| Version | Summary | Created by | Modification | Content Size | Created at | Operation |
|---|---|---|---|---|---|---|
| 1 | Frank Weinberg | + 2880 word(s) | 2880 | 2020-12-31 11:17:14 | | | |
| 2 | Karina Chen | Meta information modification | 2880 | 2021-01-04 04:44:28 | | | | |
| 3 | Karina Chen | + 1 word(s) | 2881 | 2021-01-04 05:03:20 | | |
Video Upload Options
The lung microbiome has recently emerged as a major mediator of host inflammation and pathogenesis. Understanding how the lung microbiota exerts its effects on lung cancer and the tumor microenvironment will allow for novel development of therapies.
1. The Lung Microbiome in Health
For many decades it was believed the lung represented a sterile environment. With the advent of culture-independent techniques and high-throughput sequencing of the 16S rRNA gene, a small highly conserved cistron of the bacterial genome, it became possible to identify the presence of bacterial communities at the genus or species-level phylogeny. In a landmark study in 2010, the authors published the first application of culture-independent techniques to identify microbiota present within the lungs of healthy patients as well as patients with asthma and chronic obstructive pulmonary disease (COPD) . To date, there are over 30 published studies demonstrating the presence of bacterial communities in the lower respiratory tracts of healthy persons using modern culture-independent techniques . In healthy lungs, the microbial composition is determined by the balance between immigration and elimination of bacteria. Bacterial immigration occurs through inhalation of atmospheric air which can contain upwards of 104–106 bacterial cells per cubic meter as well as microaspiration. Microaspiration is likely the primary source of immigration evidenced by overlap between the oral and lung microbiota. It is well known that subclinical microaspirations occur in healthy persons. Elimination of microbes from the lungs is a dynamic process that includes mechanical as well as immunological processes. Healthy airways contain ciliated epithelia that help to propel microbes proximally while the act of coughing represents an additional mechanism to expel microbes from the respiratory tract. It is well appreciated that the lung and airways contain specific innate and adaptive immunological defenses that allow for recognition and clearance of microbes. Further, the healthy lung is a heterogeneous environment with regional environmental variation that can influence bacterial composition. These variations include oxygen tension, pH, relative blood perfusion, relative alveolar ventilation, temperature, epithelial cell structure, deposition of inhaled particles and concentration and activity of inflammatory cells. Dickson et al. proposed the adapted island model which postulates that in health the respiratory tract is one continuous ecosystem that is dynamic and varies with microbes originating from the single source of the upper respiratory tract and immigrating to the lower respiratory tract. The number of microbial species at a specific site in the respiratory tract is a function of immigration and elimination factors. Dickson et al. validated this model, demonstrating that community richness (the number of species in a specific ecological community) decreases with increasing distance from the upper respiratory tract.
In general, studies indicate that a relatively uniform microbiota composes the healthy lung with the presence of certain dominant taxonomic groups. The most abundant phyla are Bacteroidetes and Firmicutes with prominent genera including Prevotella, Veillonella and Streptococcus. In general, the microbiota of the healthy lung most closely resembles that of the mouth, likely a result of microaspirations that occur during sleep when cough and laryngeal reflexes are diminished. Within the lung of healthy persons there are considerable variations in environmental factors (described above); however, there is a relative lack of variation in the lung microbiome suggesting that the lung microbiota is in fact determined by the balance between immigration and elimination rather than specific growth factors. It has not been determined if the lung microbiome of healthy subjects varies geographically. One study assessed the lung microbiota from healthy subjects in eight US cities and found no evidence of geographic clustering [1].
