvidzo, O.; Morse, M.; Donaldson, L.; Kwezi, L.; Gehring, C.A. The Arabidopsis Wall Associated Kinase-Like 10 Gene Encodes a Functional Guanylyl Cyclase and Is Co-Expressed with Pathogen Defense Related Genes. PLoS ONE 2010, 5, e8904.
- Newton, R.P.; Smith, C.J. Cyclic nucleotides. Phytochemistry 2004, 65, 2423–2437.
- Bot, P.; Mun, B.-G.; Imran, Q.M.; Hussain, A.; Lee, S.-U.; Loake, G.; Yun, B.-W. Differential expression of AtWAKL10 in response to nitric oxide suggests a putative role in biotic and abiotic stress responses. PeerJ 2019, 7, e7383.
- Decreux, A.; Messiaen, J. Wall-associated Kinase WAK1 Interacts with Cell Wall Pectins in a Calcium-induced Conformation. Plant Cell Physiol. 2005, 46, 268–278.
- Decreux, A.; Thomas, A.; Spies, B.; Brasseur, R.; Cutsem, P.; Messiaen, J. In vitro characterization of the homogalacturonan-binding domain of the wall-associated kinase WAK1 using site-directed mutagenesis. Phytochemistry 2006, 67, 1068–1079.
- Diévart, A.; Clark, S.E. Using mutant alleles to determine the structure and function of leucine-rich repeat receptor-like kinases. Curr. Opin. Plant Biol. 2003, 6, 507–516.
- Yu, Z.; Zhang, D.; Xu, Y.; Jin, S.; Zhang, L.; Zhang, S.; Yang, G.; Huang, J.; Yan, K.; Wu, C.; et al. CEPR2 phosphorylates and accelerates the degradation of PYR/PYLs in Arabidopsis. J. Exp. Bot. 2019, 70, 5457–5469.
- Chen, L.-J.; Wuriyanghan, H.; Zhang, Y.-Q.; Duan, K.-X.; Chen, H.-W.; Li, Q.-T.; Lu, X.; He, S.-J.; Ma, B.; Zhang, W.-K.; et al. An S-Domain Receptor-Like Kinase, OsSIK2, Confers Abiotic Stress Tolerance and Delays Dark-Induced Leaf Senescence in Rice. Plant Physiol. 2013, 163, 1752–1765.
- Xiao, D.; Cui, Y.; Xu, F.; Xu, X.; Gao, G.; Wang, Y.; Guo, Z.; Wang, D.; Wang, N.N. Senescence-Suppressed Protein Phosphatase Directly Interacts with the Cytoplasmic Domain of Senescence-Associated Receptor-Like Kinase and Negatively Regulates Leaf Senescence in Arabidopsis. Plant Physiol. 2015, 169, 1275–1291.
- Lee, I.C.; Hong, S.W.; Whang, S.S.; Lim, P.O.; Gil Nam, H.; Koo, J.C. Age-Dependent Action of an ABA-Inducible Receptor Kinase, RPK1, as a Positive Regulator of Senescence in Arabidopsis Leaves. Plant Cell Physiol. 2011, 52, 651–662.
- Li, X.; Ahmad, S.; Ali, A.; Guo, C.; Li, H.; Yu, J.; Zhang, Y.; Gao, X.; Guo, Y. Characterization of Somatic Embryogenesis Receptor-Like Kinase 4 as a Negative Regulator of Leaf Senescence in Arabidopsis. Cells 2019, 8, 50.
- Li, X.-P.; Gan, R.; Li, P.-L.; Ma, Y.-Y.; Zhang, L.-W.; Zhang, R.; Wang, Y.; Wang, N.N. Identification and functional characterization of a leucine-rich repeat receptor-like kinase gene that is involved in regulation of soybean leaf senescence. Plant Mol. Biol. 2006, 61, 829–844.
- Hou, X.; Tong, H.; Selby, J.; DeWitt, J.; Peng, X.; He, Z.-H. Involvement of a Cell Wall-Associated Kinase, WAKL4, in Arabidopsis Mineral Responses. Plant Physiol. 2005, 139, 1704–1716.