2. The Lung Microbiome and Lung Cancer
Previous studies have linked the microbiota to various malignancies. More recently, studies performed by the Wargo group have correlated gut microbial composition to response to chemotherapy and immunotherapy in colon cancer models and advanced stage melanoma patients [2][3]. There have been a number of studies suggesting the gut microbiome plays an important role in various cancers, including lung cancer [19]. Further studies showed that the composition of the gut microbiota in lung cancer patients differed significantly from that of healthy control patients [4]. A systematic review of eight studies demonstrated that NSCLC patients treated with broad spectrum antibiotics prior to or during treatment with immune checkpoint inhibitors (ICI) had poorer clinical outcomes [5]. Indeed, this suggests that disruption of the gut microbiome with antibiotics may have an effect on response to ICI in NSCLC. It should be noted that antibiotic treatment would also have an effect on the lung microbiome as well. Additionally, many recent studies demonstrate the importance of the gut microbiome in modulating immune responses and changes in the gut microbiota alter immune responses and homeostasis in the respiratory system [6]. It is surmised that this cross-talk occurs through the production of metabolites, endotoxins and cytokines by the gut microbiota which travel to the respiratory mucosa through the bloodstream [7]. Some more recent studies suggest that this cross-talk is bidirectional, with the lung microbiota potentially mediating immunologic responses in the gut [8][9]. Given the abundance of data surrounding the gut-lung axis we chose to focus more fully on the effect of the lung microbiota directly upon lung tumorigenesis.
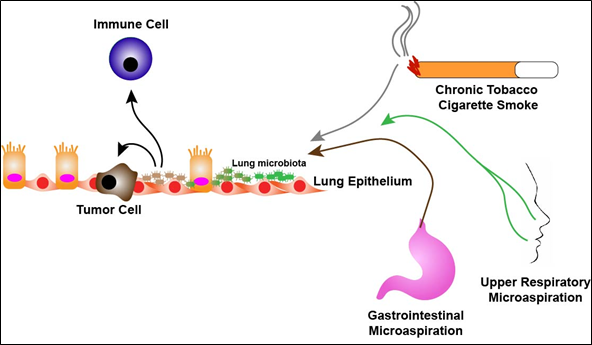
Characterization of the lung microbiome and its influences on lung cancer and treatment is in its infancy. Epidemiological studies have correlated Mycobacterium tuberculosis (TB) to lung cancer [10][11][12]. A systematic review of 41 studies demonstrated that a significantly increased lung cancer risk associated with pre-existing TB independent of tobacco use and exposure [11]. This suggests a direct link between TB and lung cancer and further suggests that the composition of the lung microbiome may have effects on tumorigenesis. In recent years, studies have demonstrated significant differences between the composition of the lung microbiome in lung cancer patients compared to healthy subjects or patients with alternative lung pathologies. In general, alpha diversity, the number (richness) and distribution (evenness) of taxa in a sample, is significantly higher in non-malignant lung tissues than tumor tissues while beta diversity, that is diversity in the microbial composition between different samples, is not significantly different between malignant and non-malignant tissues [13][14]. Other studies have determined specific taxa that are enriched in the airways of lung cancer patients. A study analyzing 216 lung aspirates collected from lung cancer patients demonstrated colonization of Gram negative bacteria such as Haemophilus influenza, Enterobacter and Escherichia coli [15]. It should be noted however, that the authors relied on culture-dependent assays for their analysis. In another study, sputum and oral samples were collected from female, never smoker, lung cancer patients (n = 8) as well as female, never smokers without cancer (n = 8). Samples taken from the lung cancer cohort had enrichment of Granulicatella, Abiotrophia and Streptococcus genera compared to the non-cancer cohort [16]. In addition, a study demonstrated enrichment of Veillonella, Neisseria, Capnocytophaga and Selenomonas in sputum collected from squamous cell and adenocarcinoma lung cancer patients (n = 20) as compared to non-cancer control subjects (n = 10) [17]. In a pilot study designed to identify potential bacterial biomarkers in lung cancer, the authors collected sputum from lung cancer patients (n = 4) and non-lung cancer subjects (n = 6) and found that seven specific bacterial species were present in all samples with significantly higher levels of enrichment of Streptococcus viridans in lung cancer samples [18]. More recently, bronchoalveolar lavage fluid (BALF) has been used to characterize the lung microbiome of the lower airways. Lee et al. collected BALF from 28 patients undergoing routine bronchoscopy for lung masses. Of the 28 patients, 20 were found to have lung cancer and 8 were diagnosed with benign diseases. Using culture-independent techniques the authors found increased relative abundance of the genera, Veillonella and Megasphaera, in lung cancer patients as well as Firmicutes and TM7 phyla as compared to patients with benign pathology [19]. Additionally, Tsay et al. collected airway brushings from patients undergoing routine diagnostic bronchoscopy for lung nodules from both the affected lung and contralateral unaffected lung. The authors report 39 subjects with lung cancer diagnoses, 36 subjects with non-cancer diagnoses and the inclusion of 10 healthy control subjects. They found that the lower airways of the lung cancer patients were enriched with oral tax (Streptococcus and Veillonella) as compared to non-cancer and healthy subjects. Interestingly, the uninvolved cancer airway samples shared many similar findings as seen in the involved cancer airway samples. It should be noted that this study included rigorous “control” specimens including buccal samples and bronchoscopy scope washes prior to procedures [36]. Together these findings support the presence of an altered lung microbiota in lung cancer patients and suggest enrichment with oral taxa likely a result of oral microaspirations (Figure 1).
Figure 1. The lung microbiome in disease. Chronic exposure to either cigarette smoke, aerodigestive or both, microaspiration over time leads to dysbiosis of health-associated lung microbiota (green) towards microbiota enriched with upper respiratory tract bacterial species (brown). Respiratory dysbiosis potentially leads to promotion/enhancement of tumorigenesis as well as alteration of the lung immune microenvironment.
3. The Lung Microbiome and the Immune System
The lung sits at the interface between the outside environment and the internal host physiology and as such plays a fundamental role in innate and initial adaptive immunity in order to protect the lung from pathogenic insults. This is not unlike other tissues, such as the gut. Multiple studies in the gut have linked the gut microbiota to mucosal immunity and modulation of host immunity. Studies have demonstrated that the gut commensal microbiota regulates the innate immune system [45,46]. It is then expected that alterations in the commensal microbiota could potentially have significant effects on immune tone in the host. Indeed, studies have demonstrated that the Gammaproteobacteria class utilizes inflammatory byproducts to survive and propagate under low oxygen conditions. During conditions such as chronic inflammation, Gammaproteobacteria can outcompete bacteria that are unable to metabolize inflammatory byproducts for survival [20]. Gammaproteobacteria can use reactive nitrogen species, a byproduct of many inflammatory cells, as a terminal electron acceptor to support growth under conditions of inflammation [20][21][22]. Therefore, the conditions of the microenvironment have the potential to enrich for potentially pathogenic bacteria (i.e., Gammaproteobacteria) which in turn promote continued or chronic inflammation. As discussed previously, the microbiota composition of the lower airways in healthy lung is dominated by Bacteroidetes phylum that shifts towards Gammaproteobacteria (class which contains many lung-associated “pathogens”) in diseased airways. It is likely that in the lung the same mechanism of bacterial overgrowth occurs as it does in the gut with Gammaproteobacteria. This is evidenced in an number of studies in humans and mice which demonstrated that increased levels of Gammaproteobacteria in the lungs is associated with disease [23].
The lung has specialized alveolar macrophages (AM) and resident dendritic cells (DC) as well as other immune cells that monitor the lower airway[24]s for pathogenic insults. They are critical mediators of lung immune homeostasis ensuring that inflammatory and immune responses are activated in response to a pathogenic insult while dampening responses to harmless environmental stimuli. In general, the lung microenvironment is one of high immune tolerance. Both AMs and DCs stimulate the proliferation of regulatory T cells (Treg) and release prostaglandin E2 (PGE2), tumor growth factor-beta (TGF-B) and interleukin-10 (IL-10) which leads to a tolerogenic state [25][26]. Furthermore, it is now appreciated that commensal lung microbiota plays an important role in promoting immune tolerance through its effect on resident lung immune cells. One fundamental question is how commensal bacteria are recognized and tolerated by the lungs and immune microenvironment. Antigen presenting cells (APCs) in the lung, namely AMs and DCs, and lung epithelial cells express pattern recognition receptors (PRRs), such as Toll-like receptors (TLRs) as well as others, that recognize molecules of host and microbial origin. Activation of PRRs induces expression of immune related genes encoding for inflammatory cytokines, type I interferons and antimicrobial peptides and leads to the initiation of innate and adaptive immune responses [25][26]. PRR ligands are present on both commensal microbiota as well as pathogenic bacteria, however, commensal PRR ligands are thought be less agonistic than pathogenic ligands [27][28]. It is also appreciated that all innate immune cells are able to decipher between pro-inflammatory or danger signals produced by a pathogenic insult versus tolerogenic signals produced by non-damaged tissue, dietary components and commensal bacteria [29]. There are several mechanisms by which the commensal microbiota could allow for host tolerance. One way is through a mechanism in which commensal microbiota protect itself from immune detection by preventing the outgrowth and spread of potentially harmful microorganisms subsequently decreasing the risk of detection by the immune system [30]. It is also appreciated that the commensal microbiota is largely prevented from access to the host epithelium by mucous production and therefore commensal microbiota cannot stimulate epithelial cell PRRs [28].