- Verica, J.A.; Chae, L.; Tong, H.; Ingmire, P.; He, Z.-H. Tissue-Specific and Developmentally Regulated Expression of a Cluster of Tandemly Arrayed Cell Wall-Associated Kinase-Like Kinase Genes in Arabidopsis. Plant Physiol. 2003, 133, 1732–1746.
- Larkan, N.J.; Ma, L.; Haddadi, P.; Buchwaldt, M.; Parkin, I.A.P.; Djavaheri, M.; Borhan, M.H. The Brassica napus wall-associated kinase-like (WAKL) gene Rlm9 provides race-specific blackleg resistance. Plant J. Cell Mol. Biol. 2020, 104, 892–900.
- Miya, A.; Albert, P.; Shinya, T.; Desaki, Y.; Ichimura, K.; Shirasu, K.; Narusaka, Y.; Kawakami, N.; Kaku, H.; Shibuya, N. CERK1, a LysM receptor kinase, is essential for chitin elicitor signaling in Arabidopsis. Proc. Natl. Acad. Sci. USA 2007, 104, 19613–19618.
- Heese, A.; Hann, D.R.; Gimenez-Ibanez, S.; Jones, A.M.E.; He, K.; Li, J.; Schroeder, J.I.; Peck, S.C.; Rathjen, J.P. The receptor-like kinase SERK3/BAK1 is a central regulator of innate immunity in plants. Proc. Natl. Acad. Sci. USA 2007, 104, 12217–12222.
- Guo, Y.; Gan, S.-S. Convergence and divergence in gene expression profiles induced by leaf senescence and 27 senescence-promoting hormonal, pathological and environmental stress treatments. Plant Cell Environ. 2011, 35, 644–655.
- Gregersen, P.L.; Holm, P.B. Transcriptome analysis of senescence in the flag leaf of wheat (Triticum aestivum L.). Plant Biotechnol. J. 2007, 5, 192–206.
- Besseau, S.; Li, J.; Palva, E.T. WRKY54 and WRKY70 co-operate as negative regulators of leaf senescence in Arabidopsis thaliana. J. Exp. Bot. 2012, 63, 2667–2679.
- Miao, Y.; Laun, T.; Zimmermann, P.; Zentgraf, U. Targets of the WRKY53 transcription factor and its role during leaf senescence in Arabidopsis. Plant Mol. Biol. 2004, 55, 853–867.
- Ülker, B.; Mukhtar, M.S.; Somssich, I.E. The WRKY70 transcription factor of Arabidopsis influences both the plant senescence and defense signaling pathways. Planta 2007, 226, 125–137.
- Zhou, X.; Jiang, Y.; Yu, D. WRKY22 transcription factor mediates dark-induced leaf senescence in Arabidopsis. Mol. Cells 2011, 31, 303–313.
- Asai, T.; Tena, G.; Plotnikova, J.; Willmann, M.R.; Chiu, W.-L.; Gomez-Gomez, L.; Boller, T.; Ausubel, F.M.; Sheen, J. MAP kinase signalling cascade in Arabidopsis innate immunity. Nat. Cell Biol. 2002, 415, 977–983.
- Van Verk, M.C.; Bol, J.F.; Linthorst, H.J. WRKY transcription factors involved in activation of SA biosynthesis genes. BMC Plant Biol. 2011, 11, 89.
- Tian, T.; Ma, L.; Liu, Y.; Xu, D.; Chen, Q.; Li, G. Arabidopsis FAR-RED ELONGATED HYPOCOTYL3 Integrates Age and Light Signals to Negatively Regulate Leaf Senescence. Plant Cell 2020, 32, 1574–1588.
- Hinckley, W.E.; Brusslan, J.A. Gene expression changes occurring at bolting time are associated with leaf senescence in Arabidopsis. Plant Direct 2020, 4, e00279.
- Morris, K.; Mackerness, S.A.-H.; Page, T.; John, C.F.; Murphy, A.M.; Carr, J.P.; Buchanan-Wollaston, V. Salicylic acid has a role in regulating gene expression during leaf senescence. Plant J. 2000, 23, 677–685.