On the other hand, pathogenic bacteria with virulence factors can easily breach the mucous layer and infiltrate the epithelium leading to an inflammatory response [31]. Together, this supports a model in which the host epithelia and immune microenvironment subscribe to an “ignorance is bliss” type model in which commensal microbiota is present but not detected. Furthermore, PRRs are not randomly distributed along mucosal surfaces and are strategically sequestered in areas where commensal bacteria are limited in their access [32][33]. To this point, the host and immune cells have developed mechanisms to tolerate commensal microbiota. Certain studies have shown that persistent PRR stimulation by microbiota derived signals preserve epithelial barrier integrity and TLR tolerance is achieved after persistent TLR stimulation [34][35][36]. Additionally, APCs continuously exposed to endotoxins leads to tolerogenic AMs and DCs [65,66]. Finally, PRR activation is also dependent on expression of dampening signals expressed by epithelial and immune cells, specificity for specific innate ligands and the cocktail of cytokines that shape a response to a specific PRR agonist [37].
As suggested, the lung microbiota is thought to play a role in immune tolerance by influencing APCs and Treg recruitment. Gollwitzer et al. found that a progressive shift from Gammaproteobacteria and Firmcutes towards Bacteroidetes in neonates induces increased expression of programmed-death ligand 1 (PD-L1) on dendritic cells which lead to necessary Treg development after birth [38]. Another study demonstrated that germ free (GF) mice sensitized and challenged with ovalbumin demonstrated increased airway reactivity and inflammation as compared with specific pathogen free (SPF) mice. Furthermore, when GF mice were reconstituted with commensal bacteria from the lungs of SPF mice there was decreased airway reactivity and inflammation. Additionally, GF mice were noted to have dysregulated DCs and AMs as compared to SPF mice, likely a result of the commensal lung microbiota’s ability to educate lung immune cells [39]. Further evidence of the lung microbiota’s ability to regulate host immune response is demonstrated in recent studies exploring the effect of viral insults and oral taxa on immune tone. Wang et al. examined the role of Staphylococcus aureus, an upper respiratory tract (URT) colonizer, and found that it was essential for augmenting resistance to lethal inflammatory responses to influenza viral challenges. SPF mice were found to have less S. aureus as compared to mice living in the natural environment and subsequently succumbed to death at higher rates due to induction of acute inflammation after influenza viral challenges. Furthermore, it was shown that S. aureus recruits monocytes into alveoli and induces polarization of AMs to an M2 phenotype leading to suppression of lethal inflammatory responses through release of anti-inflammatory molecules in response to influenza insult [40]. This study suggests that specific airway microbiota taxa can act as potential defenders against viral insults. In the age of Sars-Cov2 infection it would be interesting to analyze the lung microbiome from COVID-19 patients with hyper-inflammatory responses versus those with attenuated responses. Enrichment of specific bacterial taxa may be protective in those with attenuated inflammatory responses against Sars-Cov2 insult. Finally, Segal et al. demonstrated the presence of two distinct lower airway microbiota signatures in healthy subjects that correlated to differences in immune tone [41]. The authors found two distinct “pneumotypes”, one which is enriched with Prevotella and Veillonella (upper respiratory tract (URT) colonizers) termed supraglottic predominant taxa (SPT) and another population of persons termed background predominant taxa (BPT) characterized by low bacterial copy number whose BALF resembled background taxa environmental microbiota. Subjects with pneumotype SPT demonstrated increased numbers of lymphocytes in BALF, increased Th17 cells, IL-1α, IL-1β, fractalkine, IL-17, free fatty acids, inflammatory pathway mRNA and blunted TLR4 response. There was also decreased β-diversity and increased bacterial abundance [41]. Interestingly, the authors found that 45% of the healthy subjects are of pneumotype SPT which correlates to the percentage of microaspirators in the general population [42][43] again suggesting that microaspiration allows for the translocation of upper airway colonizers to the lower airway which affects lower airway inflammation and immune responses. This study suggests the importance of URT bacteria (Prevotella and Veillonella) in regulating the level of airway inflammation and Th17 immune activation in the lower airways. Additionally, co-culture experiments in which Prevotella and Haemophilus species were cultured with human DCs led to the observation that Prevotella could suppress Haemophilus induction of IL-12p70 in DCs leading to modulation of the immune response [44]. Another study demonstrated Prevotella-dominant airway microbiota was associated with development of more inflammatory prone macrophages [45]. This suggests that certain species of bacteria could actually modulate host immune response to other commensal or pathogenic bacteria. Given the microbiota’s effects on modulating lung inflammation, especially Th17 responses, it would not be surprising if these same upper respiratory commensals were playing a role in modulating tumor immunity. For example, a Prevotella-predominant microbiota in the lower airways could generate an immune microenvironment enriched with Th17 cell populations and IL-17 production which could support tumorigenesis. Indeed, IL-17A T cells have been linked to tumor invasion and metastasis in lung adenocarcinoma [46].
References
- Morris, A.; Beck, J.M.; Schloss, P.D.; Campbell, T.B.; Crothers, K.; Curtis, J.L.; Flores, S.C.; Fontenot, A.P.; Ghedin, E.; Huang, L.; et al. Comparison of the respiratory microbiome in healthy nonsmokers and smokers. Am. J. Respir. Crit. Care Med. 2013, 187, 1067–1075. [Google Scholar] [CrossRef] [PubMed]
- Gopalakrishnan, V.; Spencer, C.N.; Nezi, L.; Reuben, A.; Andrews, M.C.; Karpinets, T.V.; Prieto, P.A.; Vicente, D.; Hoffman, K.; Wei, S.C.; et al. Gut microbiome modulates response to anti-PD-1 immunotherapy in melanoma patients. Science 2018, 359, 97–103. [Google Scholar] [CrossRef] [PubMed]
- Geller, L.T.; Barzily-Rokni, M.; Danino, T.; Jonas, O.H.; Shental, N.; Nejman, D.; Gavert, N.; Zwang, Y.; Cooper, Z.A.; Shee, K.; et al. Potential role of intratumor bacteria in mediating tumor resistance to the chemotherapeutic drug gemcitabine. Science 2017, 357, 1156–1160. [Google Scholar] [CrossRef]