- Chen, J.; Nolan, T.M.; Ye, H.; Zhang, M.; Tong, H.; Xin, P.; Chu, J.; Chu, C.; Li, Z.; Yin, Y. Arabidopsis WRKY46, WRKY54 and WRKY70 Transcription Factors are Involved in Brassinosteroid-Regulated Plant Growth and Drought Response. Plant Cell 2017, 29, 1425–1439.
- Li, J.; Besseau, S.; Toronen, P.; Sipari, N.; Kollist, H.; Holm, L.; Palva, E.T. Defense-related transcription factors WRKY70 and WRKY54 modulate osmotic stress tolerance by regulating stomatal aperture in Arabidopsis. New Phytol. 2013, 200, 455–472.
- Li, J.; Brader, G.; Palva, E.T. The WRKY70 Transcription Factor: A Node of Convergence for Jasmonate-Mediated and Salicylate-Mediated Signals in Plant Defense. Plant Cell 2004, 16, 319–331.
- Li, J.; Zhong, R.; Palva, E.T. WRKY70 and its homolog WRKY54 negatively modulate the cell wall-associated defenses to necrotrophic pathogens in Arabidopsis. PLoS ONE 2017, 12, e0183731.
- Chen, L.; Zhang, L.; Li, D.; Wang, F.; Yu, D. WRKY8 transcription factor functions in the TMV-cg defense response by mediating both abscisic acid and ethylene signaling in Arabidopsis. Proc. Natl. Acad. Sci. USA 2013, 110, E1963–E1971.
- Ishihama, N.; Yamada, R.; Yoshioka, M.; Katou, S.; Yoshioka, H. Phosphorylation of the Nicotiana benthamiana WRKY8 Transcription Factor by MAPK Functions in the Defense Response. Plant Cell 2011, 23, 1153–1170.
- Kim, S.-H.; Lee, J.-H.; Seo, K.-I.; Ryu, B.; Sung, Y.; Chung, T.; Deng, X.W.; Lee, J.-H. Characterization of a Novel DWD protein that participates in heat stress response in Arabidopsis. Mol. Cells 2014, 37, 833–840.
- Hsu, F.-C.; Chou, M.-Y.; Chou, S.-J.; Li, Y.-R.; Peng, H.-P.; Shih, M.-C. Submergence Confers Immunity Mediated by the WRKY22 Transcription Factor in Arabidopsis. Plant Cell 2013, 25, 2699–2713.
- Liao, C.-J.; Lai, Z.; Lee, S.; Yun, D.J.; Mengiste, T. Arabidopsis HOOKLESS1 regulates responses to pathogens and abscisic acid through interaction with MED18 and acetylation of WRKY33 and ABI5 chromatin. Plant Cell 2016, 28, 1662–1681.
- Chen, L.; Xiang, S.; Chen, Y.; Li, D.; Yu, D. Arabidopsis WRKY45 Interacts with the DELLA Protein RGL1 to Positively Regulate Age-Triggered Leaf Senescence. Mol. Plant 2017, 10, 1174–1189.
- Niu, F.; Cui, X.; Zhao, P.; Sun, M.; Yang, B.; Deyholos, M.K.; Li, Y.; Zhao, X.; Jiang, Y.-Q. WRKY42 transcription factor positively regulates leaf senescence through modulating SA and ROS synthesis in Arabidopsis thaliana. Plant J. 2020, 104, 171–184.
- Guo, P.; Li, Z.; Huang, P.; Li, B.; Fang, S.; Chu, J.; Guo, H. A Tripartite Amplification Loop Involving the Transcription Factor WRKY75, Salicylic Acid, and Reactive Oxygen Species Accelerates Leaf Senescence. Plant Cell 2017, 29, 2854–2870.
- Pasternak, T.; Groot, E.P.; Kazantsev, F.V.; Teale, W.; Omelyanchuk, N.; Kovrizhnykh, V.; Palme, K.; Mironova, V.V. Salicylic Acid Affects Root Meristem Patterning via Auxin Distribution in a Concentration-Dependent Manner. Plant Physiol. 2019, 180, 1725–1739.