- Carbone, C.; Piro, G.; Di Noia, V.; D’Argento, E.; Vita, E.; Ferrara, M.G.; Pilotto, S.; Milella, M.; Cammarota, G.; Gasbarrini, A.; et al. Lung and Gut Microbiota as Potential Hidden Driver of Immunotherapy Efficacy in Lung Cancer. Mediators Inflamm. 2019, 2019, 7652014. [Google Scholar] [CrossRef]
- Elkrief, A.; Derosa, L.; Kroemer, G.; Zitvogel, L.; Routy, B. The negative impact of antibiotics on outcomes in cancer patients treated with immunotherapy: A new independent prognostic factor? Ann. Oncol. 2019, 30, 1572–1579. [Google Scholar] [CrossRef] [PubMed]
- Voynow, J.A.; Hendricks-Munoz, K.D. The Gut-Lung Axis and Pulmonary Responses to Ozone. Am. J. Respir. Cell Mol. Biol. 2018, 59, 281–282. [Google Scholar] [CrossRef] [PubMed]
- Zhou, A.; Lei, Y.; Tang, L.; Hu, S.; Yang, M.; Wu, L.; Yang, S.; Tang, B. Gut microbiota: The emerging link to lung homeostasis and disease. J. Bacteriol. 2020. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Li, F.; Wei, H.; Lian, Z.X.; Sun, R.; Tian, Z. Respiratory influenza virus infection induces intestinal immune injury via microbiota-mediated Th17 cell-dependent inflammation. J. Exp. Med. 2014, 211, 2397–2410. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Tian, Z. How lung infection leads to gut injury. Oncotarget 2015, 6, 42394–42395. [Google Scholar] [CrossRef] [PubMed]
- Pilaniya, V.; Gera, K.; Kunal, S.; Shah, A. Pulmonary tuberculosis masquerading as metastatic lung disease. Eur. Respir. Rev. 2016, 25, 97–98. [Google Scholar] [CrossRef]
- Liang, H.Y.; Li, X.L.; Yu, X.S.; Guan, P.; Yin, Z.H.; He, Q.C.; Zhou, B.S. Facts and fiction of the relationship between preexisting tuberculosis and lung cancer risk: A systematic review. Int. J. Cancer 2009, 125, 2936–2944. [Google Scholar] [CrossRef]
- Pallis, A.G.; Syrigos, K.N. Lung cancer in never smokers: Disease characteristics and risk factors. Crit. Rev. Oncol. Hematol. 2013, 88, 494–503. [Google Scholar] [CrossRef]
- Yu, G.; Gail, M.H.; Consonni, D.; Carugno, M.; Humphrys, M.; Pesatori, A.C.; Caporaso, N.E.; Goedert, J.J.; Ravel, J.; Landi, M.T. Characterizing human lung tissue microbiota and its relationship to epidemiological and clinical features. Genome Biol. 2016, 17, 163. [Google Scholar] [CrossRef]
- Morgan, X.C.; Huttenhower, C. Chapter 12: Human microbiome analysis. PLoS Comput. Biol. 2012, 8, e1002808. [Google Scholar] [CrossRef]
- Laroumagne, S.; Lepage, B.; Hermant, C.; Plat, G.; Phelippeau, M.; Bigay-Game, L.; Lozano, S.; Guibert, N.; Segonds, C.; Mallard, V.; et al. Bronchial colonisation in patients with lung cancer: A prospective study. Eur. Respir. J. 2013, 42, 220–229. [Google Scholar] [CrossRef] [PubMed]
- Hosgood, H.D., 3rd; Sapkota, A.R.; Rothman, N.; Rohan, T.; Hu, W.; Xu, J.; Vermeulen, R.; He, X.; White, J.R.; Wu, G.; et al. The potential role of lung microbiota in lung cancer attributed to household coal burning exposures. Environ. Mol. Mutagen. 2014, 55, 643–651. [Google Scholar] [CrossRef]
- Yan, X.; Yang, M.; Liu, J.; Gao, R.; Hu, J.; Li, J.; Zhang, L.; Shi, Y.; Guo, H.; Cheng, J.; et al. Discovery and validation of potential bacterial biomarkers for lung cancer. Am. J. Cancer Res. 2015, 5, 3111–3122. [Google Scholar]
- Cameron, S.J.S.; Lewis, K.E.; Huws, S.A.; Hegarty, M.J.; Lewis, P.D.; Pachebat, J.A.; Mur, L.A.J. A pilot study using metagenomic sequencing of the sputum microbiome suggests potential bacterial biomarkers for lung cancer. PLoS ONE 2017, 12, e0177062. [Google Scholar] [CrossRef]
- Lee, S.H.; Sung, J.Y.; Yong, D.; Chun, J.; Kim, S.Y.; Song, J.H.; Chung, K.S.; Kim, E.Y.; Jung, J.Y.; Kang, Y.A.; et al. Characterization of microbiome in bronchoalveolar lavage fluid of patients with lung cancer comparing with benign mass like lesions. Lung Cancer 2016, 102, 89–95. [Google Scholar] [CrossRef]
- Winter, S.E.; Winter, M.G.; Xavier, M.N.; Thiennimitr, P.; Poon, V.; Keestra, A.M.; Laughlin, R.C.; Gomez, G.; Wu, J.; Lawhon, S.D.; et al. Host-derived nitrate boosts growth of E. coli in the inflamed gut. Science 2013, 339, 708–711. [Google Scholar] [CrossRef]
- Winter, S.E.; Baumler, A.J. Dysbiosis in the inflamed intestine: Chance favors the prepared microbe. Gut Microbes 2014, 5, 71–73. [Google Scholar] [CrossRef]
- Huffnagle, G.B.; Dickson, R.P.; Lukacs, N.W. The respiratory tract microbiome and lung inflammation: A two-way street. Mucosal Immunol. 2017, 10, 299–306. [Google Scholar] [CrossRef]
- Scales, B.S.; Dickson, R.P.; Huffnagle, G.B. A tale of two sites: How inflammation can reshape the microbiomes of the gut and lungs. J. Leukoc. Biol. 2016, 100, 943–950. [Google Scholar] [CrossRef]
- Hussell, T.; Bell, T.J. Alveolar macrophages: Plasticity in a tissue-specific context. Nat. Rev. Immunol. 2014, 14, 81–93. [Google Scholar] [CrossRef] [PubMed]
- Sellge, G.; Kufer, T.A. PRR-signaling pathways: Learning from microbial tactics. Semin. Immunol. 2015, 27, 75–84. [Google Scholar] [CrossRef] [PubMed]
- Baral, P.; Batra, S.; Zemans, R.L.; Downey, G.P.; Jeyaseelan, S. Divergent functions of Toll-like receptors during bacterial lung infections. Am. J. Respir. Crit. Care Med. 2014, 190, 722–732. [Google Scholar] [CrossRef] [PubMed]
- Steimle, A.; Autenrieth, I.B.; Frick, J.S. Structure and function: Lipid A modifications in commensals and pathogens. Int. J. Med. Microbiol. 2016, 306, 290–301. [Google Scholar] [CrossRef] [PubMed]
- Sommariva, M.; Le Noci, V.; Bianchi, F.; Camelliti, S.; Balsari, A.; Tagliabue, E.; Sfondrini, L. The lung microbiota: Role in maintaining pulmonary immune homeostasis and its implications in cancer development and therapy. Cell. Mol. Life Sci. 2020. [Google Scholar] [CrossRef] [PubMed]
- Swiatczak, B.; Cohen, I.R. Gut feelings of safety: Tolerance to the microbiota mediated by innate immune receptors. Microbiol. Immunol. 2015, 59, 573–585. [Google Scholar] [CrossRef] [PubMed]
- Khan, R.; Petersen, F.C.; Shekhar, S. Commensal Bacteria: An Emerging Player in Defense Against Respiratory Pathogens. Front. Immunol. 2019, 10, 1203. [Google Scholar] [CrossRef] [PubMed]
- Artis, D. Epithelial-cell recognition of commensal bacteria and maintenance of immune homeostasis in the gut. Nat. Rev. Immunol. 2008, 8, 411–420. [Google Scholar] [CrossRef] [PubMed]
- Ortega-Cava, C.F.; Ishihara, S.; Rumi, M.A.; Kawashima, K.; Ishimura, N.; Kazumori, H.; Udagawa, J.; Kadowaki, Y.; Kinoshita, Y. Strategic compartmentalization of Toll-like receptor 4 in the mouse gut. J. Immunol. 2003, 170, 3977–3985. [Google Scholar] [CrossRef] [PubMed]
- Hornef, M.W.; Normark, B.H.; Vandewalle, A.; Normark, S. Intracellular recognition of lipopolysaccharide by toll-like receptor 4 in intestinal epithelial cells. J. Exp. Med. 2003, 198, 1225–1235. [Google Scholar] [CrossRef] [PubMed]