- Chai, J.; Liu, J.; Zhou, J.; Xing, D. Mitogen-activated protein kinase 6 regulates NPR1 gene expression and activation during leaf senescence induced by salicylic acid. J. Exp. Bot. 2014, 65, 6513–6528.
- Goh, H.-H.; Sloan, J.; Dorca-Fornell, C.; Fleming, A. Inducible Repression of Multiple Expansin Genes Leads to Growth Suppression during Leaf Development. Plant Physiol. 2012, 159, 1759–1770.
- Cho, H.T.; Kende, H. Expression of expansin genes is correlated with growth in deepwater rice. Plant Cell 1997, 9, 1661–1671.
- Cho, H.-T.; Cosgrove, D.J. Altered expression of expansin modulates leaf growth and pedicel abscission in Arabidopsis thaliana. Proc. Natl. Acad. Sci. USA 2000, 97, 9783–9788.
- Zhao, M.-R.; Li, F.; Fang, Y.; Gao, Q.; Wang, W. Expansin-regulated cell elongation is involved in the drought tolerance in wheat. Protoplasma 2011, 248, 313–323.
- Qin, L.; Kudla, U.; Roze, E.H.A.; Goverse, A.; Popeijus, H.; Nieuwland, J.; Overmars, H.; Jones, J.T.; Schots, A.; Smant, G.; et al. Plant degradation: A nematode expansin acting on plants. Nature 2004, 427, 30.
- Tan, J.; Wang, M.; Shi, Z.; Miao, X. OsEXPA10 mediates the balance between growth and resistance to biotic stress in rice. Plant Cell Rep. 2018, 37, 993–1002.
- Ju, Y.-L.; Min, Z.; Yue, X.-F.; Zhang, Y.-L.; Zhang, J.-X.; Zhang, Z.-Q.; Fang, Y.-L. Overexpression of grapevine VvNAC08 enhances drought tolerance in transgenic Arabidopsis. Plant Physiol. Biochem. 2020, 151, 214–222.
- Collin, A.; Daszkowska-Golec, A.; Kurowska, M.; Szarejko, I. Barley ABI5 (Abscisic Acid INSENSITIVE 5) is Involved in Abscisic Acid-Dependent Drought Response. Front. Plant Sci. 2020, 11, 1138.
- Zhang, H.; Mao, X.; Wang, C.; Jing, R. Overexpression of a Common Wheat Gene TaSnRK2.8 Enhances Tolerance to Drought, Salt and Low Temperature in Arabidopsis. PLoS ONE 2010, 5, e16041.
- Xu, X.; Wan, W.; Jiang, G.; Xi, Y.; Huang, H.; Cai, J.; Chang, Y.; Duan, C.-G.; Mangrauthia, S.K.; Peng, X.; et al. Nucleocytoplasmic Trafficking of the Arabidopsis WD40 Repeat Protein XIW1 Regulates ABI5 Stability and Abscisic Acid Responses. Mol. Plant 2019, 12, 1598–1611.
- Wildermuth, M.C.; Dewdney, J.; Wu, G.; Ausubel, F.M. Isochorismate synthase is required to synthesize salicylic acid for plant defence. Nature 2001, 414, 562–565.
- Kinkema, M.; Fan, W.; Dong, X. Nuclear localization of NPR1 is required for activation of PR gene expression. Plant Cell 2000, 12, 2339–2350.
- Despres, C.; DeLong, C.; Glaze, S.; Liu, E.; Fobert, P.R. The Arabidopsis NPR1/NIM1 protein enhances the DNA binding activity of a subgroup of the TGA family of bZIP transcription factors. Plant Cell 2000, 12, 279–290.
- Wang, C.; Dai, S.; Zhang, Z.; Lao, W.; Wang, R.; Meng, X.; Zhou, X. Ethylene and salicylic acid synergistically accelerate leaf senescence in Arabidopsis. J. Integr. Plant Biol. 2021, 2021.