- Park, J.H.; Kotani, T.; Konno, T.; Setiawan, J.; Kitamura, Y.; Imada, S.; Usui, Y.; Hatano, N.; Shinohara, M.; Saito, Y.; et al. Promotion of Intestinal Epithelial Cell Turnover by Commensal Bacteria: Role of Short-Chain Fatty Acids. PLoS ONE 2016, 11, e0156334. [Google Scholar] [CrossRef] [PubMed]
- Lopetuso, L.R.; Scaldaferri, F.; Petito, V.; Gasbarrini, A. Commensal Clostridia: Leading players in the maintenance of gut homeostasis. Gut Pathog. 2013, 5, 23. [Google Scholar] [CrossRef] [PubMed]
- Pena, O.M.; Pistolic, J.; Raj, D.; Fjell, C.D.; Hancock, R.E. Endotoxin tolerance represents a distinctive state of alternative polarization (M2) in human mononuclear cells. J. Immunol. 2011, 186, 7243–7254. [Google Scholar] [CrossRef]
- Couper, K.N.; Blount, D.G.; Riley, E.M. IL-10: The master regulator of immunity to infection. J. Immunol. 2008, 180, 5771–5777. [Google Scholar] [CrossRef]
- Gollwitzer, E.S.; Saglani, S.; Trompette, A.; Yadava, K.; Sherburn, R.; McCoy, K.D.; Nicod, L.P.; Lloyd, C.M.; Marsland, B.J. Lung microbiota promotes tolerance to allergens in neonates via PD-L1. Nat. Med. 2014, 20, 642–647. [Google Scholar] [CrossRef]
- Herbst, T.; Sichelstiel, A.; Schar, C.; Yadava, K.; Burki, K.; Cahenzli, J.; McCoy, K.; Marsland, B.J.; Harris, N.L. Dysregulation of allergic airway inflammation in the absence of microbial colonization. Am. J. Respir. Crit. Care Med. 2011, 184, 198–205. [Google Scholar] [CrossRef]
- Wang, J.; Li, F.; Sun, R.; Gao, X.; Wei, H.; Li, L.J.; Tian, Z. Bacterial colonization dampens influenza-mediated acute lung injury via induction of M2 alveolar macrophages. Nat. Commun. 2013, 4, 2106. [Google Scholar] [CrossRef]
- Segal, L.N.; Clemente, J.C.; Tsay, J.C.; Koralov, S.B.; Keller, B.C.; Wu, B.G.; Li, Y.; Shen, N.; Ghedin, E.; Morris, A.; et al. Enrichment of the lung microbiome with oral taxa is associated with lung inflammation of a Th17 phenotype. Nat. Microbiol. 2016, 1, 16031. [Google Scholar] [CrossRef] [PubMed]
- Gleeson, K.; Eggli, D.F.; Maxwell, S.L. Quantitative aspiration during sleep in normal subjects. Chest 1997, 111, 1266–1272. [Google Scholar] [CrossRef] [PubMed]
- Huxley, E.J.; Viroslav, J.; Gray, W.R.; Pierce, A.K. Pharyngeal aspiration in normal adults and patients with depressed consciousness. Am. J. Med. 1978, 64, 564–568. [Google Scholar] [CrossRef]
- Larsen, J.M.; Steen-Jensen, D.B.; Laursen, J.M.; Sondergaard, J.N.; Musavian, H.S.; Butt, T.M.; Brix, S. Divergent pro-inflammatory profile of human dendritic cells in response to commensal and pathogenic bacteria associated with the airway microbiota. PLoS ONE 2012, 7, e31976. [Google Scholar] [CrossRef] [PubMed]
- Bernasconi, E.; Pattaroni, C.; Koutsokera, A.; Pison, C.; Kessler, R.; Benden, C.; Soccal, P.M.; Magnan, A.; Aubert, J.D.; Marsland, B.J.; et al. Airway Microbiota Determines Innate Cell Inflammatory or Tissue Remodeling Profiles in Lung Transplantation. Am. J. Respir. Crit. Care Med. 2016, 194, 1252–1263. [Google Scholar] [CrossRef]
- Bao, Z.; Lu, G.; Cui, D.; Yao, Y.; Yang, G.; Zhou, J. IL-17A-producing T cells are associated with the progression of lung adenocarcinoma. Oncol. Rep. 2016, 36, 641–650. [Google Scholar] [CrossRef] [PubMed]
- Bao, Z.; Lu, G.; Cui, D.; Yao, Y.; Yang, G.; Zhou, J. IL-17A-producing T cells are associated with the progression of lung adenocarcinoma. Oncol. Rep. 2016, 36, 641–650. [Google Scholar] [CrossRef] [PubMed]